Journal of Southern Medical University ›› 2025, Vol. 45 ›› Issue (7): 1543-1553.doi: 10.12122/j.issn.1673-4254.2025.07.22
Previous Articles Next Articles
Kang WANG1( ), Haibin LI1, Jing YU1, Yuan MENG2, Hongli ZHANG3(
), Haibin LI1, Jing YU1, Yuan MENG2, Hongli ZHANG3( )
)
Received:2025-02-26
Online:2025-07-20
Published:2025-07-17
Contact:
Hongli ZHANG
E-mail:wk19950303@smu.edu.cn;13926231602@163.com
Kang WANG, Haibin LI, Jing YU, Yuan MENG, Hongli ZHANG. High expression of ELFN1 is a prognostic biomarker and promotes proliferation and metastasis of colorectal cancer cells[J]. Journal of Southern Medical University, 2025, 45(7): 1543-1553.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.j-smu.com/EN/10.12122/j.issn.1673-4254.2025.07.22
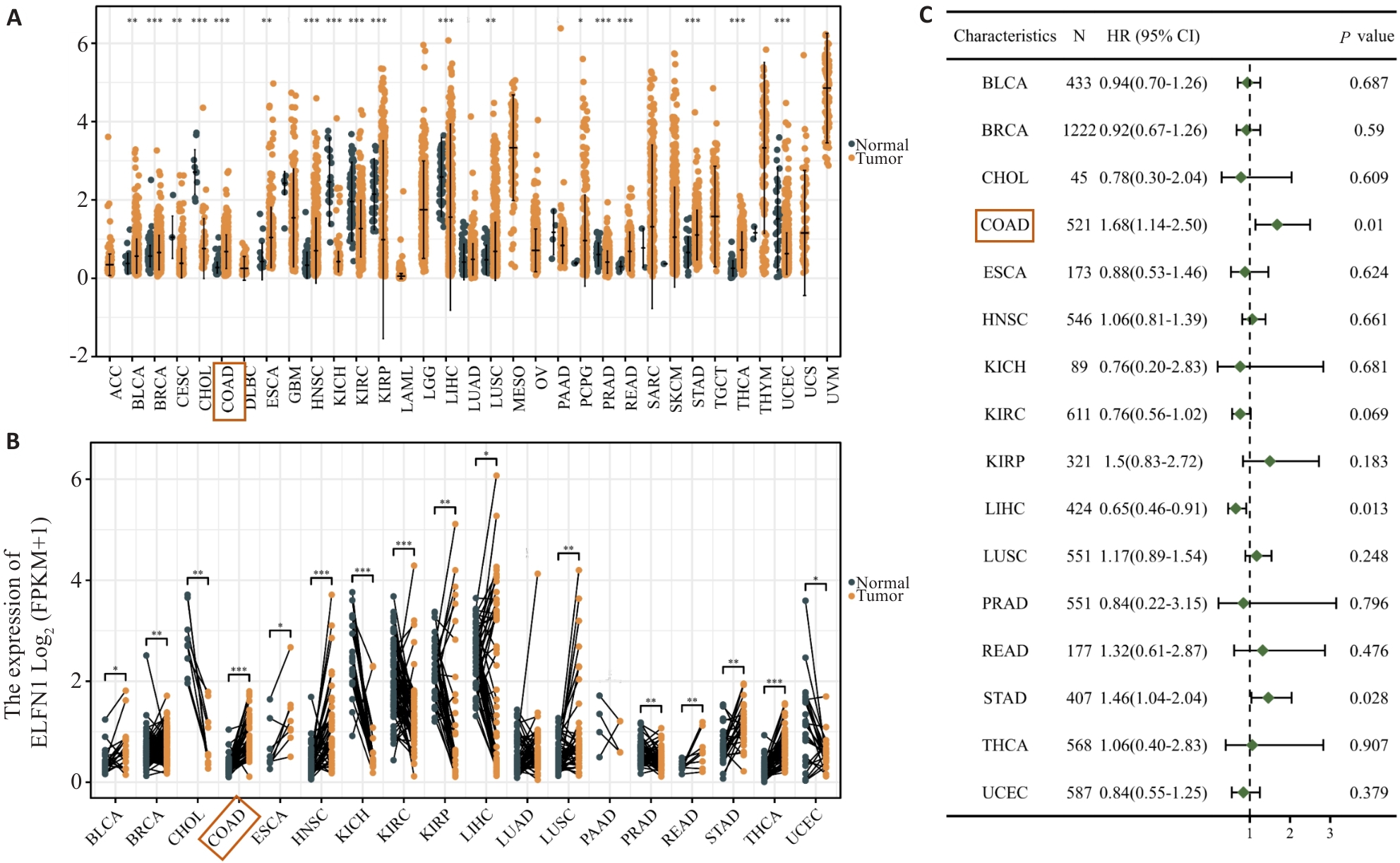
Fig.1 Data from the TCGA database show high expression of ELFN1 in a variety of tumors, which has different prognostic implications. A: TCGA diffuse carcinoma analysis. B: TCGA pan-cancer pairing analysis. C: Forest map of the prognostic value of ELFN1 in different tumors. *P<0.05, **P<0.01, ***P<0.001.
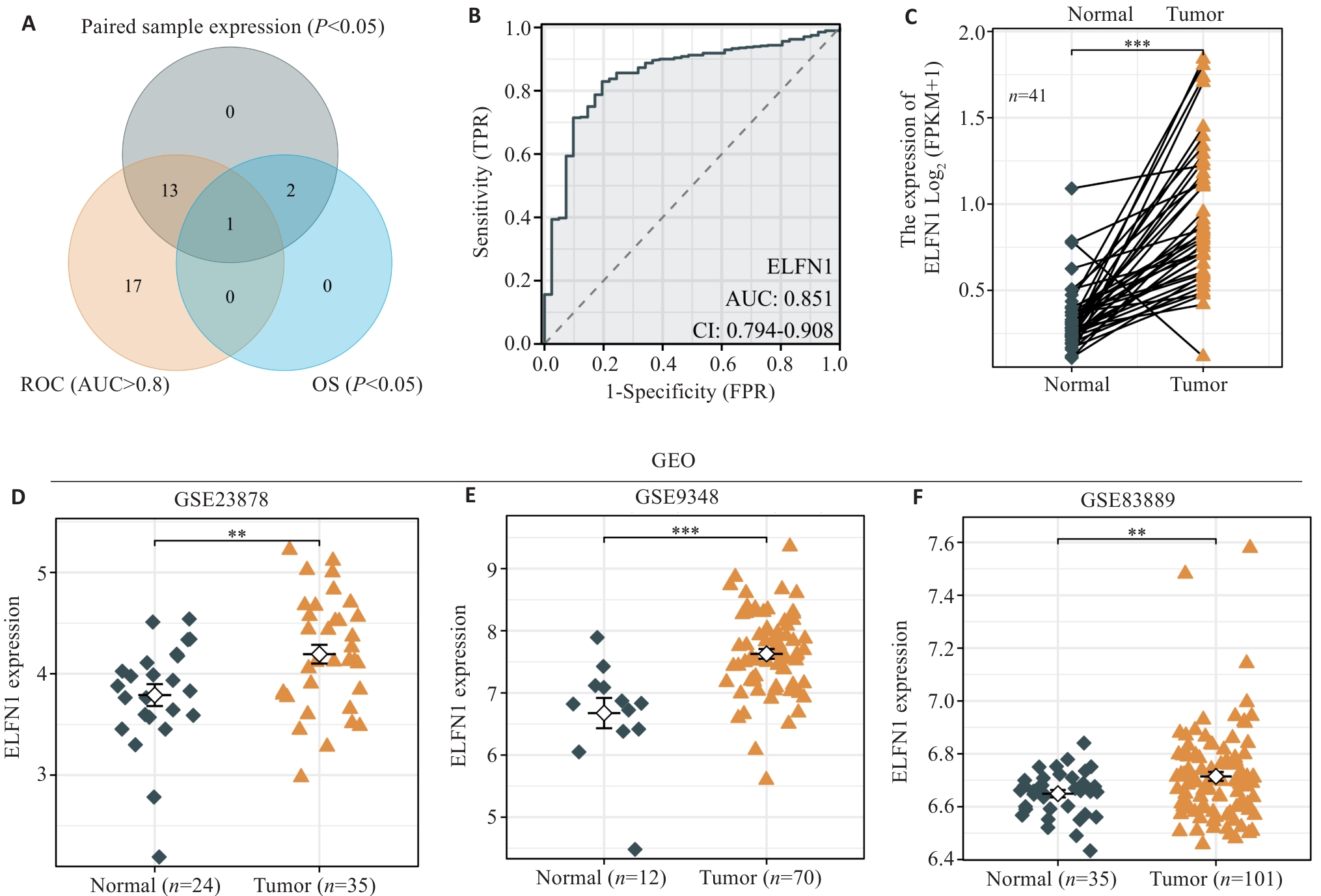
