Journal of Southern Medical University ›› 2026, Vol. 46 ›› Issue (3): 693-706.doi: 10.12122/j.issn.1673-4254.2026.03.23
Kelin XIANG1( ), Xiaoyu ZHANG2, Zhengpeng LI1, Zhiwei XU2, Sujie LIU1, Yuan CHAI2,3(
), Xiaoyu ZHANG2, Zhengpeng LI1, Zhiwei XU2, Sujie LIU1, Yuan CHAI2,3( )
)
Received:2025-08-14
Online:2026-03-20
Published:2026-03-26
Contact:
Yuan CHAI
E-mail:xiangkelin1024@163.com;chaizxy@163.com
Kelin XIANG, Xiaoyu ZHANG, Zhengpeng LI, Zhiwei XU, Sujie LIU, Yuan CHAI. Identification of efferocytosis-related genes in osteoarthritis and prediction of traditional Chinese medicines based on bioinformatics and machine learning[J]. Journal of Southern Medical University, 2026, 46(3): 693-706.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.j-smu.com/EN/10.12122/j.issn.1673-4254.2026.03.23
| GEO ID | Platform | Control (n) | Case (n) | Group |
|---|---|---|---|---|
| GSE55235 | GPL96 | 10 | 10 | Training set |
| GSE55457 | GPL96 | 10 | 10 | Validation set |
| GSE117999 | GPL20844 | 12 | 12 | Validation set |
Tab.1 Basic information and grouping of GEO datasets
| GEO ID | Platform | Control (n) | Case (n) | Group |
|---|---|---|---|---|
| GSE55235 | GPL96 | 10 | 10 | Training set |
| GSE55457 | GPL96 | 10 | 10 | Validation set |
| GSE117999 | GPL20844 | 12 | 12 | Validation set |
| Gene | Primer sequence (5'-3') |
|---|---|
| UCP2-S | СТСССААТGТТGСССGАААТ |
| UCP2-A | GAAGTGGCAAGGGAGGTCGT |
| EGLN3-S | CGCCAAGTTACATGGAGGGG |
| EGLN3-A | CCAGACAGTCATGGCGTACC |
| IL1B-S | GAACAACAAAAATGCCTCGTGC |
| IL1B-A | GACAAACCGCTTTTCCATCTTCT |
| MMP13-S | TGCATACGAGCATCCATCCC |
| MMP13-A | CGTGTCCTCAAAGTGAACCGC |
| COL2A1-S | CAGACAGTACCTTGAGACAGCATGA |
| COL2A1-A | AGGTGCGAGCGGGATTCTT |
| GAPDH-S | CTGGAGAAACCTGCCAAGTATG |
| GAPDH-A | GGTGGAAGAATGGGAGTTGCT |
Tab.2 PCR primer sequences
| Gene | Primer sequence (5'-3') |
|---|---|
| UCP2-S | СТСССААТGТТGСССGАААТ |
| UCP2-A | GAAGTGGCAAGGGAGGTCGT |
| EGLN3-S | CGCCAAGTTACATGGAGGGG |
| EGLN3-A | CCAGACAGTCATGGCGTACC |
| IL1B-S | GAACAACAAAAATGCCTCGTGC |
| IL1B-A | GACAAACCGCTTTTCCATCTTCT |
| MMP13-S | TGCATACGAGCATCCATCCC |
| MMP13-A | CGTGTCCTCAAAGTGAACCGC |
| COL2A1-S | CAGACAGTACCTTGAGACAGCATGA |
| COL2A1-A | AGGTGCGAGCGGGATTCTT |
| GAPDH-S | CTGGAGAAACCTGCCAAGTATG |
| GAPDH-A | GGTGGAAGAATGGGAGTTGCT |
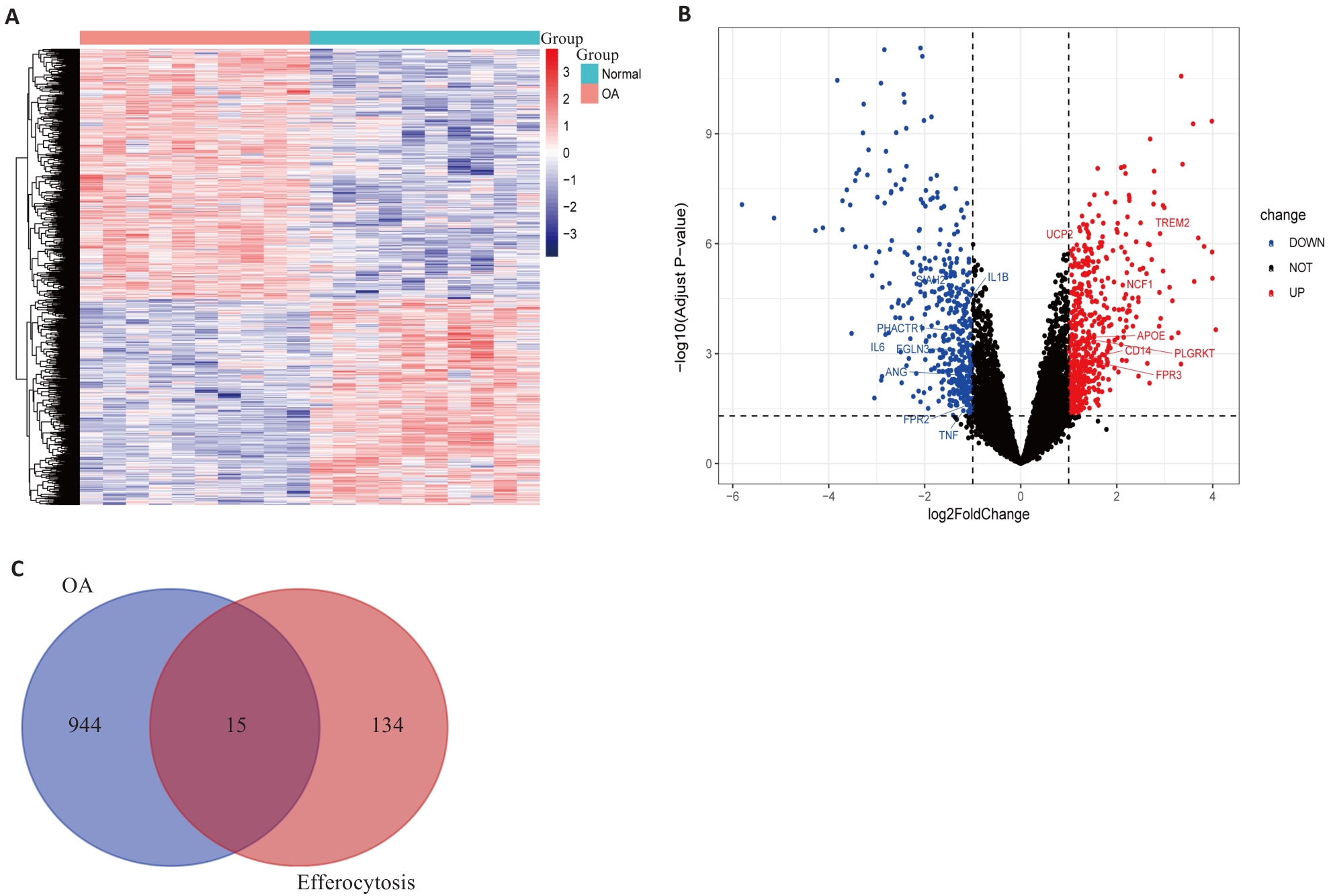
Fig.1 Differential gene expression analysis between osteoarthritis (OA) patients and healthy controls. A: Gene expression heatmap between OA patients and healthy controls. Red represents up-regulated expression, and blue represents down-regulated expression. B: Volcano plot of differentially expressed genes (DEGs). Black dots indicate genes with no significant differences, red dots indicate significantly up-regulated DEGs, and blue dots indicate significantly down-regulated DEGs. The figure displays efferocytosis-related DEGs. C: Venn diagram of the intersection between DEGs in OA and efferocytosis-related gene sets, obtaining 15 efferocytosis-related DEGs.
