南方医科大学学报 ›› 2025, Vol. 45 ›› Issue (7): 1543-1553.doi: 10.12122/j.issn.1673-4254.2025.07.22
收稿日期:2025-02-26
出版日期:2025-07-20
发布日期:2025-07-17
通讯作者:
张虹丽
E-mail:wk19950303@smu.edu.cn;13926231602@163.com
作者简介:王 康,硕士,医师,E-mail: wk19950303@smu.edu.cn
基金资助:
Kang WANG1( ), Haibin LI1, Jing YU1, Yuan MENG2, Hongli ZHANG3(
), Haibin LI1, Jing YU1, Yuan MENG2, Hongli ZHANG3( )
)
Received:2025-02-26
Online:2025-07-20
Published:2025-07-17
Contact:
Hongli ZHANG
E-mail:wk19950303@smu.edu.cn;13926231602@163.com
摘要:
目的 探讨细胞外富含亮氨酸重复序列和纤连蛋白III型结构域的蛋白质1(ELFN1)在结肠癌中的表达、临床预后、基因功能、信号通路和生物学表型。 方法 运用R语言,通过公共数据库TCGA、GEO数据库分析33种癌谱ELFN1表达水平,并鉴定结肠癌中ELFN1的差异基因,通过基因功能注释和富集分析鉴定其相关信号通路;分析并评价ELFN1表达与临床病理特征的相关性;qPCR及Western blotting验证ELFN1在细胞株以及肠癌组织中的表达水平;Transwell、EDU实验分析ELFN1在SW480细胞的生物学表型。 结果 ELFN1在14种肿瘤中高表达,结肠癌中ELFN1的表达高于癌周组织(P<0.001);临床特征与预后分析结果显示,ELFN1 mRNA高表达的患者OS较差;Cox回归分析显示,ELFN1表达是OS的独立预后因素(P<0.05);ELFN1在肿瘤转移增殖中显著富集,进一步GSEA分析识别了与之相关的肿瘤信号通路;实验分析结果显示,肠癌细胞系ELFN1的表达水平高于正常细胞(P<0.05);细胞功能实验显示,敲低ELFN1可阻止肠癌SW480细胞增殖和转移(P<0.05)。 结论 ELFN1在结肠癌中异常高表达,并代表了不良的临床预后;并与肠癌增殖转移表型相关,提示ELFN1可能为结肠癌的预后标记物。
王康, 李海宾, 余靖, 孟源, 张虹丽. ELFN1高表达是结肠癌的预后生物标志物并促进结肠癌细胞的增殖转移[J]. 南方医科大学学报, 2025, 45(7): 1543-1553.
Kang WANG, Haibin LI, Jing YU, Yuan MENG, Hongli ZHANG. High expression of ELFN1 is a prognostic biomarker and promotes proliferation and metastasis of colorectal cancer cells[J]. Journal of Southern Medical University, 2025, 45(7): 1543-1553.
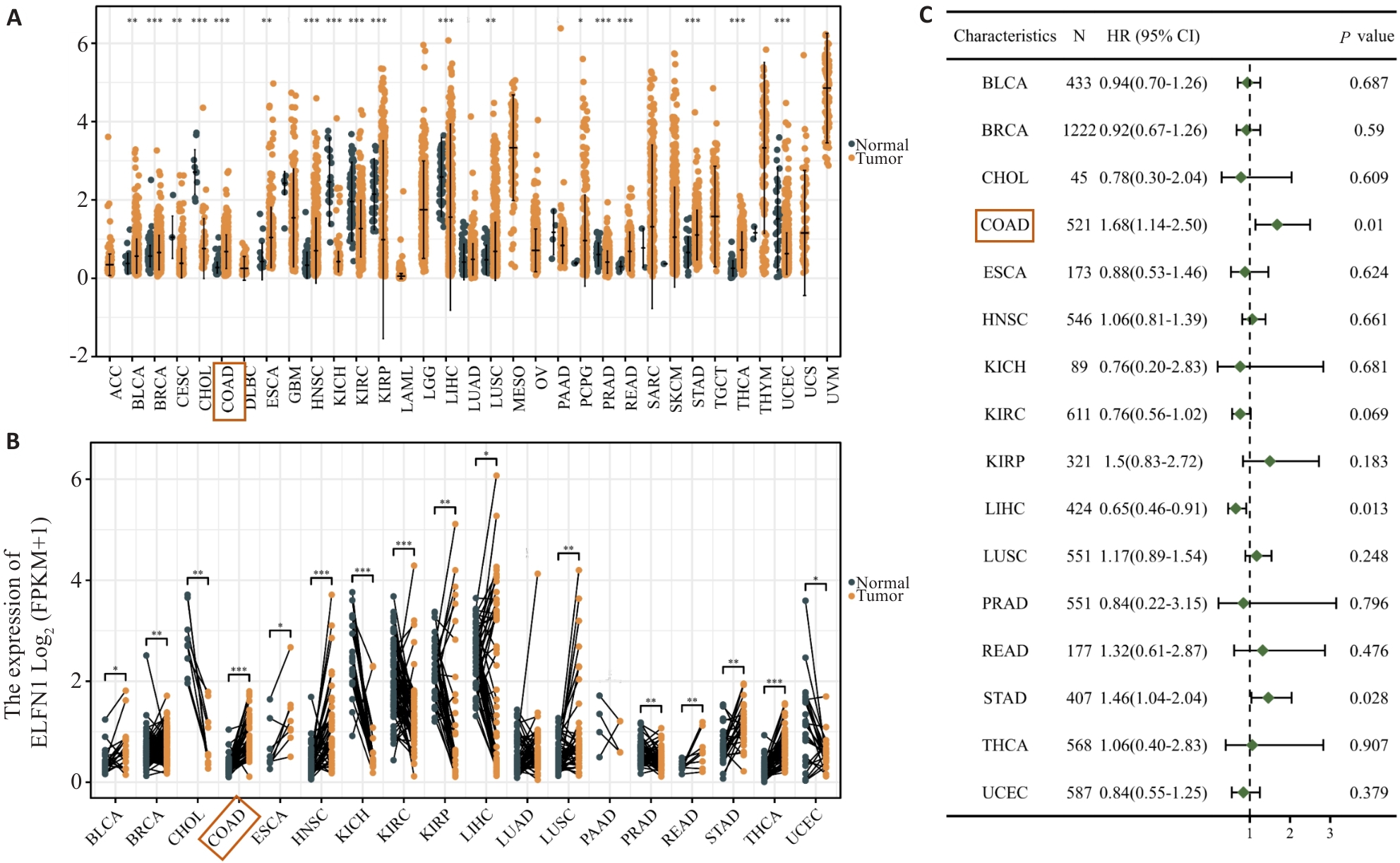
图1 基于TCGA数据库显示ELFN1在多种肿瘤中高表达并具有预后差异
Fig.1 Data from the TCGA database show high expression of ELFN1 in a variety of tumors, which has different prognostic implications. A: TCGA diffuse carcinoma analysis. B: TCGA pan-cancer pairing analysis. C: Forest map of the prognostic value of ELFN1 in different tumors. *P<0.05, **P<0.01, ***P<0.001.
