南方医科大学学报 ›› 2024, Vol. 44 ›› Issue (9): 1776-1782.doi: 10.12122/j.issn.1673-4254.2024.09.18
收稿日期:2024-02-24
出版日期:2024-09-20
发布日期:2024-09-30
通讯作者:
张庆玲
E-mail:yemengnan3@163.com;zhangqingling@gdph.org.cn
作者简介:叶梦楠,硕士,E-mail: yemengnan3@163.com
基金资助:
Mengnan YE1,2( ), Hongmei WU2, Yan MEI2, Qingling ZHANG2(
), Hongmei WU2, Yan MEI2, Qingling ZHANG2( )
)
Received:2024-02-24
Online:2024-09-20
Published:2024-09-30
Contact:
Qingling ZHANG
E-mail:yemengnan3@163.com;zhangqingling@gdph.org.cn
Supported by:摘要:
目的 分析CREM在胃癌中的表达及其与临床预后的相关性。 方法 利用TCGA和GEO数据库分析胃癌和癌旁组织CREM mRNA表达差异,免疫组化染色(IHC)分析43例胃癌和配对癌旁组织的CREM蛋白表达差异,分析CREM表达与胃癌患者临床病理特征之间的关系;采用Kaplan-Meier生存分析探讨CREM表达水平与胃癌患者生存期之间的关系;利用LinkedOmics数据库注释CREM相关基因的GO功能和KEGG通路的富集情况。 结果 数据库分析显示,CREM在胃癌组织中高表达(P<0.05),与胃癌患者的不良预后正相关(P=0.01)。IHC结果显示,与癌旁组织相比,胃癌组织中CREM高表达(P<0.0001),CREM表达水平与T分期和N分期有关(P<0.05),CREM高表达的胃癌患者总生存期更短(RR=4.02,P=0.0046)。基因富集分析显示,CREM可能通过细胞黏附分子介导的信号通路促进胃癌发生发展。 结论 CREM在胃癌中高表达提示不良预后,CREM可作为候选的胃癌预后指标。
叶梦楠, 武鸿美, 梅琰, 张庆玲. CREM在胃癌中高表达并与患者的不良预后相关[J]. 南方医科大学学报, 2024, 44(9): 1776-1782.