Fig.2 ELFN1 has a high diagnostic value in colon cancer and is highly expressed in colon cancer. A: Venn diagram of ELFN1 expression patterns and its prognosis and diagnostic value of colon cancer. B: ROC curve analysis showing an AUC value of ELFN1 in COAD of 0.851. C: High expression levels of ELFN1 in paired samples of TCGA. D-F: Multiple datasets in the GEO database indicate that the expression level of ELFN1 is higher tumor tissues than in normal tissues. *P<0.05, **P<0.01, ***P<0.001.
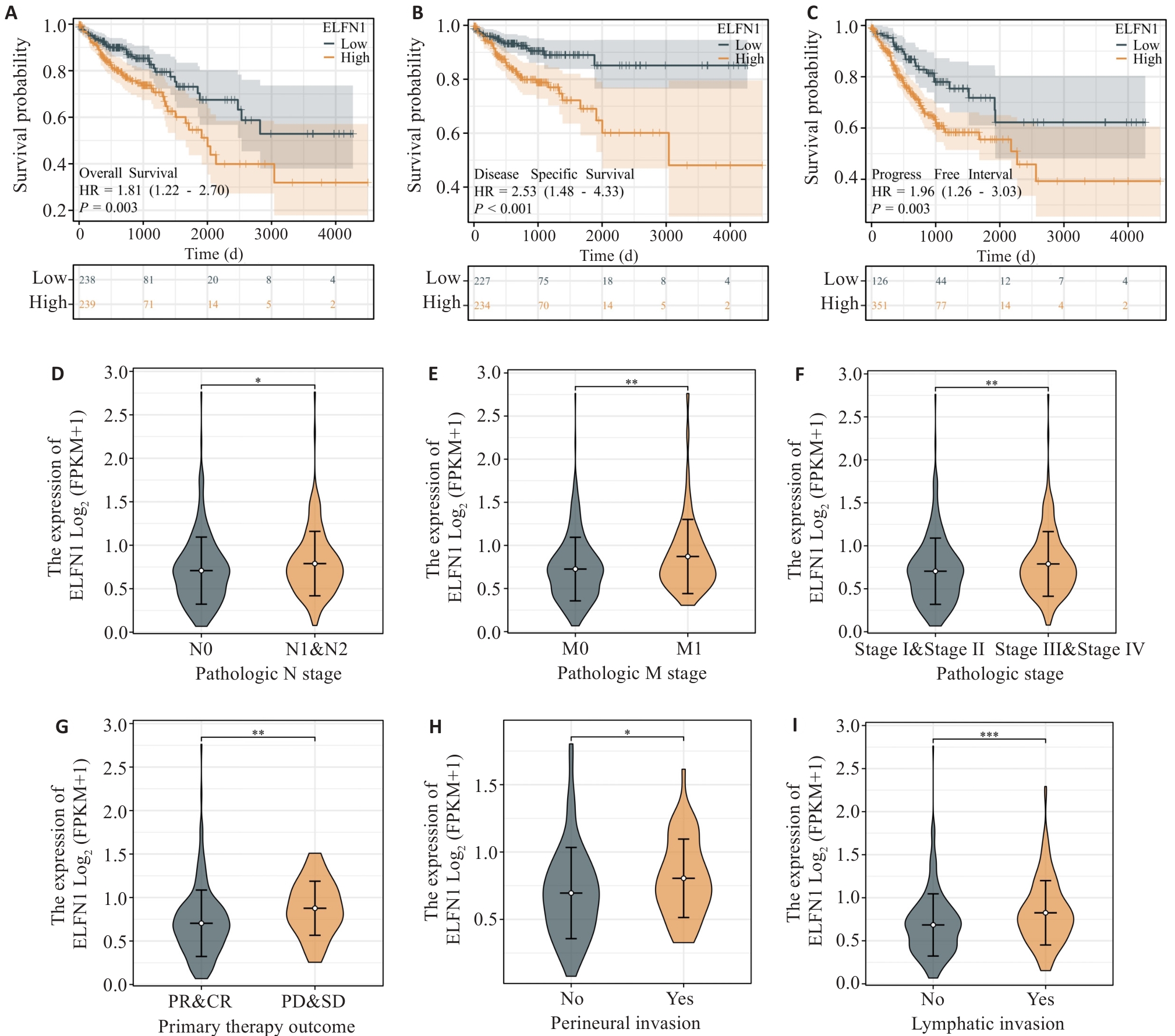
Fig.3 Prognostic and clinical significance of ELFN1 in colon cancer. A-C: Kaplan-Meier survival analysis reveals that COAD patients with high ELFN1 expression have shorter overall survival (A), disease-specific survival (B) and progression-free interval (C) than those with low ELFN1 expression. D-I: mRNA expression level of ELFN1 in COAD patients with different clinicopathological features. *P<0.05, **P<0.01, ***P<0.001.
| Characteristics | Total (n) | Univariate analysis | Multivariate analysis | |||
|---|---|---|---|---|---|---|
| Hazard ratio (95%CI) | P | Hazard ratio (95%CI) | P | |||
| Age (year) | 477 | |||||
| ≤65 | 194 | Reference | ||||
| >65 | 283 | 1.610 (1.052-2.463) | 0.028 | 3.417 (1.309-8.920) | 0.012 | |