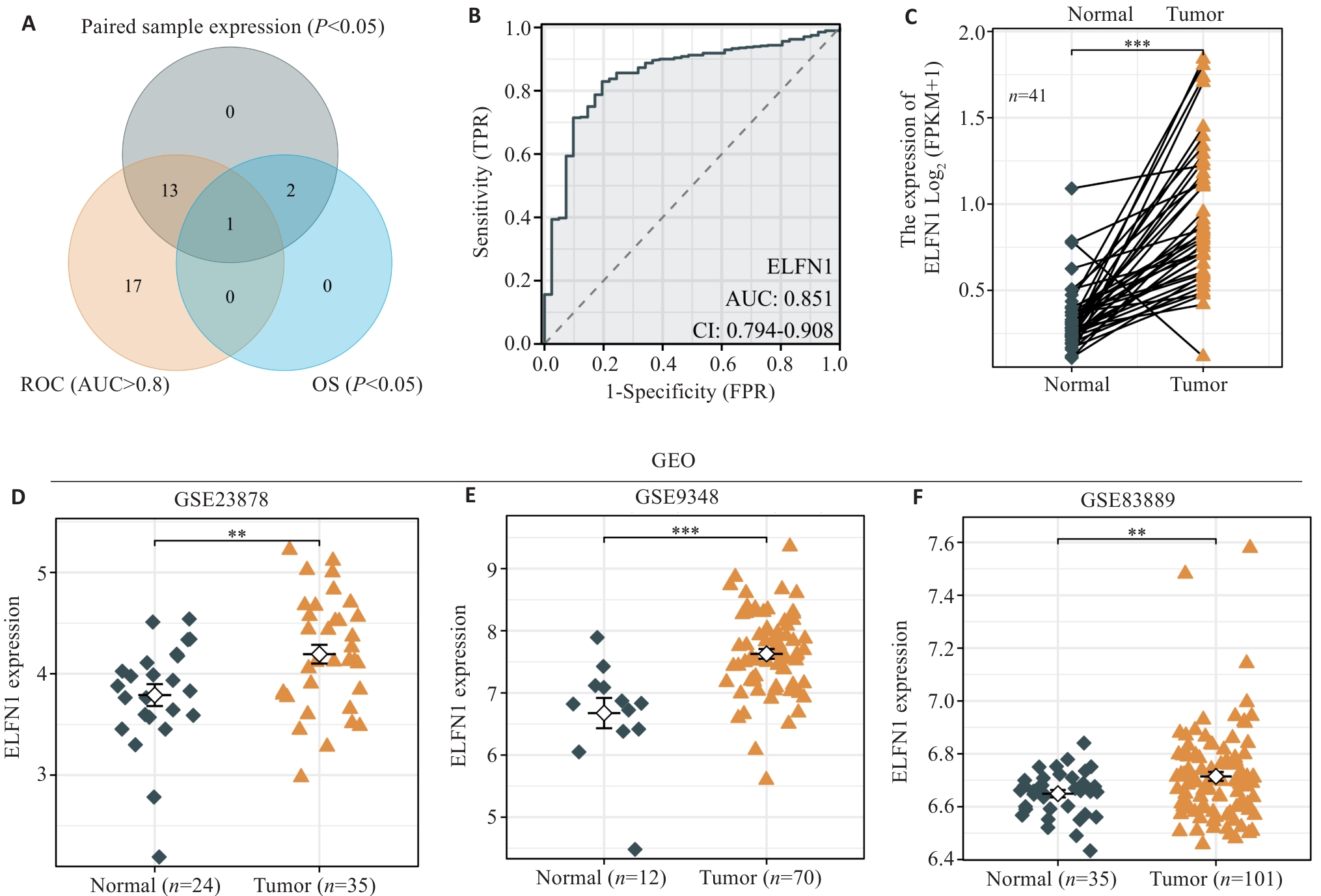
图2 ELFN1在结肠癌中具有较高诊断价值并在结肠癌中高表达
Fig.2 ELFN1 has a high diagnostic value in colon cancer and is highly expressed in colon cancer. A: Venn diagram of ELFN1 expression patterns and its prognosis and diagnostic value of colon cancer. B: ROC curve analysis showing an AUC value of ELFN1 in COAD of 0.851. C: High expression levels of ELFN1 in paired samples of TCGA. D-F: Multiple datasets in the GEO database indicate that the expression level of ELFN1 is higher tumor tissues than in normal tissues. *P<0.05, **P<0.01, ***P<0.001.