Mengnan YE, Hongmei WU, Yan MEI, Qingling ZHANG. High expression of CREM is associated with poor prognosis in gastric cancer patients[J]. Journal of Southern Medical University, 2024, 44(9): 1776-1782.
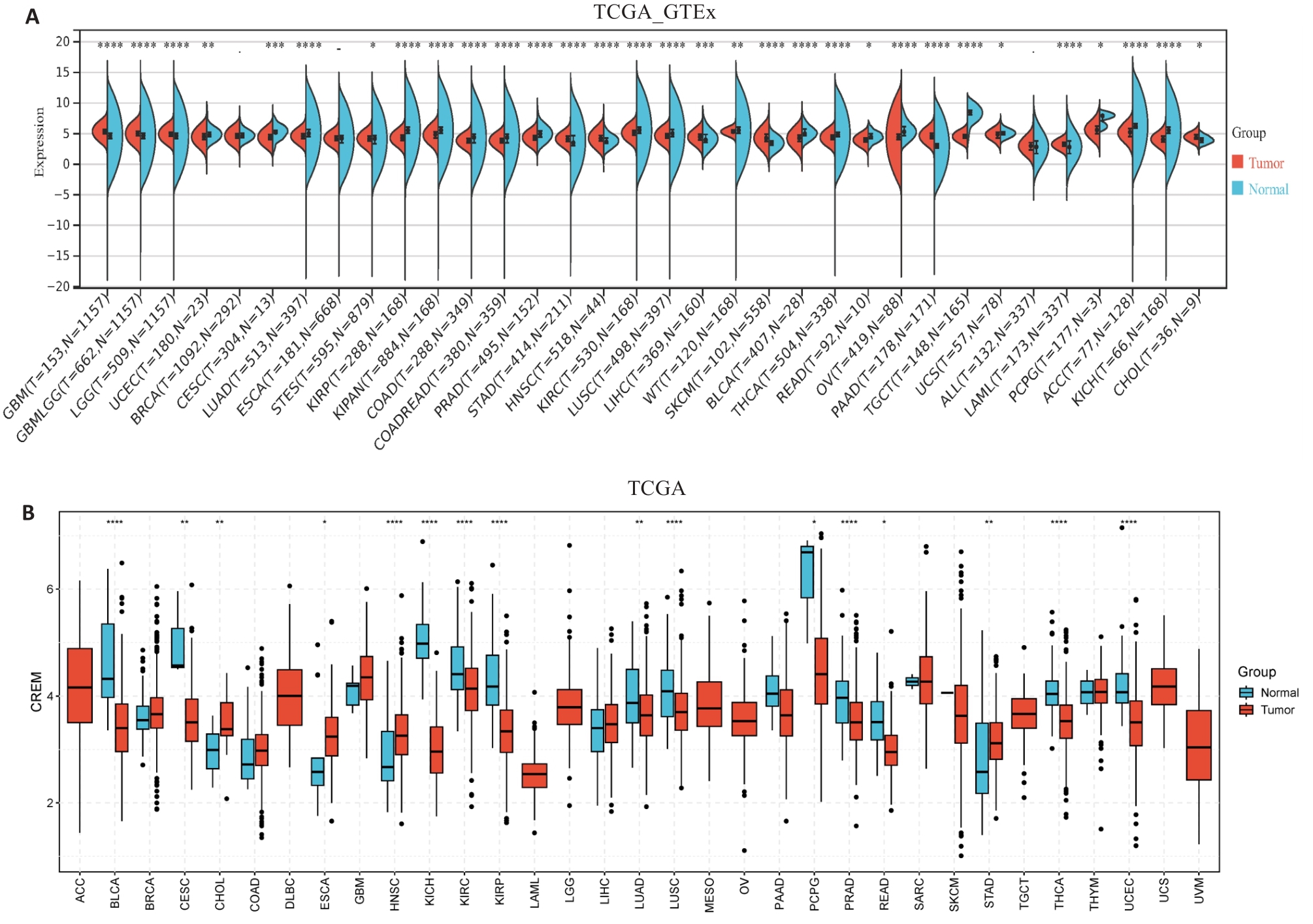
图1 CREM在多种恶性肿瘤中的差异性表达
Fig.1 Differential expression of CREM in different cancers based on data from TCGA-GTEx (A) and TCGA (B) databases. *P<0.05, **P<0.01, ***P<0.001, ****P<0.0001.
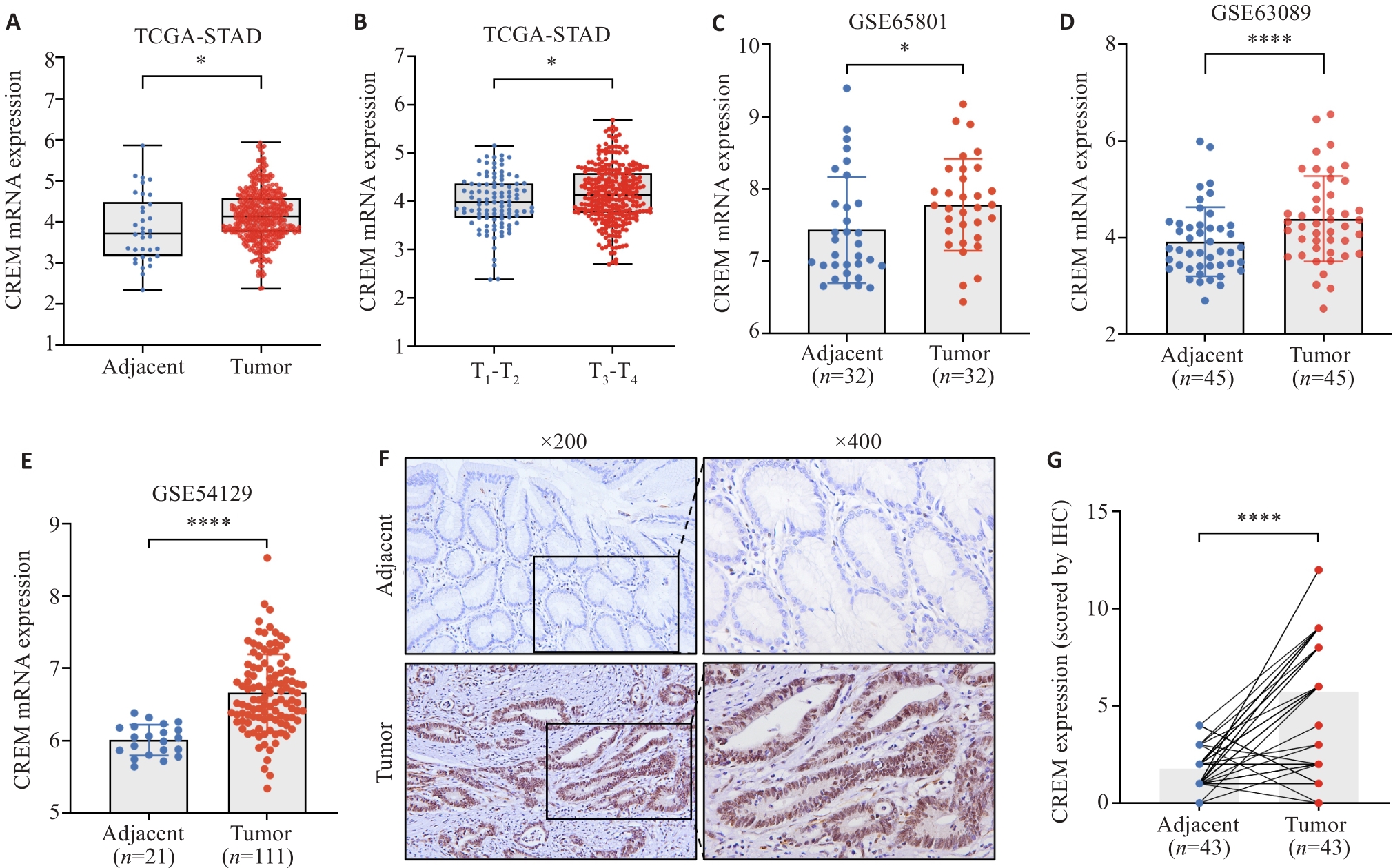
图3 CREM在胃癌组织中高表达
Fig.3 CREM is highly expressed in gastric cancer tissues. A: Expression of CREM in gastric cancer and adjacent tissues in TCGA database. B: Differential expression of CREM in tumors in different T-stages. C-E: Differential expression of CREM in gastric cancer tissues and adjacent tissues in GSE65801, GSE63089, and GSE54129 datasets. F, G: Immunohistochemical staining of CREM in gastric cancer and adjacent tissues and the immunohistochemical scores. *P<0.05, ****P<0.0001.
| Characteristic | n | CREM Expression | χ2 | P | |
|---|---|---|---|---|---|
| Low (n=11) | High (n=32) | ||||
| Age (year) | 1.815 | 0.178 | |||