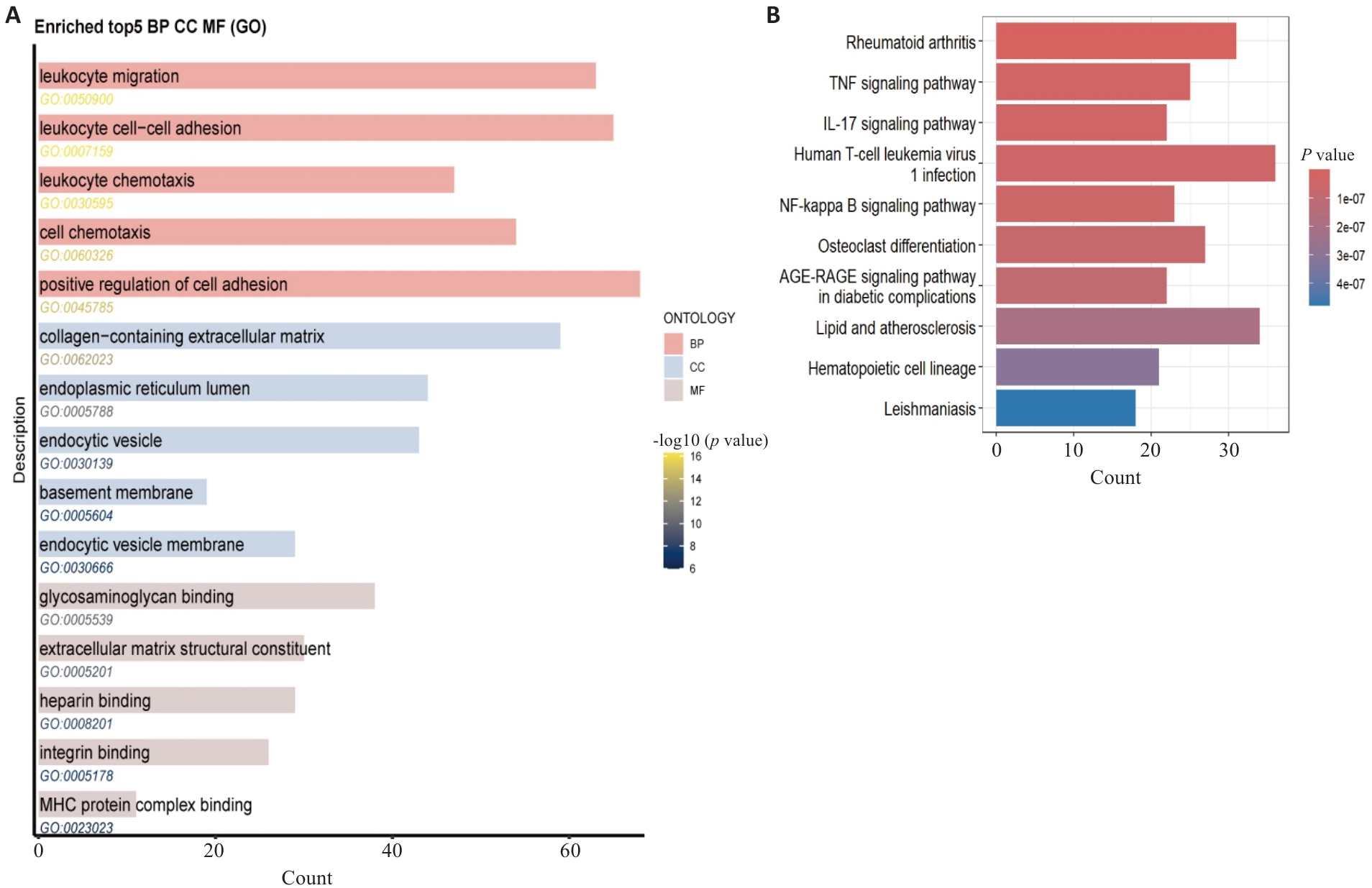
Fig.2 GO functional and KEGG pathway enrichment analysis of osteoarthritis DEGs. A: GO enrichment analysis results. B: KEGG pathway enrichment analysis results, displaying the signaling pathways significantly enriched by the DEGs.
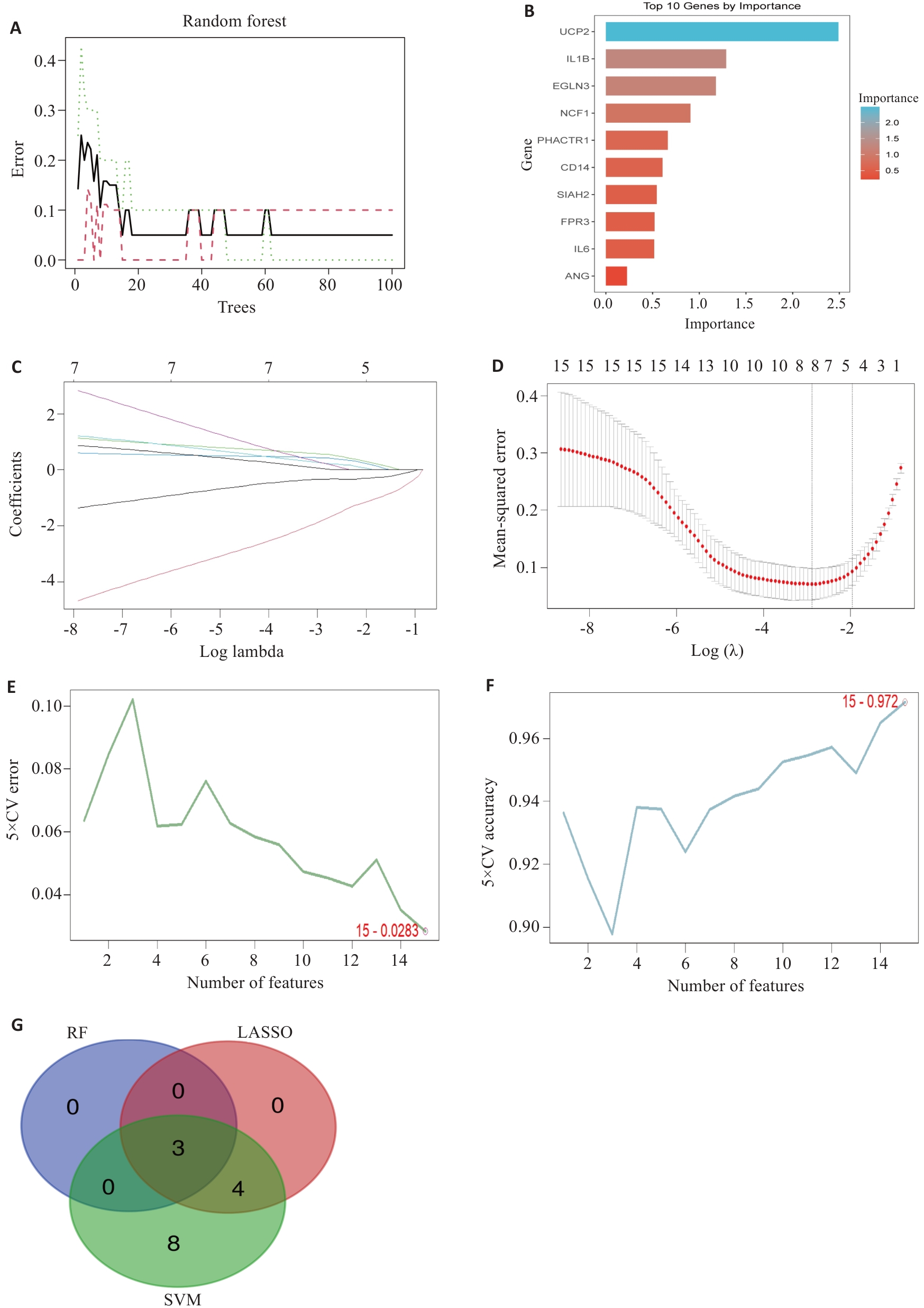
Fig.3 Machine learning screening of feature genes. A, B: RF screening process and results, showing trend of model error with increasing number of decision trees (A) and gene importance ranking (B). C, D: LASSO regression screening process and results showing model coefficient path diagram (C) and trend of cross-validation error with penalty coefficient λ (D). E, F: SVM screening process and results showing model error rate with varying number of features (E) and model accuracy with varying number of features (F). G: Venn diagram of the intersection of screening results from three machine learning algorithms, ultimately obtaining 3 consensus feature genes.