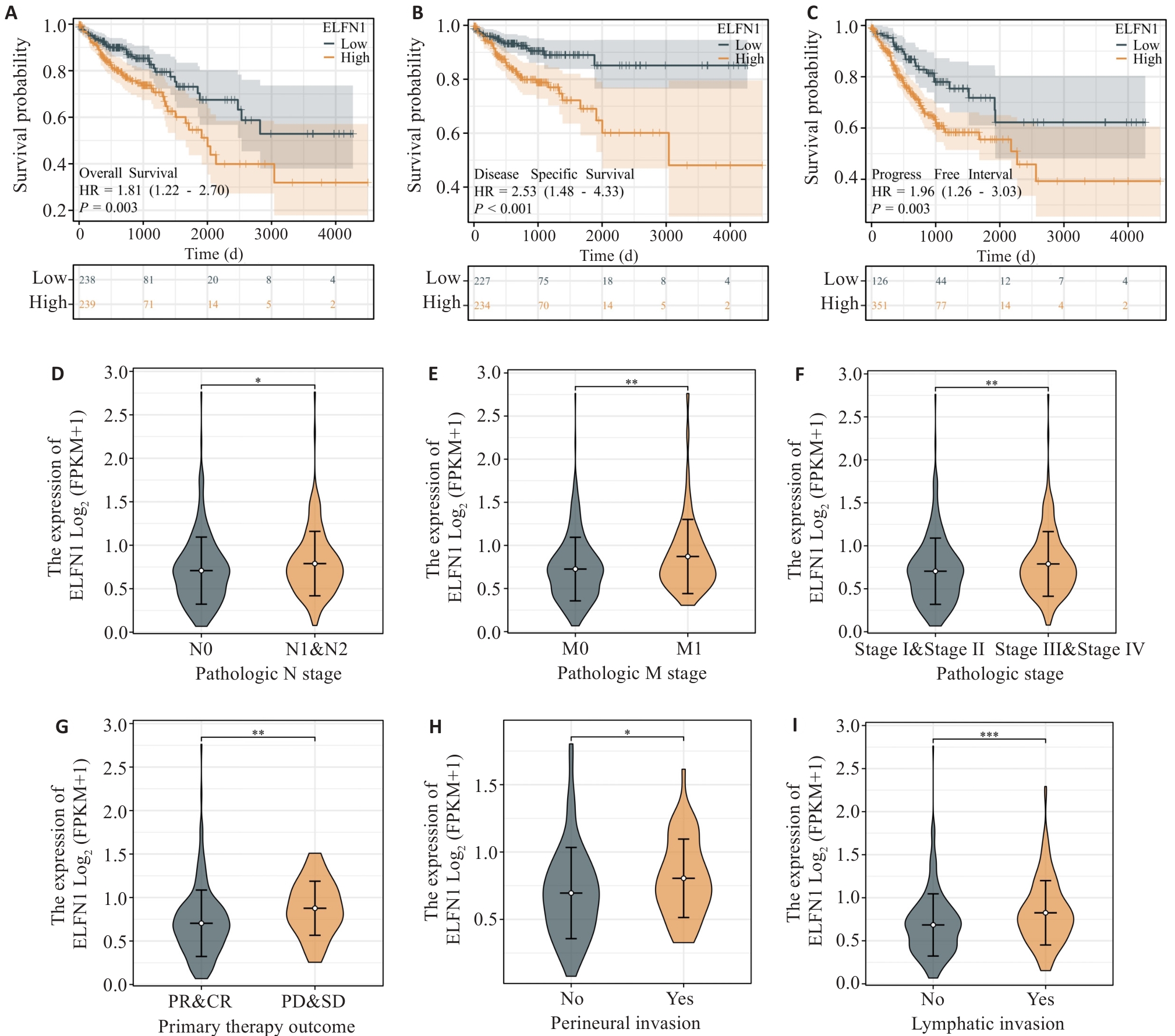
图3 ELFN1在结肠癌中的预后及临床价值
Fig.3 Prognostic and clinical significance of ELFN1 in colon cancer. A-C: Kaplan-Meier survival analysis reveals that COAD patients with high ELFN1 expression have shorter overall survival (A), disease-specific survival (B) and progression-free interval (C) than those with low ELFN1 expression. D-I: mRNA expression level of ELFN1 in COAD patients with different clinicopathological features. *P<0.05, **P<0.01, ***P<0.001.
| Characteristics | Total (n) | Univariate analysis | Multivariate analysis | |||
|---|---|---|---|---|---|---|
| Hazard ratio (95%CI) | P | Hazard ratio (95%CI) | P | |||
| Age (year) | 477 | |||||
| ≤65 | 194 | Reference | ||||
| >65 | 283 | 1.610 (1.052-2.463) | 0.028 | 3.417 (1.309-8.920) | 0.012 | |
| T stage | 476 | |||||
| T1&T2 | 94 | Reference | ||||
| T3&T4 | 382 | 3.072 (1.423-6.631) | 0.004 | 0.661 (0.120-3.637) | 0.634 | |
| N stage | 477 | |||||
| N0 | 283 | Reference | ||||
| N1&N2 | 194 | 2.592 (1.743-3.855) | <0.001 | 0.174 (0.016-1.839) | 0.146 | |
| M stage | 414 | |||||
| M0 | 348 | Reference | ||||
| M1 | 66 | 4.193 (2.683-6.554) | <0.001 | 5.463 (1.807-16.518) | 0.003 | |
| Pathologic stage | 466 | |||||
| Stage I&Stage II | 267 | Reference | ||||
| Stage III&Stage IV | 199 | 2.947 (1.942-4.471) | <0.001 | 14.791(1.177-185.875) | 0.037 | |
| Perineural invasion | 181 | |||||
| No | 135 | Reference | ||||
| Yes | 46 | 1.940 (0.982-3.832) | 0.056 | 0.736 (0.280-1.935) | 0.534 | |
| Lymphatic invasion | 433 | |||||
| No | 265 | Reference | ||||
| Yes | 168 | 2.450 (1.614-3.720) | <0.001 | 1.408 (0.557-3.556) | 0.470 | |
| ELFN1 | 477 | |||||
| Low | 238 | Reference | ||||
| High | 239 | 1.683 (1.135-2.496) | 0.010 | 3.284 (1.154-9.339) | 0.026 | |
表1 ELFN1在COAD中的单因素和多因素Cox回归分析
Tab.1 Univariate and multivariate Cox regression analyses of overall survival of COAD patients based on ELFN1 expression levels
| Characteristics | Total (n) | Univariate analysis | Multivariate analysis | |||
|---|---|---|---|---|---|---|
| Hazard ratio (95%CI) | P | Hazard ratio (95%CI) | P | |||
| Age (year) | 477 | |||||
| ≤65 | 194 | Reference | ||||
| >65 | 283 | 1.610 (1.052-2.463) | 0.028 | 3.417 (1.309-8.920) | 0.012 | |
| T stage | 476 | |||||
| T1&T2 | 94 | Reference | ||||
| T3&T4 | 382 | 3.072 (1.423-6.631) | 0.004 | 0.661 (0.120-3.637) | 0.634 | |
| N stage | 477 | |||||
| N0 | 283 | Reference | ||||