| <60 | 15 | 2 (13.3) | 13 (86.7) | ||
| ≥60 | 28 | 9 (32.1) | 19 (67.9) | ||
| Gender | 1.120 | 0.290 | |||
| Male | 29 | 6 (20.7) | 23 (79.3) | ||
| Female | 14 | 5 (35.7) | 9 (64.3) | ||
| Ki-67(%) | 1.120 | 0.290 | |||
| <50 | 14 | 5 (35.7) | 9 (64.3) | ||
| ≥50 | 29 | 6 (20.7) | 23 (79.3) | ||
| Tumor size (cm) | 2.516 | 0.113 | |||
| <4 | 15 | 6 (40.0) | 9 (60.0) | ||
| ≥4 | 28 | 5 (17.9) | 23 (82.1) | ||
| T stage | 10.862 | 0.001 | |||
| T1-T2 | 14 | 8 (57.1) | 6 (42.9) | ||
| T3-T4 | 29 | 3 (10.3) | 26 (89.7) | ||
| N stage | 4.418 | 0.036 | |||
| N0 | 16 | 7 (43.8) | 9 (56.2) | ||
| N1-N3 | 27 | 4 (14.8) | 23 (85.2) | ||
| TNM staging | 3.376 | 0.066 | |||
| I-II | 21 | 8 (38.1) | 13 (61.9) | ||
| III-IV | 22 | 3 (13.6) | 19 (86.4) | ||
表1 43例胃癌患者CREM的表达与临床病理特征的关系
Tab. 1 Relationship between CREM expression and clinicopathological features of 43 patients with STAD
| Characteristic | n | CREM Expression | χ2 | P | |
|---|---|---|---|---|---|
| Low (n=11) | High (n=32) | ||||
| Age (year) | 1.815 | 0.178 | |||
| <60 | 15 | 2 (13.3) | 13 (86.7) | ||
| ≥60 | 28 | 9 (32.1) | 19 (67.9) | ||