| Dataset | Accuracy | Precision | Recall | F1-Score | AUC |
|---|---|---|---|---|---|
| GSE55235 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 |
| GSE55457 | 0.850 | 0.818 | 0.900 | 0.857 | 0.900 |
| GSE117999 | 0.850 | 0.818 | 0.900 | 0.857 | 0.910 |
Tab.3 Performance evaluation of classification model based on the feature genes
| Dataset | Accuracy | Precision | Recall | F1-Score | AUC |
|---|---|---|---|---|---|
| GSE55235 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 |
| GSE55457 | 0.850 | 0.818 | 0.900 | 0.857 | 0.900 |
| GSE117999 | 0.850 | 0.818 | 0.900 | 0.857 | 0.910 |
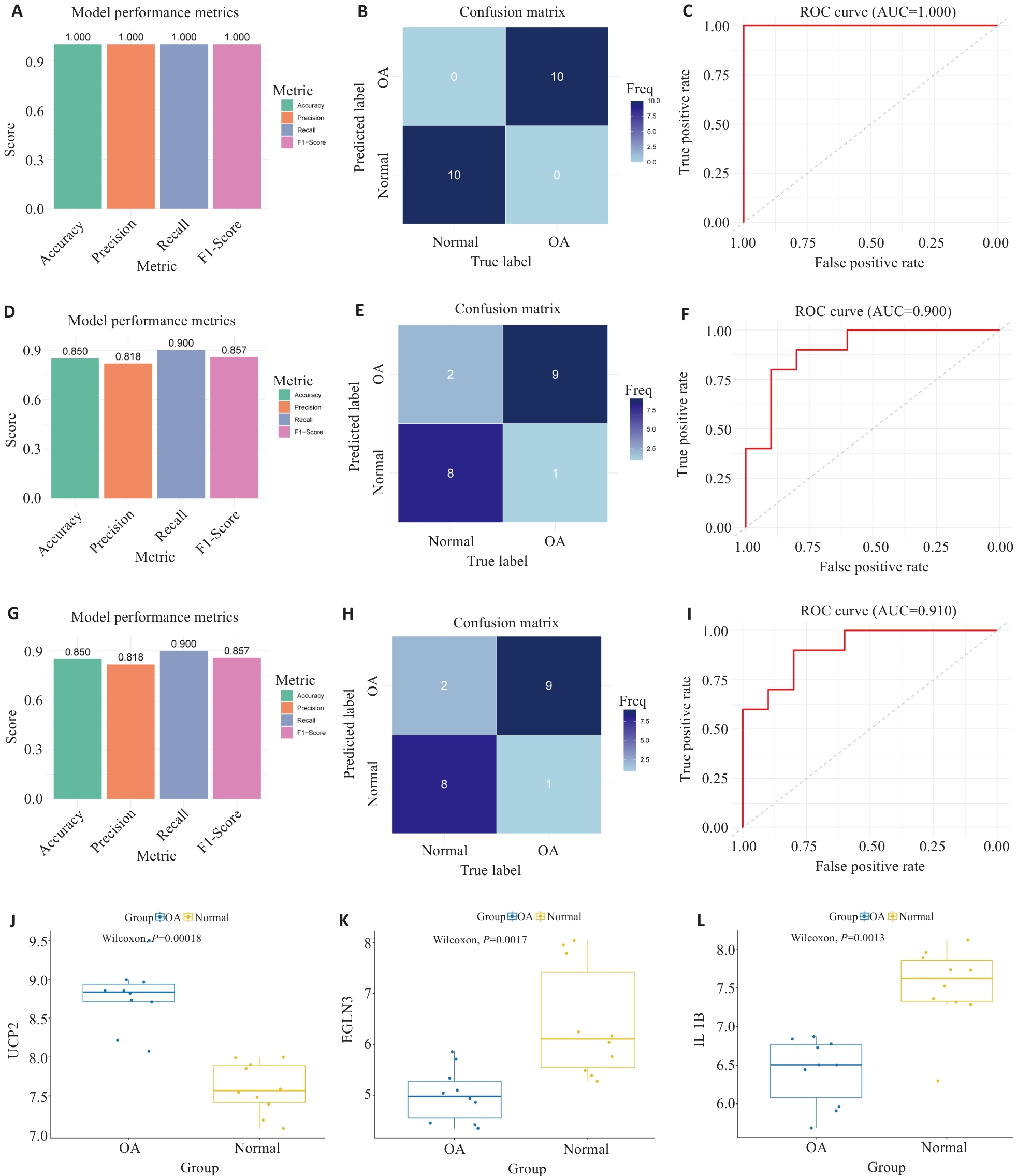
Fig.4 Comprehensive analysis of model performance evaluation. A-C: Performance assessment of the diagnostic model in the training set (GSE55235), showing the accuracy, precision, recall, and F1-score diagram (A), confusion matrix (B), and ROC curve and AUC value (C). D-F: Performance assessment of the diagnostic model in the validation set (GSE55457) showing the accuracy, precision, recall, and F1-score diagram (D), confusion matrix (E), and ROC curve and AUC value (F). G-I: Performance of the diagnostic model in the validation set (GSE117999), showing the accuracy, precision, recall, and F1-score diagram (G), confusion matrix (H), and ROC curve and AUC value (I). J-L: Comparison of expression levels of the 3 feature genes between OA patients and healthy controls (J: UCP2; K: EGLN3; L: IL1B).