| N1&N2 | 194 | 2.592 (1.743-3.855) | <0.001 | 0.174 (0.016-1.839) | 0.146 | |
| M stage | 414 | |||||
| M0 | 348 | Reference | ||||
| M1 | 66 | 4.193 (2.683-6.554) | <0.001 | 5.463 (1.807-16.518) | 0.003 | |
| Pathologic stage | 466 | |||||
| Stage I&Stage II | 267 | Reference | ||||
| Stage III&Stage IV | 199 | 2.947 (1.942-4.471) | <0.001 | 14.791(1.177-185.875) | 0.037 | |
| Perineural invasion | 181 | |||||
| No | 135 | Reference | ||||
| Yes | 46 | 1.940 (0.982-3.832) | 0.056 | 0.736 (0.280-1.935) | 0.534 | |
| Lymphatic invasion | 433 | |||||
| No | 265 | Reference | ||||
| Yes | 168 | 2.450 (1.614-3.720) | <0.001 | 1.408 (0.557-3.556) | 0.470 | |
| ELFN1 | 477 | |||||
| Low | 238 | Reference | ||||
| High | 239 | 1.683 (1.135-2.496) | 0.010 | 3.284 (1.154-9.339) | 0.026 | |
| Characteristic | Total (n) | OR (95%CI) | P |
|---|---|---|---|
| Pathologic T stage (T3&T4 vs T1&T2) | 477 | 1.831 (0.975-3.600) | 0.069 |
| Pathologic N stage (N1&N2 vs N0) | 478 | 1.743 (1.078-2.851) | 0.025 |
| Pathologic M stage (M1 vs M0) | 415 | 2.425 (1.286-4.572) | 0.006 |
| Pathologic stage (Stage III&IV vs Stage I&II) | 467 | 1.787 (1.104-2.936) | 0.020 |
| Primary therapy outcome (PD&SD vs PR&CR) | 250 | 2.834 (1.120-7.152) | 0.025 |
| Perineural invasion (Yes vs No) | 181 | 2.675 (0.984-7.424) | 0.044 |
| Lymphatic invasion (Yes vs No) | 434 | 2.823 (1.653-4.934) | <0.001 |
表2 ELFN1在COAD中的Logistic回归分析
Tab.2 Logistic regression analysis of ELFN1
| Characteristic | Total (n) | OR (95%CI) | P |
|---|---|---|---|
| Pathologic T stage (T3&T4 vs T1&T2) | 477 | 1.831 (0.975-3.600) | 0.069 |
| Pathologic N stage (N1&N2 vs N0) | 478 | 1.743 (1.078-2.851) | 0.025 |
| Pathologic M stage (M1 vs M0) | 415 | 2.425 (1.286-4.572) | 0.006 |
| Pathologic stage (Stage III&IV vs Stage I&II) | 467 | 1.787 (1.104-2.936) | 0.020 |
| Primary therapy outcome (PD&SD vs PR&CR) | 250 | 2.834 (1.120-7.152) | 0.025 |
| Perineural invasion (Yes vs No) | 181 | 2.675 (0.984-7.424) | 0.044 |
| Lymphatic invasion (Yes vs No) | 434 | 2.823 (1.653-4.934) | <0.001 |
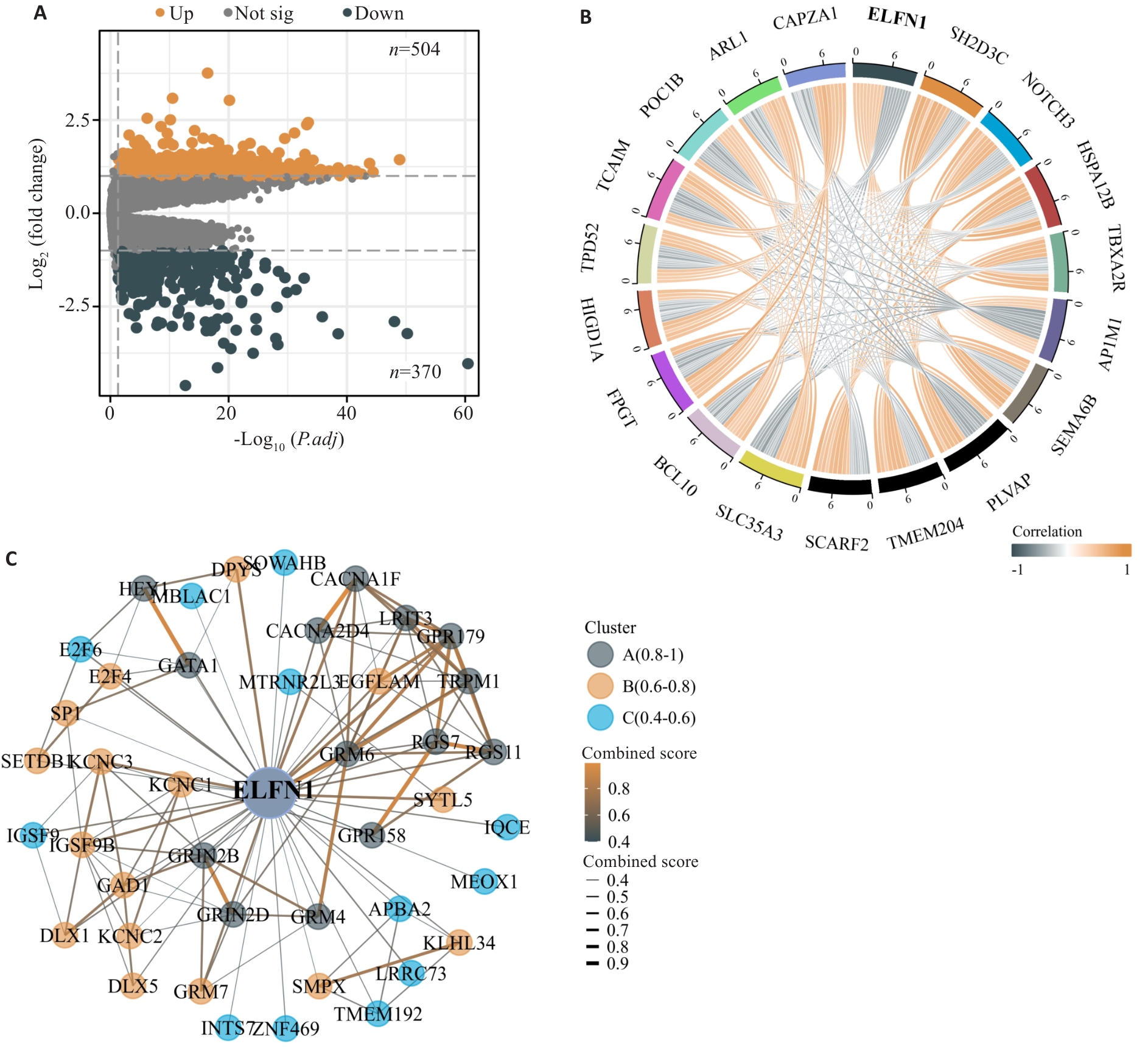
图4 ELFN1差异表达基因分析及PPI网络
Fig.4 Analysis of differentially expressed genes related to ELFN1 and construction of the PPI network. A: Volcano plot of differentially expressed genes (DEGs, |log2(FC)|>1, P.adj<0.05). B: The co-expression chord diagram showing the top 10 genes positively and negatively correlated with ELFN1. C: Molecular protein interaction network of ELFN1.