| Gender | 1.120 | 0.290 | |||
| Male | 29 | 6 (20.7) | 23 (79.3) | ||
| Female | 14 | 5 (35.7) | 9 (64.3) | ||
| Ki-67(%) | 1.120 | 0.290 | |||
| <50 | 14 | 5 (35.7) | 9 (64.3) | ||
| ≥50 | 29 | 6 (20.7) | 23 (79.3) | ||
| Tumor size (cm) | 2.516 | 0.113 | |||
| <4 | 15 | 6 (40.0) | 9 (60.0) | ||
| ≥4 | 28 | 5 (17.9) | 23 (82.1) | ||
| T stage | 10.862 | 0.001 | |||
| T1-T2 | 14 | 8 (57.1) | 6 (42.9) | ||
| T3-T4 | 29 | 3 (10.3) | 26 (89.7) | ||
| N stage | 4.418 | 0.036 | |||
| N0 | 16 | 7 (43.8) | 9 (56.2) | ||
| N1-N3 | 27 | 4 (14.8) | 23 (85.2) | ||
| TNM staging | 3.376 | 0.066 | |||
| I-II | 21 | 8 (38.1) | 13 (61.9) | ||
| III-IV | 22 | 3 (13.6) | 19 (86.4) | ||
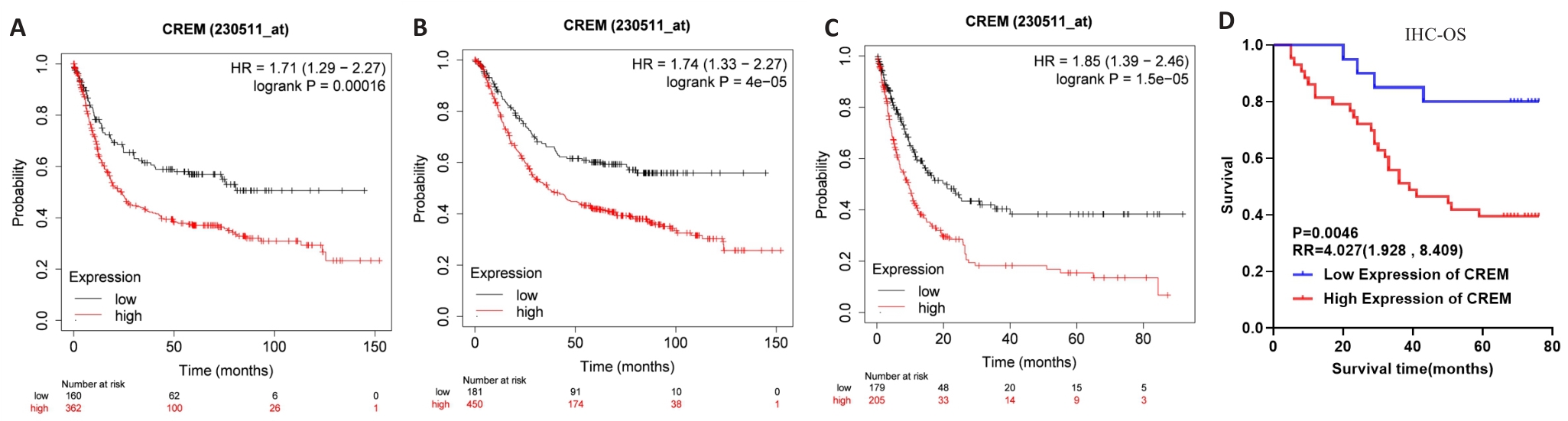
图4 CREM高表达与胃癌患者预后不良相关
Fig.4 High CREM expression is associated with poor prognosis in gastric cancer patients. A-C: FP, OS, and PPS curves of gastric cancer patients with different CREM mRNA expression levels in the Kaplan-Meier Plotter database. D: OS curves of patients with high and low CREM protein expression in clinical gastric cancer samples.