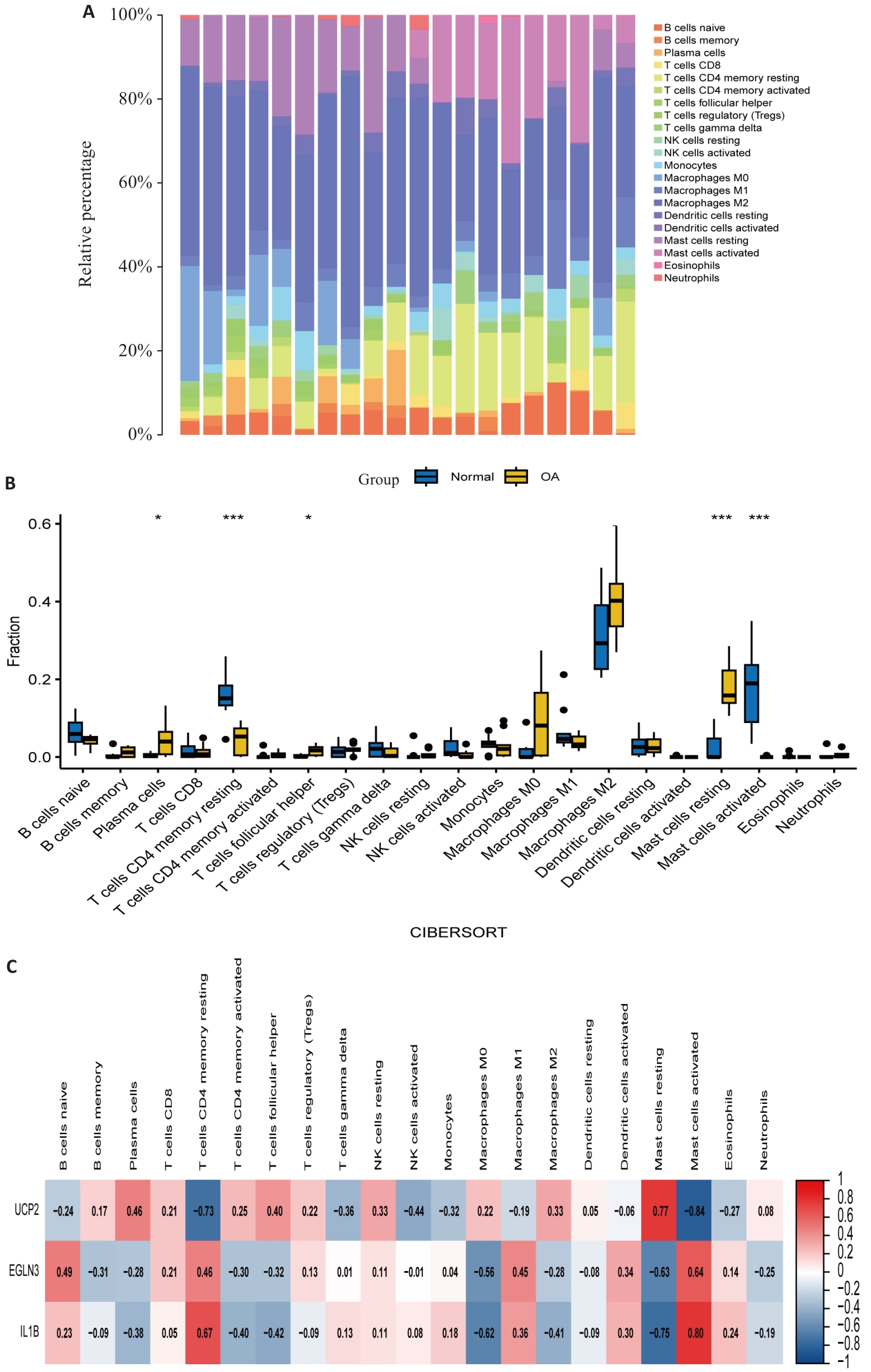
Fig.6 Immune infiltration analysis. A: Infiltration of immune cells in OA patient samples and healthy control samples, with different colors representing different immune cells. B: Comparison of immune cell infiltration between OA patient samples and healthy control samples. C: Correlation between feature genes and immune cells, with color depth representing the strength of correlation. *P<0.05, ***P<0.001.
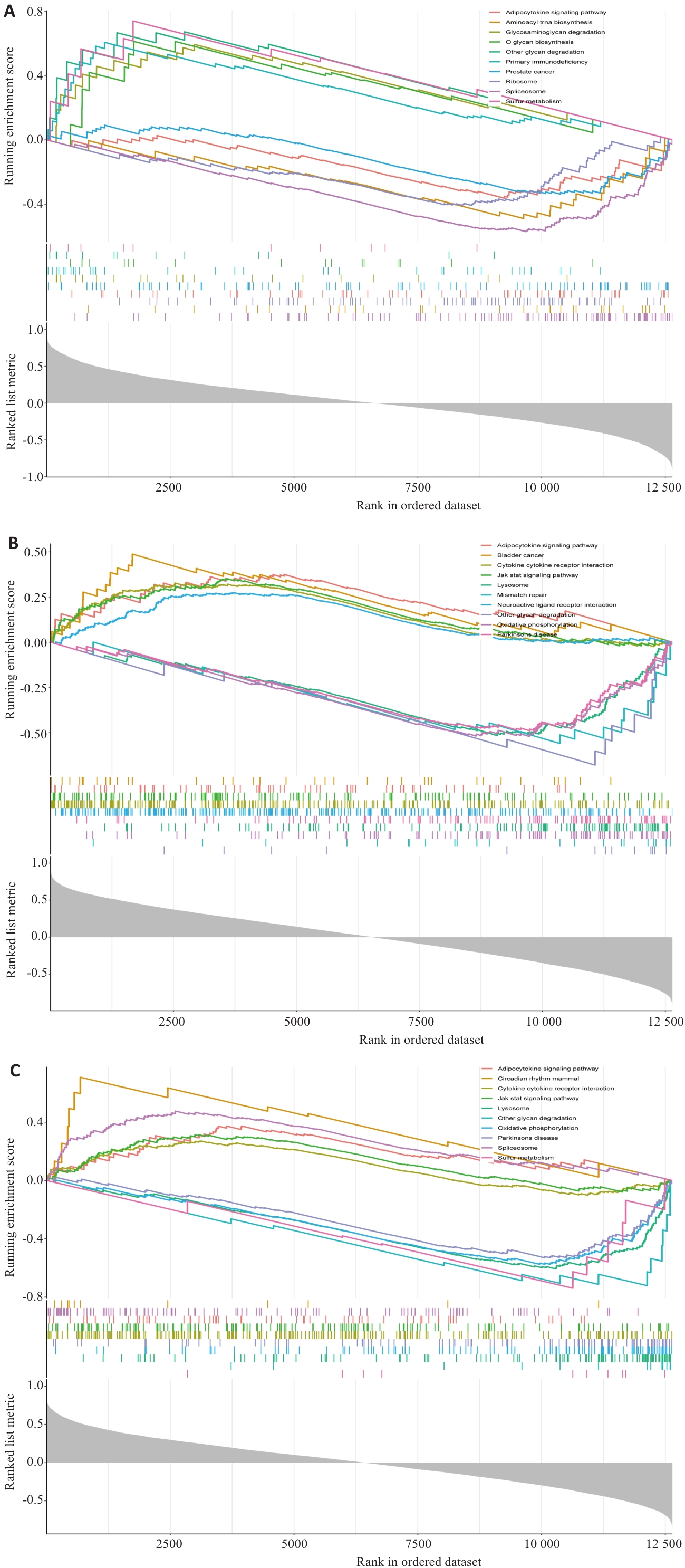
Fig.7 GSEA enrichment analysis results of the feature genes. A: GSEA enrichment analysis results of UCP2. B: GSEA enrichment analysis results of EGLN3. C: GSEA enrichment analysis results of IL1B.