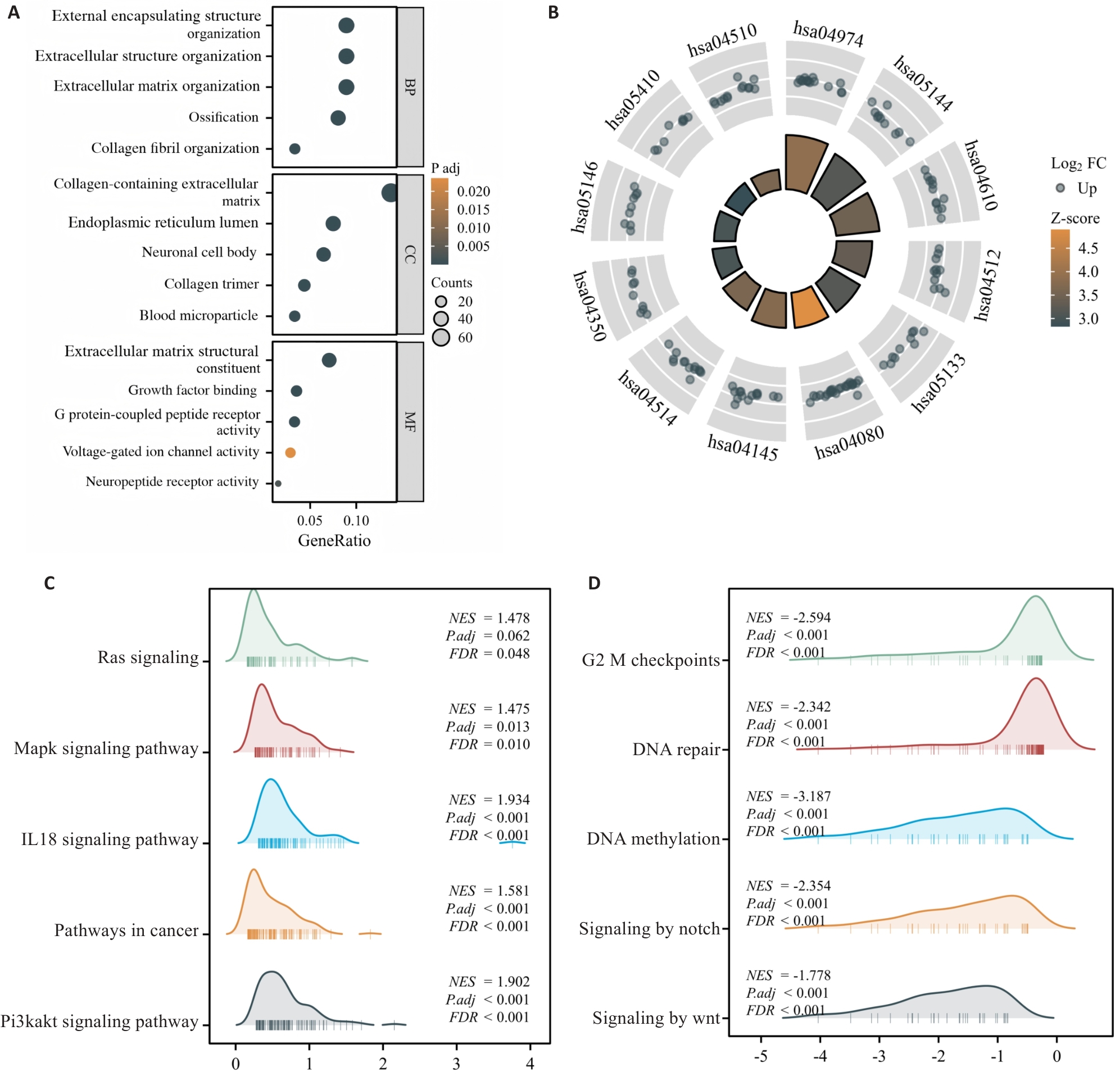
图5 ELFN1互作蛋白鉴定及基因功能注释分析
Fig.5 ELFN1 interaction protein identification and gene function annotation analysis. A: GO functional annotation analysis of biological process (BP), cellular component (CC) and molecular function (MF). B: Analysis of 10 KEGG signaling pathways related to ELFN1. C, D: The 10 GSEA pathways associated with ELFN1 in COAD.
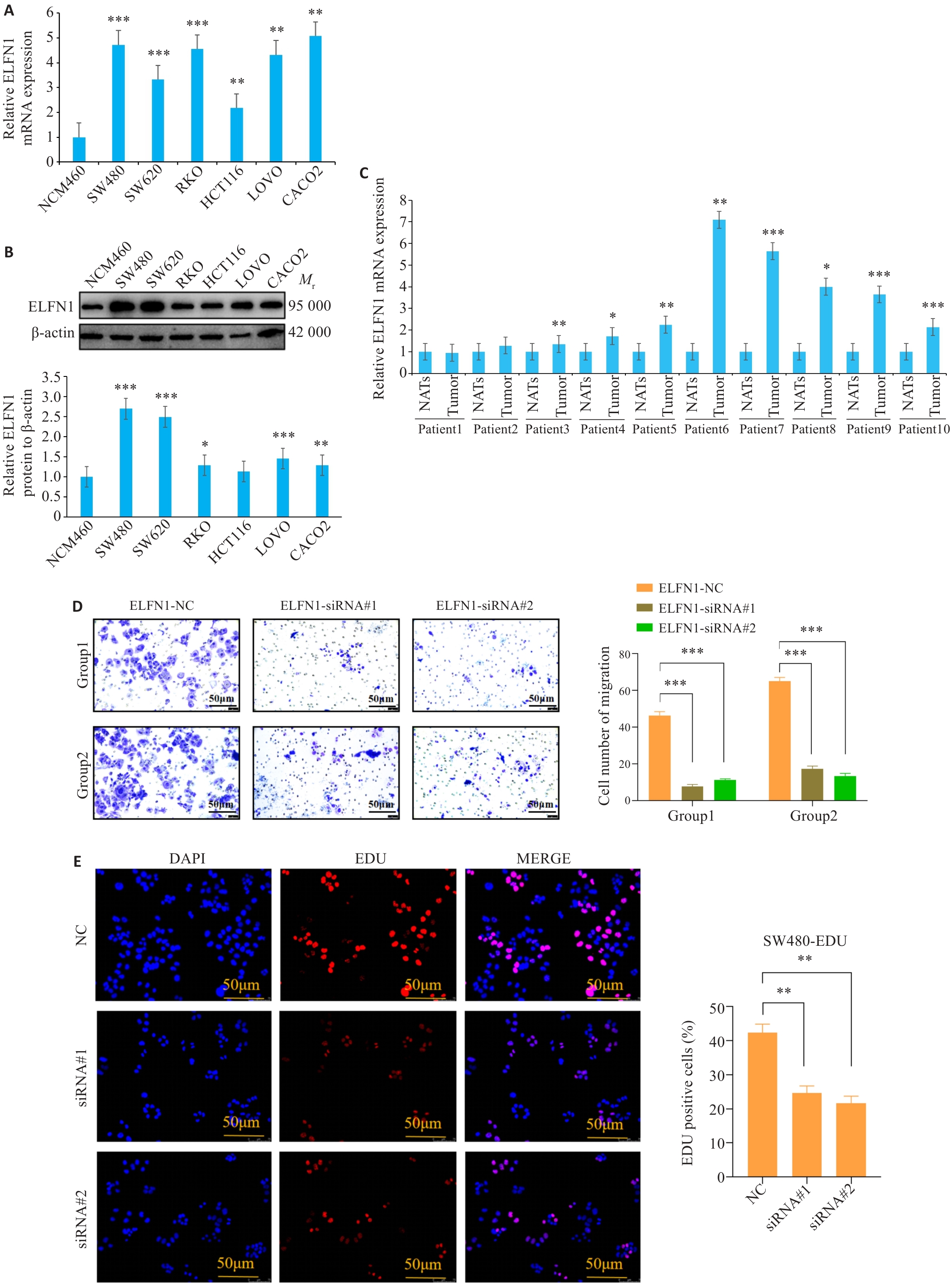
图6 ELFN1在COAD细胞系中的表达及功能
Fig.6 Expression and function of ELFN1 in COAD cell lines. A, B: ELFN1 expression levels in NCM460, SW480, SW620, RKO, HCT116, LOVO, and CACO2 cells detected by qRT-PCR (A) and Western blotting (B). *P<0.05, **P<0.01, ***P<0.001 vs NCM460 cells. C: Expression levels of ELFN1 in 10 pairs of colon cancer tissue samples (*P<0.05, **P<0.01, ***P<0.001 vs NATs). D: Transwell assay of SW480 cells transfected with si-ELFN1. E: EDU proliferation assay of SW480 cells transfected with si-ELFN1. **P<0.01, ***P<0.001.