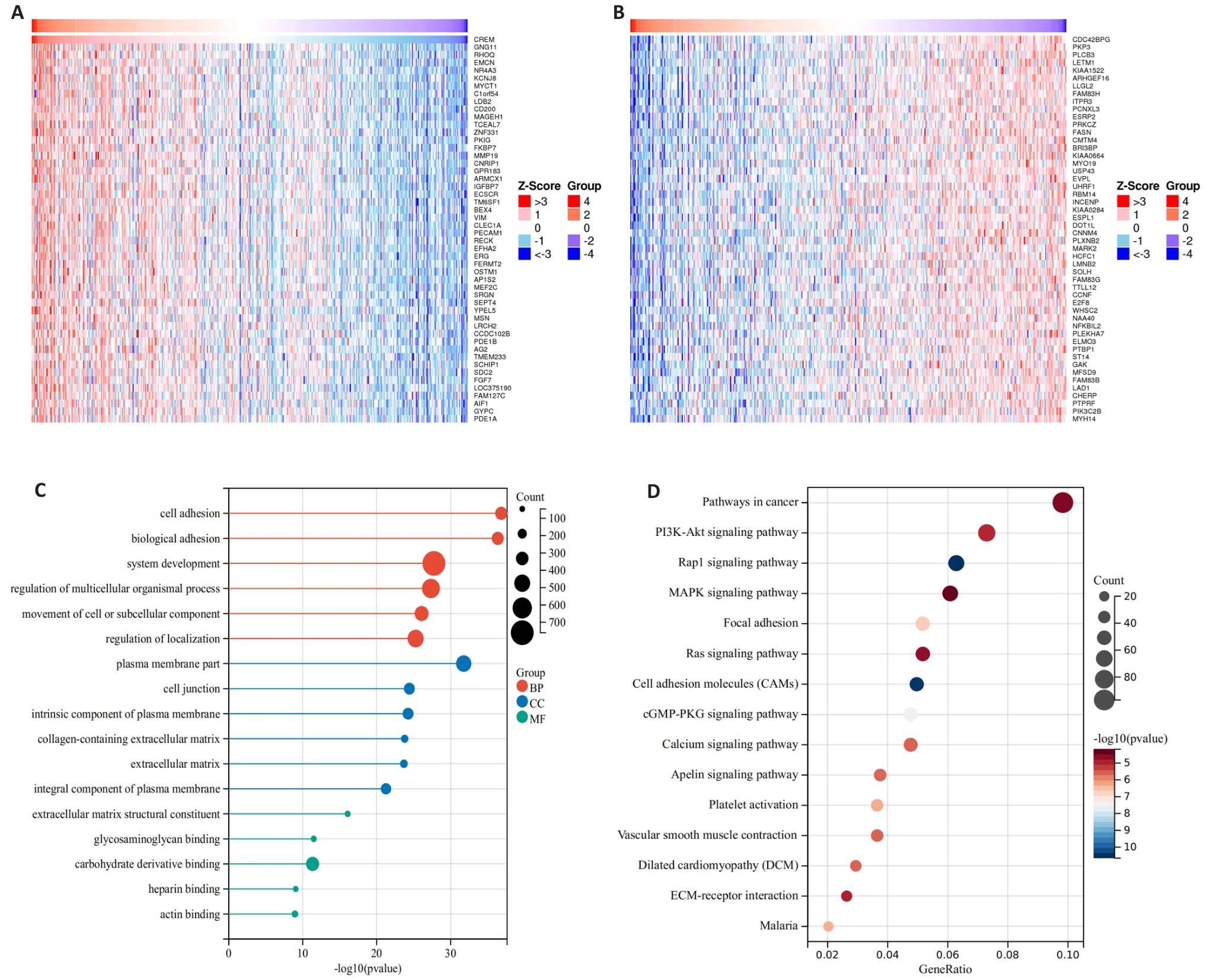
图5 胃癌中CREM相关基因分析及KEGG、GO富集分析结果
Fig.5 Analysis of CREM-related genes and results of KEGG and GO enrichment analysis in gastric cancer. A, B: LinkedOmics database analysis of genes positively and negatively correlated with CREM expression in the TCGA gastric cancer dataset. C: GO enrichment analysis to explore the role of CREM in gastric cancer, including the biological process, cellular composition, and molecular function. D: KEGG enrichment analysis to explore the relevant molecular pathways of CREM in gastric cancer development.