| Category | TCM herbs |
|---|---|
| Exterior-releasing | Saposhnikovia divaricata, Cinnamomum cassia, Ligusticum sinense, Arctium lappa, Centipeda minima, Peucedanum praeruptorum, Angelica dahurica, Ephedra sinica |
| Heat-clearing | Rehmannia glutinosa, Sophora flavescens, Scutellaria baicalensis, Coptis chinensis, Forsythia suspensa, Portulaca oleracea, Lonicera japonica, Houttuynia cordata, Lonicera japonica stem, Arnebia euchroma, Anemarrhena asphodeloides, Morus alba leaf, Phellodendron chinense, Prunella vulgaris, Chrysanthemum indicum, Scutellaria barbata, Terminalia chebula |
| Purgative | Rheum palmatum, Euphorbia lathyris |
| Wind-damp dispelling | Angelica pubescens, Centella asiatica, Dioscorea nipponica, Sinomenium acutum, Clematis chinensis, Gentiana macrophylla, Morus alba twig, Siegesbeckia orientalis, Stephania tetrandra, Chaenomeles speciosa |
| Dampness-resolving | Pogostemon cablin |
| Dampness-draining | Sargassum, Poria cocos, Smilax glabra, Alisma orientale, Akebia quinata |
| Internal-warming | Fresh Ginger Rhizome, Dried Ginger Rhizome, Aconitum carmichaelii, Evodia rutaecarpa, Cnidium monnieri, Cinnamomum cassia |
| Qi-regulating | Magnolia officinalis flower, Magnolia officinalis bark, Inula racemosa, Aucklandia lappa, Citrus aurantium |
| Hemostatic | Artemisia argyi |
| Blood-activating | Ligusticum chuanxiong, Achyranthes bidentata, Salvia miltiorrhiza, Paeonia lactiflora, Carthamus tinctorius, Panax notoginseng, Curcuma longa, Panax notoginseng flower, Crocus sativus, Sargentodoxa cuneata, Caesalpinia sappan, Cyathula officinalis, Anemone raddeana |
| Phlegm-resolving | Ginkgo biloba |
| Tranquilizing | Polygala tenuifolia |
| Wind-extinguishing | Gastrodia elata |
| Tonic | Lycium barbarum, Epimedium brevicornum, Panax ginseng leaf, Panax ginseng, Atractylodes macrocephala, Fish maw, Angelica sinensis, Ganoderma lucidum, Rehmannia glutinosa, Red ginseng, Glycyrrhiza uralensis, Cordyceps sinensis, Eucommia ulmoides, Cornus officinalis, Polygonatum sibiricum, Ophiopogon japonicus, Gynostemma pentaphyllum, Coriolus versicolor, Dioscorea opposita, Houttuynia cordata, Hippophae rhamnoides, Bletilla striata, Agrimonia pilosa, Ligustrum lucidum |
| Astringent | Schisandra chinensis, Rosa laevigata, Punica granatum peel, Ephedra sinica root, Terminalia chebula |
Tab.4 Classification of predicted traditional Chinese medicines
| Category | TCM herbs |
|---|---|
| Exterior-releasing | Saposhnikovia divaricata, Cinnamomum cassia, Ligusticum sinense, Arctium lappa, Centipeda minima, Peucedanum praeruptorum, Angelica dahurica, Ephedra sinica |
| Heat-clearing | Rehmannia glutinosa, Sophora flavescens, Scutellaria baicalensis, Coptis chinensis, Forsythia suspensa, Portulaca oleracea, Lonicera japonica, Houttuynia cordata, Lonicera japonica stem, Arnebia euchroma, Anemarrhena asphodeloides, Morus alba leaf, Phellodendron chinense, Prunella vulgaris, Chrysanthemum indicum, Scutellaria barbata, Terminalia chebula |
| Purgative | Rheum palmatum, Euphorbia lathyris |
| Wind-damp dispelling | Angelica pubescens, Centella asiatica, Dioscorea nipponica, Sinomenium acutum, Clematis chinensis, Gentiana macrophylla, Morus alba twig, Siegesbeckia orientalis, Stephania tetrandra, Chaenomeles speciosa |
| Dampness-resolving | Pogostemon cablin |
| Dampness-draining | Sargassum, Poria cocos, Smilax glabra, Alisma orientale, Akebia quinata |
| Internal-warming | Fresh Ginger Rhizome, Dried Ginger Rhizome, Aconitum carmichaelii, Evodia rutaecarpa, Cnidium monnieri, Cinnamomum cassia |
| Qi-regulating | Magnolia officinalis flower, Magnolia officinalis bark, Inula racemosa, Aucklandia lappa, Citrus aurantium |
| Hemostatic | Artemisia argyi |
| Blood-activating | Ligusticum chuanxiong, Achyranthes bidentata, Salvia miltiorrhiza, Paeonia lactiflora, Carthamus tinctorius, Panax notoginseng, Curcuma longa, Panax notoginseng flower, Crocus sativus, Sargentodoxa cuneata, Caesalpinia sappan, Cyathula officinalis, Anemone raddeana |
| Phlegm-resolving | Ginkgo biloba |
| Tranquilizing | Polygala tenuifolia |
| Wind-extinguishing | Gastrodia elata |
| Tonic | Lycium barbarum, Epimedium brevicornum, Panax ginseng leaf, Panax ginseng, Atractylodes macrocephala, Fish maw, Angelica sinensis, Ganoderma lucidum, Rehmannia glutinosa, Red ginseng, Glycyrrhiza uralensis, Cordyceps sinensis, Eucommia ulmoides, Cornus officinalis, Polygonatum sibiricum, Ophiopogon japonicus, Gynostemma pentaphyllum, Coriolus versicolor, Dioscorea opposita, Houttuynia cordata, Hippophae rhamnoides, Bletilla striata, Agrimonia pilosa, Ligustrum lucidum |
| Astringent | Schisandra chinensis, Rosa laevigata, Punica granatum peel, Ephedra sinica root, Terminalia chebula |
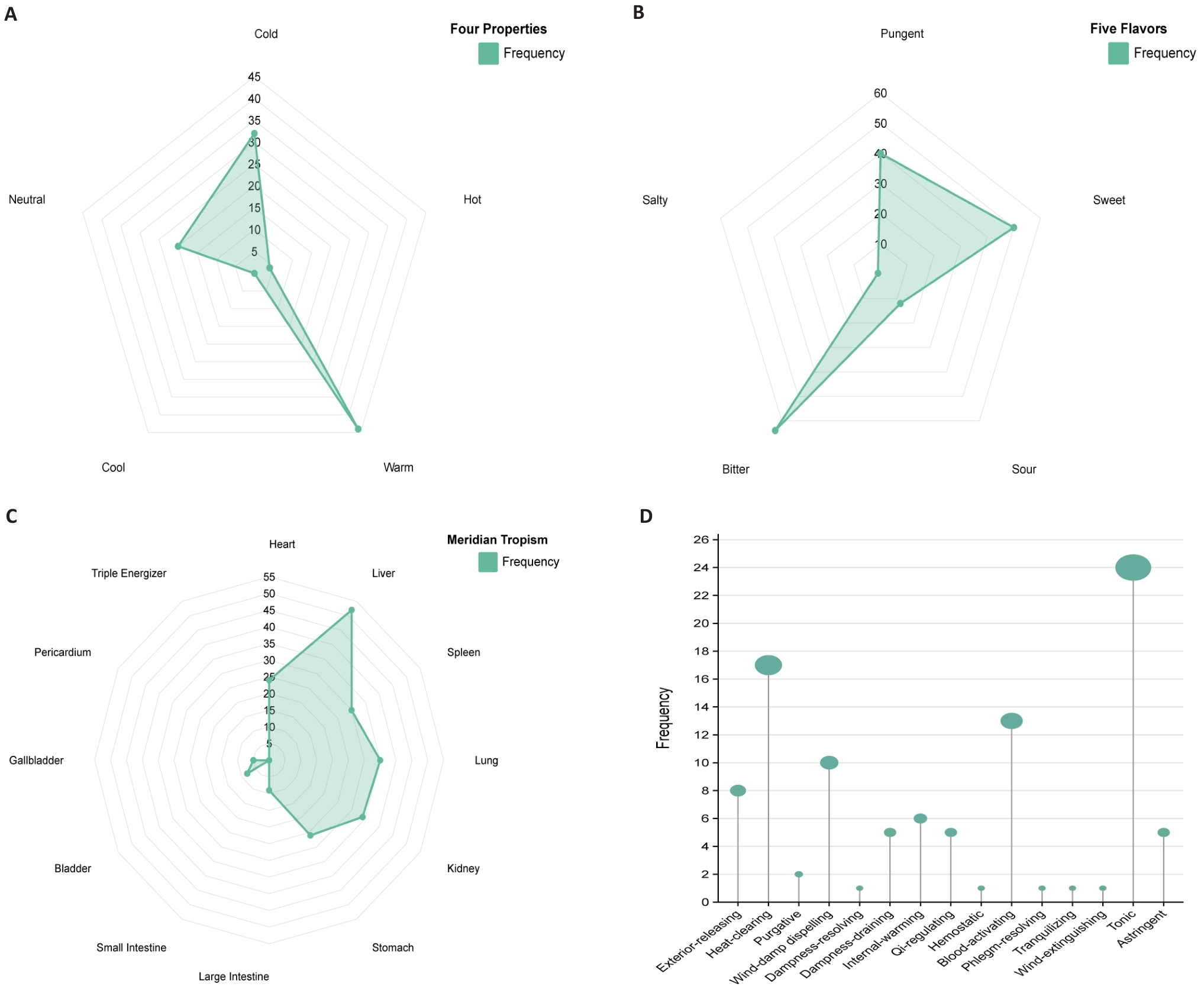
Fig.8 Prediction of traditional Chinese medicines. A: Radar chart of four properties frequency. B: Radar chart of five flavors frequency. C: Radar chart of meridian tropism frequency. D: Frequency diagram of efficacy classification.