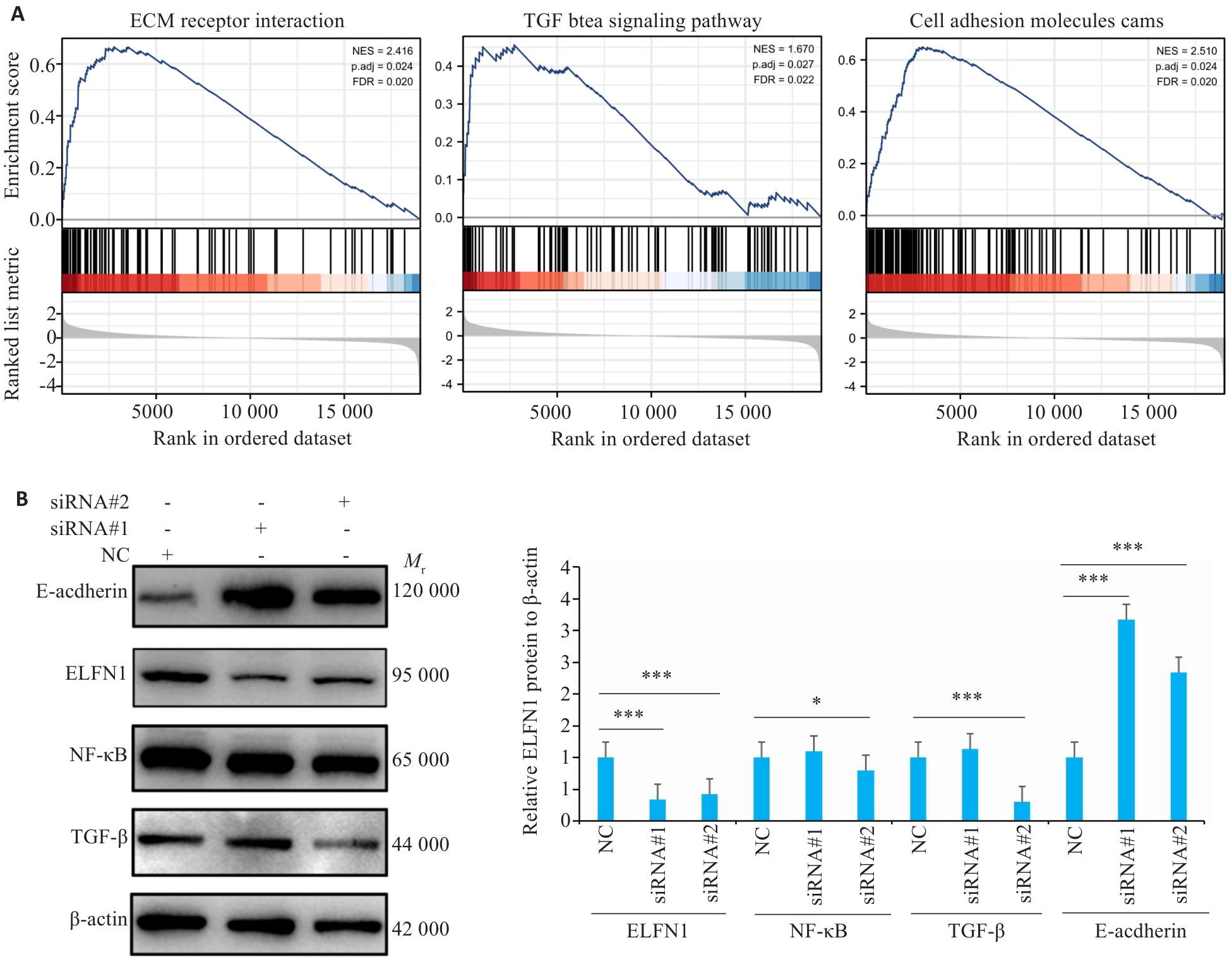
图7 ELFN1在结肠癌致癌信号通路分子机制研究
Fig.7 Molecular mechanism of ELFN1 in the oncogenic signaling pathway of colorectal cancer. A: Analysis of relevant carcinogenic pathways of ELFN1 in COAD. B: Western blotting for detecting protein expressions in COAD cells with ELFN1 knockdown. *P<0.05, ***P<0.001.