| T stage | 476 | |||||
| T1&T2 | 94 | Reference | ||||
| T3&T4 | 382 | 3.072 (1.423-6.631) | 0.004 | 0.661 (0.120-3.637) | 0.634 | |
| N stage | 477 | |||||
| N0 | 283 | Reference | ||||
| N1&N2 | 194 | 2.592 (1.743-3.855) | <0.001 | 0.174 (0.016-1.839) | 0.146 | |
| M stage | 414 | |||||
| M0 | 348 | Reference | ||||
| M1 | 66 | 4.193 (2.683-6.554) | <0.001 | 5.463 (1.807-16.518) | 0.003 | |
| Pathologic stage | 466 | |||||
| Stage I&Stage II | 267 | Reference | ||||
| Stage III&Stage IV | 199 | 2.947 (1.942-4.471) | <0.001 | 14.791(1.177-185.875) | 0.037 | |
| Perineural invasion | 181 | |||||
| No | 135 | Reference | ||||
| Yes | 46 | 1.940 (0.982-3.832) | 0.056 | 0.736 (0.280-1.935) | 0.534 | |
| Lymphatic invasion | 433 | |||||
| No | 265 | Reference | ||||
| Yes | 168 | 2.450 (1.614-3.720) | <0.001 | 1.408 (0.557-3.556) | 0.470 | |
| ELFN1 | 477 | |||||
| Low | 238 | Reference | ||||
| High | 239 | 1.683 (1.135-2.496) | 0.010 | 3.284 (1.154-9.339) | 0.026 | |
Tab.1 Univariate and multivariate Cox regression analyses of overall survival of COAD patients based on ELFN1 expression levels
| Characteristics | Total (n) | Univariate analysis | Multivariate analysis | |||
|---|---|---|---|---|---|---|