| 1 | Sung H, Ferlay J, Siegel RL, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries[J]. CA Cancer J Clin, 2021, 71(3): 209-49. |
| 2 | Qiu HB, Cao SM, Xu RH. Cancer incidence, mortality, and burden in China: a time-trend analysis and comparison with the United States and United Kingdom based on the global epidemiological data released in 2020[J]. Cancer Commun, 2021, 41(10): 1037-48. |
| 3 | 赫 捷, 陈万青, 李兆申, 等. 中国胃癌筛查与早诊早治指南(2022,北京)[J]. 中华肿瘤杂志, 2022, 44(7): 634-66. |
| 4 | Cao W, Chen HD, Yu YW, et al. Changing profiles of cancer burden worldwide and in China: a secondary analysis of the global cancer statistics 2020[J]. Chin Med J, 2021, 134(7): 783-91. |
| 5 | Thrift AP, Wenker TN, El-Serag HB. Global burden of gastric cancer: epidemiological trends, risk factors, screening and prevention[J]. Nat Rev Clin Oncol, 2023, 20(5): 338-49. |
| 6 | Shitara K, Chin K, Yoshikawa T, et al. Phase II study of adjuvant chemotherapy of S-1 plus oxaliplatin for patients with stage III gastric cancer after D2 gastrectomy[J]. Gastric Cancer, 2017, 20(1): 175-81. |
| 7 | Cancer Genome Atlas Research Network. Comprehensive molecular characterization of gastric adenocarcinoma[J]. Nature, 2014, 513(7517): 202-9. |
| 8 | Canale M, Casadei-Gardini A, Ulivi P, et al. Epigenetic mechanisms in gastric cancer: potential new therapeutic opportunities[J]. Int J Mol Sci, 2020, 21(15): 5500-9. |
| 9 | Persengiev SP, Green MR. The role of ATF/CREB family members in cell growth, survival and apoptosis[J]. Apoptosis, 2003, 8(3): 225-8. |
| 10 | Brindle PK, Montminy MR. The CREB family of transcription activators[J]. Curr Opin Genet Dev, 1992, 2(2): 199-204. |
| 11 | Sassone-Corsi P. Coupling gene expression to cAMP signalling: role of CREB and CREM[J]. Int J Biochem Cell Biol, 1998, 30(1): 27-38. |
| 12 | Sánchez-Jasso DE, López-Guzmán SF, Bermúdez-Cruz RM, et al. Novel aspects of cAMP-response element modulator (CREM) role in spermatogenesis and male fertility[J]. Int J Mol Sci, 2023, 24(16): 12558-67. |
| 13 | Wu XM, Jin W, Liu XJ, et al. Cyclic AMP response element modulator-1 (CREM-1) involves in neuronal apoptosis after traumatic brain injury[J]. J Mol Neurosci, 2012, 47(2): 357-67. |
| 14 | Kaprio H, Siddiqui A, Saustila L, et al. The oncogenic properties of the EWSR1: CREM fusion gene are associated with polyamine metabolism[J]. Sci Rep, 2023, 13(1): 4884-93. |