| [1] | Stachniak TJ, Sylwestrak EL, Scheiffele P, et al. Elfn1-induced constitutive activation of mGluR7 determines frequency-dependent recruitment of somatostatin interneurons[J]. J Neurosci, 2019, 39(23): 4461-74. doi:10.1523/jneurosci.2276-18.2019 |
| [2] | Dunn HA, Patil DN, Cao Y, et al. Synaptic adhesion protein ELFN1 is a selective allosteric modulator of group III metabotropic glutamate receptors in trans [J]. Proc Natl Acad Sci USA, 2018, 115(19): 5022-7. doi:10.1073/pnas.1722498115 |
| [3] | Tomioka NH, Yasuda H, Miyamoto H, et al. Elfn1 recruits presynaptic mGluR7 in trans and its loss results in seizures[J]. Nat Commun, 2014, 5: 4501. doi:10.1038/ncomms5501 |
| [4] | Girgenti MJ, Wang JW, Ji DJ, et al. Transcriptomic organization of the human brain in post-traumatic stress disorder[J]. Nat Neurosci, 2021, 24(1): 24-33. doi:10.1038/s41593-020-00748-7 |
| [5] | Zhang C, Lian H, Xie L, et al. LncRNA ELFN1-AS1 promotes esophageal cancer progression by up-regulating GFPT1 via sponging miR-183-3p[J]. Biol Chem, 2020, 401(9): 1053-61. doi:10.1515/hsz-2019-0430 |
| [6] | Jie YK, Ye L, Chen H, et al. ELFN1-AS1 accelerates cell proliferation, invasion and migration via regulating miR-497-3p/CLDN4 axis in ovarian cancer[J]. Bioengineered, 2020, 11(1): 872-82. doi:10.1080/21655979.2020.1797281 |
| [7] | Feng WG, Zhu RX, Ma JL, et al. LncRNA ELFN1-AS1 promotes retinoblastoma growth and invasion via regulating miR-4270/SBK1 axis[J]. Cancer Manag Res, 2021, 13: 1067-73. doi:10.2147/cmar.s281536 |
| [8] | Ma G, Li GC, Gou AJ, et al. Long non-coding RNA ELFN1-AS1 in the pathogenesis of pancreatic cancer[J]. Ann Transl Med, 2021, 9(10): 877. doi:10.21037/atm-21-2376 |
| [9] | Office FE. Retraction: Long non-coding RNA ELFN1-AS1 promoted colon cancer cell growth and migration via the miR-191-5p/special AT-rich sequence-binding protein 1 axis[J]. Front Oncol, 2023, 13: 1322464. doi:10.3389/fonc.2023.1322464 |
| [10] | Lei R, Feng LC, Hong D. ELFN1-AS1 accelerates the proliferation and migration of colorectal cancer via regulation of miR-4644/TRIM44 axis[J]. Cancer Biomark, 2020, 27(4): 433-43. doi:10.3233/cbm-190559 |
| [11] | Li YM, Gan YQ, Liu JX, et al. Downregulation of MEIS1 mediated by ELFN1-AS1/EZH2/DNMT3a axis promotes tumorigenesis and oxaliplatin resistance in colorectal cancer[J]. Signal Transduct Target Ther, 2022, 7(1): 87. doi:10.1038/s41392-022-00902-6 |
| [12] | Dekker E, Tanis PJ, Vleugels JLA, et al. Colorectal cancer[J]. Lancet, 2019, 394(10207): 1467-80. doi:10.1016/s0140-6736(19)32319-0 |
| [13] | Eng C, Jácome AA, Agarwal R, et al. A comprehensive framework for early-onset colorectal cancer research[J]. Lancet Oncol, 2022, 23(3): e116-28. doi:10.1016/s1470-2045(21)00588-x |
| [14] | Libé-Philippot B, Lejeune A, Wierda K, et al. LRRC37B is a human modifier of voltage-gated sodium channels and axon excitability in cortical neurons[J]. Cell, 2023, 186(26): 5766-83.e25. doi:10.1016/j.cell.2023.11.028 |
| [15] | Huang JX, Yuan WW, Chen BB, et al. lncRNA ELFN1-AS1 upregulates TRIM29 by suppressing miR-211-3p to promote gastric cancer progression[J]. Acta Biochim Biophys Sin (Shanghai), 2023, 55(3): 484-97. doi:10.3724/abbs.2023023 |
| [16] | Deng X, Kong FY, Li S, et al. A KLF4/PiHL/EZH2/HMGA2 regulatory axis and its function in promoting oxaliplatin-resistance of colorectal cancer[J]. Cell Death Dis, 2021, 12(5): 485. doi:10.1038/s41419-021-03753-1 |
| [17] | Huang L, Xiao W, Wang Y, et al. Metabotropic glutamate receptors (mGluRs) in epileptogenesis: an update on abnormal mGluRs signaling and its therapeutic implications[J]. Neural Regen Res, 2024, 19(2): 360-8. doi:10.4103/1673-5374.379018 |
| [18] | Becker JAJ, Pellissier LP, Corde Y, et al. Facilitating mGluR4 activity reverses the long-term deleterious consequences of chronic morphine exposure in male mice[J]. Neuropsychopharmacology, 2021, 46(7): 1373-85. doi:10.1038/s41386-020-00927-x |
| [19] | Qin X, Cardoso Rodriguez F, Sufi J, et al. An oncogenic phenoscape of colonic stem cell polarization[J]. Cell, 2023, 186(25): 5554-68.e18. doi:10.1016/j.cell.2023.11.004 |
| [20] | Chen CK, Ng CS, Wu SM, et al. Regulatory differences in natal down development between altricial Zebra finch and precocial chicken[J]. Mol Biol Evol, 2016, 33(8): 2030-43. doi:10.1093/molbev/msw085 |
| [21] | Taylor KR, Barron T, Hui A, et al. Glioma synapses recruit mechanisms of adaptive plasticity[J]. Nature, 2023, 623(7986): 366-74. doi:10.1038/s41586-023-06678-1 |