| Hazard ratio (95%CI) | P | Hazard ratio (95%CI) | P | |||
| Age (year) | 477 | |||||
| ≤65 | 194 | Reference | ||||
| >65 | 283 | 1.610 (1.052-2.463) | 0.028 | 3.417 (1.309-8.920) | 0.012 | |
| T stage | 476 | |||||
| T1&T2 | 94 | Reference | ||||
| T3&T4 | 382 | 3.072 (1.423-6.631) | 0.004 | 0.661 (0.120-3.637) | 0.634 | |
| N stage | 477 | |||||
| N0 | 283 | Reference | ||||
| N1&N2 | 194 | 2.592 (1.743-3.855) | <0.001 | 0.174 (0.016-1.839) | 0.146 | |
| M stage | 414 | |||||
| M0 | 348 | Reference | ||||
| M1 | 66 | 4.193 (2.683-6.554) | <0.001 | 5.463 (1.807-16.518) | 0.003 | |
| Pathologic stage | 466 | |||||
| Stage I&Stage II | 267 | Reference | ||||
| Stage III&Stage IV | 199 | 2.947 (1.942-4.471) | <0.001 | 14.791(1.177-185.875) | 0.037 | |
| Perineural invasion | 181 | |||||
| No | 135 | Reference | ||||
| Yes | 46 | 1.940 (0.982-3.832) | 0.056 | 0.736 (0.280-1.935) | 0.534 | |
| Lymphatic invasion | 433 | |||||
| No | 265 | Reference | ||||
| Yes | 168 | 2.450 (1.614-3.720) | <0.001 | 1.408 (0.557-3.556) | 0.470 | |
| ELFN1 | 477 | |||||
| Low | 238 | Reference | ||||
| High | 239 | 1.683 (1.135-2.496) | 0.010 | 3.284 (1.154-9.339) | 0.026 | |
| Characteristic | Total (n) | OR (95%CI) | P |