| 15 | Wang YC, Zhou SH, Yang XJ, et al. Low expression of cyclic AMP response element modulator-1 can increase the migration and invasion of esophageal squamous cell carcinoma[J]. Tumour Biol, 2013, 34(6): 3649-57. |
| 16 | Kaprio H, Heuser VD, Orte K, et al. Expression of transcription factor CREM in human tissues[J]. J Histochem Cytochem, 2021, 69(8): 495-509. |
| 17 | Shen WT, Song ZG, Zhong X, et al. Sangerbox: a comprehensive, interaction-friendly clinical bioinformatics analysis platform[J]. Imeta, 2022, 1(3): e36. |
| 18 | Hanahan D. Hallmarks of cancer: new dimensions[J]. Cancer Discov, 2022, 12(1): 31-46. |
| 19 | Alsina M, Arrazubi V, Diez M, et al. Current developments in gastric cancer: from molecular profiling to treatment strategy[J]. Nat Rev Gastroenterol Hepatol, 2023, 20(3): 155-70. |
| 20 | Sun JM, Shen L, Shah MA, et al. Pembrolizumab plus chemotherapy versus chemotherapy alone for first-line treatment of advanced oesophageal cancer (KEYNOTE-590): a randomised, placebo-controlled, phase 3 study[J]. Lancet, 2021, 398(10302): 759-71. |
| 21 | Augustin JE, Soussan P, Bass AJ. Targeting the complexity of ERBB2 biology in gastroesophageal carcinoma[J]. Ann Oncol, 2022, 33(11): 1134-48. |
| 22 | Kawazoe A, Fukuoka S, Nakamura Y, et al. Lenvatinib plus pembrolizumab in patients with advanced gastric cancer in the first-line or second-line setting (EPOC1706): an open-label, single-arm, phase 2 trial[J]. Lancet Oncol, 2020, 21(8): 1057-65. |
| 23 | Wainberg ZA, Enzinger PC, Kang YK, et al. Bemarituzumab in patients with FGFR2b-selected gastric or gastro-oesophageal junction adenocarcinoma (FIGHT): a randomised, double-blind, placebo-controlled, phase 2 study[J]. Lancet Oncol, 2022, 23(11): 1430-40. |
| 24 | Qi CS, Gong JF, Li J, et al. Claudin18.2-specific CAR T cells in gastrointestinal cancers: phase 1 trial interim results[J]. Nat Med, 2022, 28(6): 1189-98. |
| 25 | Bardia A, Messersmith WA, Kio EA, et al. Sacituzumab govitecan, a Trop-2-directed antibody-drug conjugate, for patients with epithelial cancer: final safety and efficacy results from the phase I/II IMMU-132-01 basket trial[J]. Ann Oncol, 2021, 32(6): 746-56. |
| 26 | Servillo G, Della Fazia MA, Sassone-Corsi P. Coupling cAMP signaling to transcription in the liver: pivotal role of CREB and CREM[J]. Exp Cell Res, 2002, 275(2): 143-54. |