| [1] | Hunter DJ, March L, Chew M. Osteoarthritis in 2020 and beyond: a lancet commission[J]. Lancet, 2020, 396(10264): 1711-2. doi:10.1016/s0140-6736(20)32230-3 |
| [2] | Katz JN, Arant KR, Loeser RF. Diagnosis and treatment of hip and knee osteoarthritis: a review[J]. JAMA, 2021, 325(6): 568-78. doi:10.1001/jama.2020.22171 |
| [3] | Richard MJ, Driban JB, McAlindon TE. Pharmaceutical treatment of osteoarthritis[J]. Osteoarthritis Cartilage, 2023, 31(4): 458-66. doi:10.1016/j.joca.2022.11.005 |
| [4] | Del Sordo L, Blackler GB, Philpott HT, et al. Impaired efferocytosis by synovial macrophages in patients with knee osteoarthritis[J]. Arthritis Rheumatol, 2023, 75(5): 685-96. doi:10.1002/art.42412 |
| [5] | Xiong W, Han Z, Ding SL, et al. In situ remodeling of efferocytosis via lesion-localized microspheres to reverse cartilage senescence[J]. Adv Sci: Weinh, 2024, 11(19): e2400345. doi:10.1002/advs.202400345 |
| [6] | Guillem-Llobat P, Marín M, Rouleau M, et al. New insights into the pro-inflammatory and osteoclastogenic profile of circulating monocytes in osteoarthritis patients[J]. Int J Mol Sci, 2024, 25(3): 1710. doi:10.3390/ijms25031710 |
| [7] | Kourtzelis I, Mitroulis I, von Renesse J, et al. From leukocyte recruitment to resolution of inflammation: the cardinal role of integrins[J]. J Leukoc Biol, 2017, 102(3): 677-83. doi:10.1189/jlb.3mr0117-024r |
| [8] | Wang Z, Wang B, Zhang J, et al. Chemokine (C-C motif) ligand 2/chemokine receptor 2 (CCR2) axis blockade to delay chondrocyte hypertrophy as a therapeutic strategy for osteoarthritis[J]. Med Sci Monit, 2021, 27: e930053. doi:10.12659/msm.930053 |
| [9] | Koch AE, Burrows JC, Haines GK, et al. Immunolocalization of endothelial and leukocyte adhesion molecules in human rheumatoid and osteoarthritic synovial tissues[J]. Lab Invest, 1991, 64(3): 313-20. doi:doi:10.1084/jem.173.3.779 |
| [10] | 韩建建, 徐 鹏, 韩礼纲, 等. 关节镜清理术联合透明质酸钠对膝骨关节炎病人关节液细胞间黏附分子-1、白细胞介素-6的影响[J]. 蚌埠医学院学报, 2021, 46(2): 222-5. doi:10.13898/j.cnki.issn.1000-2200.2021.02.022 |
| [11] | Wojdasiewicz P, Poniatowski ŁA, Kotela A, et al. The chemokine CX3CL1 (fractalkine) and its receptor CX3CR1: occurrence and potential role in osteoarthritis[J]. Arch Immunol Ther Exp, 2014, 62(5): 395-403. doi:10.1007/s00005-014-0275-0 |
| [12] | Chen Y, Mehmood K, Chang YF, et al. The molecular mechanisms of glycosaminoglycan biosynthesis regulating chondrogenesis and endochondral ossification[J]. Life Sci, 2023, 335: 122243. doi:10.1016/j.lfs.2023.122243 |
| [13] | Okamura Y, Mishima S, Kashiwakura JI, et al. The dual regulation of substance P-mediated inflammation via human synovial mast cells in rheumatoid arthritis[J]. Allergol Int, 2017, 66: S9-S20. doi:10.1016/j.alit.2017.03.002 |
| [14] | Mustonen AM, Nieminen P. Fatty acids and oxylipins in osteoarthritis and rheumatoid arthritis: a complex field with significant potential for future treatments[J]. Curr Rheumatol Rep, 2021, 23(6): 41. doi:10.1007/s11926-021-01007-9 |
| [15] | Wang Y, Liu J, Huang B, et al. Mathematical modeling and application of IL-1β/TNF signaling pathway in regulating chondrocyte apoptosis[J]. Front Cell Dev Biol, 2023, 11: 1288431. doi:10.3389/fcell.2023.1288431 |
| [16] | Malemud CJ. Matrix metalloproteinases and synovial joint pathology[M]//Matrix Metalloproteinases and Tissue Remodeling in Health and Disease: Target Tissues and Therapy. Amsterdam: Elsevier, 2017: 305-25. doi:10.1016/bs.pmbts.2017.03.003 |
| [17] | Miao Y, Dong Y, Huang P, et al. Increasing UCP2 expression and decreasing NOX1/4 expression maintain chondrocyte phenotype by reducing reactive oxygen species production[J]. Oncotarget, 2017, 8(38): 63750-63. doi:10.18632/oncotarget.18908 |
| [18] | Park D, Han CZ, Elliott MR, et al. Continued clearance of apoptotic cells critically depends on the phagocyte Ucp2 protein[J]. Nature, 2011, 477(7363): 220-4. doi:10.1038/nature10340 |
| [19] | Charlier E, Relic B, Deroyer C, et al. Insights on molecular mechanisms of chondrocytes death in osteoarthritis[J]. Int J Mol Sci, 2016, 17(12): E2146. doi:10.3390/ijms17122146 |
| [20] | Hu S, Zhang C, Ni L, et al. Stabilization of HIF-1α alleviates osteoarthritis via enhancing mitophagy[J]. Cell Death Dis, 2020, 11(6): 481. doi:10.1038/s41419-020-2680-0 |
| [21] | Xia X, Wang S, Ni B, et al. Hypoxic gastric cancer-derived exosomes promote progression and metastasis via miR-301a-3p/PHD3/HIF-1α positive feedback loop[J]. Oncogene, 2020, 39(39): 6231-44. doi:10.1038/s41388-020-01425-6 |
| [22] | Jin Y, Pan Y, Zheng S, et al. Inactivation of EGLN3 hydroxylase facilitates Erk3 degradation via autophagy and impedes lung cancer growth[J]. Oncogene, 2022, 41(12): 1752-66. doi:10.1038/s41388-022-02203-2 |
| [23] | Escribese MM, Sierra-Filardi E, Nieto C, et al. The prolyl hydroxylase PHD3 identifies proinflammatory macrophages and its expression is regulated by activin A[J]. J Immunol, 2012, 189(4): 1946-54. doi:10.4049/jimmunol.1201064 |
| [24] | Samad TA, Moore KA, Sapirstein A, et al. Interleukin-1β-mediated induction of Cox-2 in the CNS contributes to inflammatory pain hypersensitivity[J]. Nature, 2001, 410(6827): 471-5. doi:10.1038/35068566 |