| [22] | Iijima N, Sato K, Kuranaga E, et al. Differential cell adhesion implemented by Drosophila Toll corrects local distortions of the anterior-posterior compartment boundary[J]. Nat Commun, 2020, 11(1): 6320. doi:10.1038/s41467-020-20118-y |
| [23] | Valle JW, Lamarca A, Goyal L, et al. New horizons for precision medicine in biliary tract cancers[J]. Cancer Discov, 2017, 7(9): 943-62. doi:10.1158/2159-8290.cd-17-0245 |
| [24] | Dai J, Su YZ, Zhong SY, et al. Exosomes: key players in cancer and potential therapeutic strategy[J]. Signal Transduct Target Ther, 2020, 5(1): 145. doi:10.1038/s41392-020-00261-0 |
| [25] | Dunn HA, Dhaliwal SK, Chang CT, et al. Distinct autoregulatory roles of ELFN1 intracellular and extracellular domains on membrane trafficking, synaptic localization, and dimerization[J]. J Biol Chem, 2025, 301(1): 108073. doi:10.1016/j.jbc.2024.108073 |
| [26] | Ayhan S, Dursun A. ELFN1 is a new extracellular matrix (ECM)-associated protein[J]. Life Sci, 2024, 352: 122900. doi:10.1016/j.lfs.2024.122900 |
| [27] | Wishart AL, Conner SJ, Guarin JR, et al. Decellularized extracellular matrix scaffolds identify full-length collagen VI as a driver of breast cancer cell invasion in obesity and metastasis[J]. Sci Adv, 2020, 6(43): eabc3175. doi:10.1126/sciadv.abc3175 |
| [28] | Dunn HA, Orlandi C, Martemyanov KA. Beyond the ligand: extracellular and transcellular G protein-coupled receptor complexes in physiology and pharmacology[J]. Pharmacol Rev, 2019, 71(4): 503-19. doi:10.1124/pr.119.018044 |
| [29] | Kumar US, Natarajan A, Massoud TF, et al. FN3 linked nanobubbles as a targeted contrast agent for US imaging of cancer-associated human PD-L1[J]. J Control Release, 2022, 346: 317-27. doi:10.1016/j.jconrel.2022.04.030 |
| [30] | Winkler J, Abisoye-Ogunniyan A, Metcalf KJ, et al. Concepts of extracellular matrix remodelling in tumour progression and metastasis[J]. Nat Commun, 2020, 11(1): 5120. doi:10.1038/s41467-020-18794-x |
| [31] | Glaviano A, Foo ASC, Lam HY, et al. PI3K/AKT/mTOR signaling transduction pathway and targeted therapies in cancer[J]. Mol Cancer, 2023, 22(1): 138. doi:10.1186/s12943-023-01827-6 |
| [32] | Deng ZQ, Fan T, Xiao C, et al. TGF-β signaling in health, disease and therapeutics[J]. Sig Transduct Target Ther, 2024, 9: 61. doi:10.1038/s41392-024-01764-w |
| [33] | Rigillo G, Belluti S, Campani V, et al. The NF-Y splicing signature controls hybrid EMT and ECM-related pathways to promote aggressiveness of colon cancer[J]. Cancer Lett, 2023, 567: 216262. doi:10.1016/j.canlet.2023.216262 |
| [34] | Strating E, Verhagen MP, Wensink E, et al. Co-cultures of colon cancer cells and cancer-associated fibroblasts recapitulate the aggressive features of mesenchymal-like colon cancer[J]. Front Immunol, 2023, 14: 1053920. doi:10.3389/fimmu.2023.1053920 |
| [1] | 陈鑫源, 吴成挺, 李瑞迪, 潘雪芹, 张耀丹, 陶俊宇, 林才志. 双术汤通过P53/SLC7A11/GPX4通路诱导胃癌细胞铁死亡[J]. 南方医科大学学报, 2025, 45(7): 1363-1371. |
| [2] | 庞金龙, 赵新丽, 张振, 王豪杰, 周星琦, 杨玉梅, 李姗姗, 常小强, 李锋, 李娴. 皮肤黑色素瘤中MMRN2高表达促进肿瘤细胞的侵袭和迁移并与不良预后相关[J]. 南方医科大学学报, 2025, 45(7): 1479-1489. |
| [3] | 高志, 吴傲, 胡仲翔, 孙培养. 类风湿性关节炎中氧化应激与免疫浸润的生物信息学分析[J]. 南方医科大学学报, 2025, 45(4): 862-870. |
| [4] | 许怀文, 翁丽, 薛鸿. CXCL12可作为2型糖尿病合并慢性阻塞性肺疾病的潜在治疗靶点[J]. 南方医科大学学报, 2025, 45(1): 100-109. |
| [5] | 叶梦楠, 武鸿美, 梅琰, 张庆玲. CREM在胃癌中高表达并与患者的不良预后相关[J]. 南方医科大学学报, 2024, 44(9): 1776-1782. |
| [6] | 纪凯, 于冠宇, 周乐其, 张天帅, 凌潜龙, 满文江, 朱冰, 张卫. HNRNPA1基因在结直肠癌组织中高表达及其潜在的诊断和治疗价值[J]. 南方医科大学学报, 2024, 44(9): 1685-1695. |
| [7] | 陈莉莉, 吴天宇, 张铭, 丁子夏, 张妍, 杨依清, 郑佳倩, 张小楠. 类风湿关节炎的潜在生物标志物及其免疫调控机制:基于GEO数据库[J]. 南方医科大学学报, 2024, 44(6): 1098-1108. |
| [8] | 周伟, 聂军, 胡佳, 蒋艺枝, 张大发. 内质网应激相关基因在主动脉夹层疾病中的差异性表达及与免疫浸润的相关性[J]. 南方医科大学学报, 2024, 44(5): 859-866. |
| [9] | 裴蓓, 张艺, 魏思源, 梅语, 宋标, 董港, 温子昂, 李学军. 基于转录组学测序及生物信息学方法鉴定肠上皮化生的潜在致病基因[J]. 南方医科大学学报, 2024, 44(5): 941-949. |
| [10] | 王沁智, 宋冰, 郝诗睿, 肖志远, 金连辉, 郑通, 柴芳. 基于生物信息学分析CCND2在甲状腺乳头状癌中的表达及其对免疫浸润的影响[J]. 南方医科大学学报, 2024, 44(5): 981-988. |
| [11] | 梁一豪, 赖颖君, 袁燕文, 袁 炜, 张锡波, 张拔山, 卢志锋. 基于GEO数据库筛选胃癌差异表达基因及其功能和通路富集分析[J]. 南方医科大学学报, 2024, 44(3): 605-616. |
| [12] | 任自敬, 周佩洋, 田静. 基于ceRNA芯片研究帕金森病患者血浆长链非编码RNA表达谱并验证lnc-CTSD-5∶1在PD细胞模型中的作用[J]. 南方医科大学学报, 2024, 44(11): 2146-2155. |
| [13] | 成佳聪, 李智慧, 刘 鳐, 李 成, 黄 鑫, 田颖鑫, 沈富兵. 早幼粒细胞白血病蛋白与TAK1结合蛋白相互作用的生物信息学分析与验证[J]. 南方医科大学学报, 2024, 44(1): 179-186. |
| [14] | 孙 硕, 黄 鑫, 李国东, 张春云, 卢泽梅, 张伟伟, 李泽彦, 杨清竹. 敲低结肠癌转移相关基因1促进RSL3诱导的结直肠癌细胞铁死亡[J]. 南方医科大学学报, 2024, 44(1): 173-178. |
| [15] | 王秋生, 张 震, 王 炼, 汪 煜, 姚新宇, 王月月, 张小凤, 葛思堂, 左芦根. 高表达DAP5促进胃癌细胞的糖代谢并与不良预后相关[J]. 南方医科大学学报, 2023, 43(7): 1063-1070. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||