| 27 | Behr R, Weinbauer GF. cAMP response element modulator (CREM): an essential factor for spermatogenesis in Primates?[J]. Int J Androl, 2001, 24(3): 126-35. |
| 28 | Feng F, Wu JT, Gao ZL, et al. Screening the key microRNAs and transcription factors in prostate cancer based on microRNA functional synergistic relationships[J]. Medicine, 2017, 96(1): e5679. |
| 29 | Janiszewska M, Primi MC, Izard T. Cell adhesion in cancer: beyond the migration of single cells[J]. J Biol Chem, 2020, 295(8): 2495-505. |
| 30 | Chen JZ, Qin HY, Hao JZ, et al. Cardiac-specific overexpression of CREM-IbΔC-X via CRISPR/Cas9 in mice presents a new model of atrial cardiomyopathy with spontaneous atrial fibrillation[J]. Transl Res, 2024, 267: 54-66. |
| [1] | 陈鑫源, 吴成挺, 李瑞迪, 潘雪芹, 张耀丹, 陶俊宇, 林才志. 双术汤通过P53/SLC7A11/GPX4通路诱导胃癌细胞铁死亡[J]. 南方医科大学学报, 2025, 45(7): 1363-1371. |
| [2] | 庞金龙, 赵新丽, 张振, 王豪杰, 周星琦, 杨玉梅, 李姗姗, 常小强, 李锋, 李娴. 皮肤黑色素瘤中MMRN2高表达促进肿瘤细胞的侵袭和迁移并与不良预后相关[J]. 南方医科大学学报, 2025, 45(7): 1479-1489. |
| [3] | 吴璇, 方家敏, 韩玮玮, 陈琳, 孙菁, 金齐力. 高表达PRELID1促进胃癌细胞上皮间质转化并与不良预后相关[J]. 南方医科大学学报, 2025, 45(7): 1535-1542. |
| [4] | 王康, 李海宾, 余靖, 孟源, 张虹丽. ELFN1高表达是结肠癌的预后生物标志物并促进结肠癌细胞的增殖转移[J]. 南方医科大学学报, 2025, 45(7): 1543-1553. |
| [5] | 侯鑫睿, 张振东, 曹明远, 杜予心, 王小平. 红景天苷靶向miR-1343-3p-OGDHL/PDHB糖代谢轴抑制胃癌细胞的体内外增殖[J]. 南方医科大学学报, 2025, 45(6): 1226-1239. |
| [6] | 张毅, 沈昱, 万志强, 陶嵩, 柳亚魁, 王栓虎. CDKN3高表达促进胃癌细胞的迁移和侵袭:基于调控p53/NF-κB信号通路和抑制胃癌细胞凋亡[J]. 南方医科大学学报, 2025, 45(4): 853-861. |
| [7] | 高志, 吴傲, 胡仲翔, 孙培养. 类风湿性关节炎中氧化应激与免疫浸润的生物信息学分析[J]. 南方医科大学学报, 2025, 45(4): 862-870. |
| [8] | 黄晴晴, 张文静, 张小凤, 王炼, 宋雪, 耿志军, 左芦根, 王月月, 李静, 胡建国. 高表达MYO1B促进胃癌细胞增殖、迁移和侵袭并与患者的不良预后有关[J]. 南方医科大学学报, 2025, 45(3): 622-631. |
| [9] | 李华莉, 宋婷, 刘嘉雯, 李永宝, 姜兆静, 窦文, 周凌宏. 预后导向的肺癌调强放疗计划优化新方法[J]. 南方医科大学学报, 2025, 45(3): 643-649. |
| [10] | 宋雪, 陈悦, 张敏, 张诺, 左芦根, 李静, 耿志军, 张小凤, 王月月, 王炼, 胡建国. GPSM2在胃癌组织中高表达并通过促进肿瘤细胞的增殖影响患者预后[J]. 南方医科大学学报, 2025, 45(2): 229-238. |
| [11] | 唐天威, 李路安, 陈源汉, 张丽, 徐丽霞, 李志莲, 冯仲林, 张辉林, 华瑞芳, 叶智明, 梁馨苓, 李锐钊. 高血清胱抑素C水平是IgA肾病不良预后的独立危险因素[J]. 南方医科大学学报, 2025, 45(2): 379-386. |
| [12] | 许怀文, 翁丽, 薛鸿. CXCL12可作为2型糖尿病合并慢性阻塞性肺疾病的潜在治疗靶点[J]. 南方医科大学学报, 2025, 45(1): 100-109. |
| [13] | 陈晓睿, 魏青政, 张宗亮, 原江水, 宋卫青. 过表达带电多泡体蛋白2B基因抑制肾透明细胞癌细胞的增殖[J]. 南方医科大学学报, 2025, 45(1): 126-136. |
| [14] | 王耀彬, 陈柳燕, 罗伊凌, 申继清, 周素芳. NUF2对泛癌的预后和免疫治疗效果的预测价值[J]. 南方医科大学学报, 2025, 45(1): 137-149. |
| [15] | 周超, 张晶晶, 唐巧, 付双楠, 张宁, 何召云, 张瑾, 张田义, 刘鹏程, 宫嫚. 血清色氨酸用于乙肝相关慢加急性肝衰竭90 d死亡风险分层管理的潜在价值:一项多中心回顾性研究[J]. 南方医科大学学报, 2025, 45(1): 59-64. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||