| [25] | Kapoor M, Martel-Pelletier J, Lajeunesse D, et al. Role of proinflammatory cytokines in the pathophysiology of osteoarthritis[J]. Nat Rev Rheumatol, 2011, 7(1): 33-42. doi:10.1038/nrrheum.2010.196 |
| [26] | Sun K, Jing X, Guo J, et al. Mitophagy in degenerative joint diseases[J]. Autophagy, 2021, 17(9): 2082-92. doi:10.1080/15548627.2020.1822097 |
| [27] | Liu Y, Lu T, Liu Z, et al. Six macrophage-associated genes in synovium constitute a novel diagnostic signature for osteoarthritis[J]. Front Immunol, 2022, 13: 936606. doi:10.3389/fimmu.2022.936606 |
| [28] | Wang XY, Han XL, Ma JJ, et al. 5-hydroxymethylfurfural attenuates osteoarthritis by upregulating of glucose metabolism in chondrocytes[J]. Phytomedicine, 2025, 139: 156499. doi:10.1016/j.phymed.2025.156499 |
| [29] | Hao GH, Han SQ, Xiao ZG, et al. Synovial mast cells and osteoarthritis: current understandings and future perspectives[J]. Heliyon, 2024, 10(24): e41003. doi:10.1016/j.heliyon.2024.e41003 |
| [30] | Metcalfe DD, Baram D, Mekori YA. Mast cells[J]. Physiol Rev, 1997, 77(4): 1033-79. doi:10.1152/physrev.1997.77.4.1033 |
| [31] | Fang L, Liu K, Liu C, et al. Tumor accomplice: T cell exhaustion induced by chronic inflammation[J]. Front Immunol, 2022, 13: 979116. doi:10.3389/fimmu.2022.979116 |
| [32] | Holter DB, Zahalka S, Brösamlen J, et al. Mast cells activated in vitro can modulate macrophage polarization and antibacterial responses[J]. J Allergy Clin Immunol, 2025, 156(3): 754-73. doi:10.1016/j.jaci.2025.02.040 |
| [33] | Mihlan M, Wissmann S, Gavrilov A, et al. Neutrophil trapping and nexocytosis, mast cell-mediated processes for inflammatory signal relay[J]. Cell, 2024, 187(19): 5316-35.e28. doi:10.1016/j.cell.2024.07.014 |
| [34] | Poon IKH, Ravichandran KS. Targeting efferocytosis in inflam-maging[J]. Annu Rev Pharmacol Toxicol, 2024, 64: 339-57. doi:10.1146/annurev-pharmtox-032723-110507 |
| [35] | Wherry EJ, Kurachi M. Molecular and cellular insights into T cell exhaustion[J]. Nat Rev Immunol, 2015, 15(8): 486-99. doi:10.1038/nri3862 |
| [36] | Wang TT, He CQ. Pro-inflammatory cytokines: The link between obesity and osteoarthritis[J]. Cytokine Growth Factor Rev, 2018, 44: 38-50. doi:10.1016/j.cytogfr.2018.10.002 |
| [37] | Zhang C, Lin Y, Yan CH, et al. Adipokine signaling pathways in osteoarthritis[J]. Front Bioeng Biotechnol, 2022, 10: 865370. doi:10.3389/fbioe.2022.865370 |
| [38] | Zheng S, Li YT, Yin L, et al. Identification of sulfur metabolism-related gene signature in osteoarthritis and TM9SF2's sustenance effect on M2 macrophages' phagocytic activity[J]. J Orthop Surg Res, 2024, 19(1): 62. doi:10.1186/s13018-023-04384-2 |
| [39] | Peffers MJ, Chabronova A, Balaskas P, et al. SnoRNA signatures in cartilage ageing and osteoarthritis[J]. Sci Rep, 2020, 10(1): 10641. doi:10.1038/s41598-020-67446-z |
| [40] | Song M, Pang L, Zhang M, et al. Cdc2-like kinases: structure, biological function, and therapeutic targets for diseases[J]. Signal Transduct Target Ther, 2023, 8(1): 148. doi:10.1038/s41392-023-01409-4 |
| [41] | 杨 雷, 伍搏宇, 熊 辉, 等. 基于基因芯片技术结合CIBERSORT反卷积算法研究骨关节炎的免疫机制及潜在中药靶向预测[J]. 中国免疫学杂志, 2022, 38(3): 288-95. doi:10.3969/j.issn.1000-484X.2022.03.006 |
| [42] | 唐仕荣, 巫永华, 刘恩岐, 等. 枸杞多糖的提取分级及其氧自由基清除能力分析[J]. 食品科技, 2018, 43(10): 251-6. |
| [43] | 蔡松涛, 孙京涛, 魏 瑄. 枸杞多糖抑制核因子κB(NF-κB)通路降低骨关节炎软骨细胞炎性细胞因子水平[J]. 细胞与分子免疫学杂志, 2018, 34(11): 989-93. |
| [44] | 赵 飞, 丁 冬, 巩 凡, 等. 枸杞多糖干预膝骨关节炎模型兔关节软骨组织CD151和基质金属蛋白酶3的表达[J]. 中国组织工程研究, 2020, 24(17): 2648-53. doi:10.3969/j.issn.2095-4344.2562 |
| [45] | Mi B, Wang J, Liu Y, et al. Icariin activates autophagy via down-regulation of the NF‑κB signaling-mediated apoptosis in chondro-cytes[J]. Front Pharmacol, 2018, 9: 605. doi:10.3389/fphar.2018.00605 |
| [46] | Huang H, Zhang ZF, Qin FW, et al. Icariin inhibits chondrocyte apoptosis and angiogenesis by regulating the TDP-43 signaling pathway[J]. Molec Gen & Gen Med, 2019, 7(4): e00586. doi:10.1002/mgg3.586 |
| [47] | Zhu PT, Wu Y, Yang AH, et al. Catalpol suppressed proliferation, growth and invasion of CT26 colon cancer by inhibiting inflammation and tumor angiogenesis[J]. Biomed Pharmacother, 2017, 95: 68-76. doi:10.1016/j.biopha.2017.08.049 |
| [48] | Zeng YF, Wang R, Bian Y, et al. Catalpol attenuates IL-1β induced matrix catabolism, apoptosis and inflammation in rat chondrocytes and inhibits cartilage degeneration[J]. Med Sci Monit, 2019, 25: 6649-59. doi:10.12659/msm.916209 |
| [49] | Lu S, Xiao X, Cheng M. Matrine inhibits IL-1β-induced expression of matrix metalloproteinases by suppressing the activation of MAPK and NF-κB in human chondrocytes in vitro [J]. Int J Clin Exp Pathol, 2015, 8(5): 4764-72. doi:10.1016/S1359-6349(09)71986-X |
| [50] | Wu DY, Zhu XY, Kang XD, et al. The protective effect of sophocarpine in osteoarthritis: an in vitro and in vivo study[J]. Int Immunopharmacol, 2019, 67: 145-51. doi:10.1016/j.intimp.2018.11.046 |
| [51] | 梁桂洪, 曾令烽, 潘建科, 等. 川芎嗪干预骨性关节炎的机制研究进展[J]. 中华中医药杂志, 2020, 35(12): 6228-32. |
| [52] | 李飞龙, 谢平金, 柴生颋, 等. 川芎嗪对膝骨性关节炎大鼠软骨VEGF表达的影响[J]. 中国骨质疏松杂志, 2018, 24(7): 904-9. |
| [53] | 朱海泉, 刘子敏, 孟祥圣, 等. 川芎嗪抑制NF-κB P65磷酸化对LPS诱导的骨关节炎软骨细胞凋亡和炎症反应的调节作用[J]. 中国免疫学杂志, 2019, 35(2): 181-5. |
| [54] | 高 坤, 张 勇, 陈大宇, 等. 牛膝总皂苷干预兔膝骨关节炎滑膜液来源细胞因子的表达[J]. 中国组织工程研究, 2019, 23(33): 5317-21. |
| [55] | 孙雪莲, 刘 渊, 周红海. 牛膝总皂苷对兔膝骨关节炎软骨组织形态变化及关节液中IL-1β、TGF-β1含量的影响[J]. 中药新药与临床药理, 2016, 27(3): 321-6. |
| [56] | Xu X, Zhang X, Diao Y, et al. Achyranthes bidentate saponins protect rat articular chondrocytes against interleukin-1β-induced inflammation and apoptosis in vitro[J]. Kaohsiung J Med Sci, 2017,33(2):62-8. doi:10.1016/j.kjms.2016.11.004 |
| [57] | Wei B, Zhang Y, Tang L, et al. Protective effects of quercetin against inflammation and oxidative stress in a rabbit model of knee osteoarthritis[J]. Drug Dev Res, 2019, 80(3): 360-7. doi:10.1002/ddr.21510 |
| [1] | Hongcheng TAO, Fukai LIANG, Wenbo HUANG, Siqi FAN, Ping ZENG. Analysis of potential association of alcohol exposure with femoral head osteonecrosis and construction of a diagnostic model using machine learning [J]. Journal of Southern Medical University, 2026, 46(3): 559-569. |
| [2] | Jiahui QIU, Meng CHEN, Ru MAN, Xin CHEN, Dongrui QIU, Qitong CHANG, Hongyu MA. Huayu Tongbian Decoction promotes efferocytosis of interstitial Cajal cells in rats with slow transit constipation by inhibiting the PI3K/Akt signaling pathway [J]. Journal of Southern Medical University, 2026, 46(2): 293-300. |
| [3] | Junyao CHEN, Zeyu CHEN, Zhaojie LIN, Menghao FANG, Chaoying SHEN, Qi XU, Xiaoyi ZHANG, Lu LU. Dual role of tea consumption in gastrointestinal disease risks: analysis using a risk prediction model integrating interpretable machine learning and large language model [J]. Journal of Southern Medical University, 2026, 46(2): 353-361. |
| [4] | Yunneng CUI, Minqing FENG, Liangfeng YAO, Jiewen YAN, Wenhan LI, Yanping HUANG. Enhancement of radiomics-based machine learning models for predicting efficacy of high-intensity focused ultrasound ablation of uterine fibroids using undersampling methods [J]. Journal of Southern Medical University, 2026, 46(1): 141-149. |
| [5] | Zhe WANG, Keyu KONG, Minghao JIN, Sonu NG, Wenxuan FAN, Zanjing ZHAI, Zihao HU, Lin NIU, Yansong QI, Yongsheng XU. Exosomes from folic acid-treated subpatellar fat pad-derived mesenchymal stem cells promote M2 polarization of macrophages in vitro [J]. Journal of Southern Medical University, 2026, 46(1): 166-174. |
| [6] | Qizhi HUANG, Daipeng XIE, Lintong YAO, Qiaxuan LI, Shaowei WU, Haiyu ZHOU. Tumor microenvironment-specific CT radiomics signature for predicting immunotherapy response in non-small cell lung cancer [J]. Journal of Southern Medical University, 2025, 45(9): 1903-1918. |
| [7] | Ying WANG, Jing LI, Yidi WANG, Mingyu HUA, Weibin HU, Xiaozhi ZHANG. Construction and verification of a prognostic model combining anoikis and immune prognostic signatures for primary liver cancer [J]. Journal of Southern Medical University, 2025, 45(9): 1967-1979. |
| [8] | Jun JIANG, Shuo FENG, Yingui SUN, Yan AN. Construction of risk prediction models of hypothermia after transurethral holmium laser enucleation of the prostate based on three machine learning algorithms [J]. Journal of Southern Medical University, 2025, 45(9): 2019-2025. |
| [9] | Xinyuan CHEN, Chengting WU, Ruidi LI, Xueqin PAN, Yaodan ZHANG, Junyu TAO, Caizhi LIN. Shuangshu Decoction inhibits growth of gastric cancer cell xenografts by promoting cell ferroptosis via the P53/SLC7A11/GPX4 axis [J]. Journal of Southern Medical University, 2025, 45(7): 1363-1371. |
| [10] | Kang WANG, Haibin LI, Jing YU, Yuan MENG, Hongli ZHANG. High expression of ELFN1 is a prognostic biomarker and promotes proliferation and metastasis of colorectal cancer cells [J]. Journal of Southern Medical University, 2025, 45(7): 1543-1553. |
| [11] | Meimei CHEN, Yang WANG, Huangwei LEI, Fei ZHANG, Ruina HUANG, Zhaoyang YANG. Construction of recognition models for subthreshold depression based on multiple machine learning algorithms and vocal emotional characteristics [J]. Journal of Southern Medical University, 2025, 45(4): 711-717. |
| [12] | Zhi GAO, Ao WU, Zhongxiang HU, Peiyang SUN. Bioinformatics analysis of oxidative stress and immune infiltration in rheumatoid arthritis [J]. Journal of Southern Medical University, 2025, 45(4): 862-870. |
| [13] | Liping FU, Lixia YUAN, Jie WANG, Xuelan CHEN, Guizhi KE, Yu HUANG, Xinyi YANG, Gang LIU. Advances of low-intensity pulsed ultrasound for treatment of musculoskeletal disorders in the past decade [J]. Journal of Southern Medical University, 2025, 45(3): 661-668. |
| [14] | Changlong FU, Ruolan CHEN, Shiqi XU, Jinxin YOU, Qing LIN, Yanfeng HUANG. Morinda officinalis polysaccharide delays osteoarthritis mouse chondrocyte degeneration by modulating the glycolysis-pyroptosis axis via targeting the lncRNA XIST [J]. Journal of Southern Medical University, 2025, 45(12): 2541-2550. |
| [15] | Yongxin MAI, Shuting ZHOU, Ruijia WEN, Jinfang ZHANG, Dongxiang ZHAN. Aucubin alleviates knee osteoarthritis in mice by suppressing the NF‑κB signaling pathway [J]. Journal of Southern Medical University, 2025, 45(10): 2104-2110. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||