|---|---|---|---|
| Pathologic T stage (T3&T4 vs T1&T2) | 477 | 1.831 (0.975-3.600) | 0.069 |
| Pathologic N stage (N1&N2 vs N0) | 478 | 1.743 (1.078-2.851) | 0.025 |
| Pathologic M stage (M1 vs M0) | 415 | 2.425 (1.286-4.572) | 0.006 |
| Pathologic stage (Stage III&IV vs Stage I&II) | 467 | 1.787 (1.104-2.936) | 0.020 |
| Primary therapy outcome (PD&SD vs PR&CR) | 250 | 2.834 (1.120-7.152) | 0.025 |
| Perineural invasion (Yes vs No) | 181 | 2.675 (0.984-7.424) | 0.044 |
| Lymphatic invasion (Yes vs No) | 434 | 2.823 (1.653-4.934) | <0.001 |
Tab.2 Logistic regression analysis of ELFN1
| Characteristic | Total (n) | OR (95%CI) | P |
|---|---|---|---|
| Pathologic T stage (T3&T4 vs T1&T2) | 477 | 1.831 (0.975-3.600) | 0.069 |
| Pathologic N stage (N1&N2 vs N0) | 478 | 1.743 (1.078-2.851) | 0.025 |
| Pathologic M stage (M1 vs M0) | 415 | 2.425 (1.286-4.572) | 0.006 |
| Pathologic stage (Stage III&IV vs Stage I&II) | 467 | 1.787 (1.104-2.936) | 0.020 |
| Primary therapy outcome (PD&SD vs PR&CR) | 250 | 2.834 (1.120-7.152) | 0.025 |
| Perineural invasion (Yes vs No) | 181 | 2.675 (0.984-7.424) | 0.044 |
| Lymphatic invasion (Yes vs No) | 434 | 2.823 (1.653-4.934) | <0.001 |
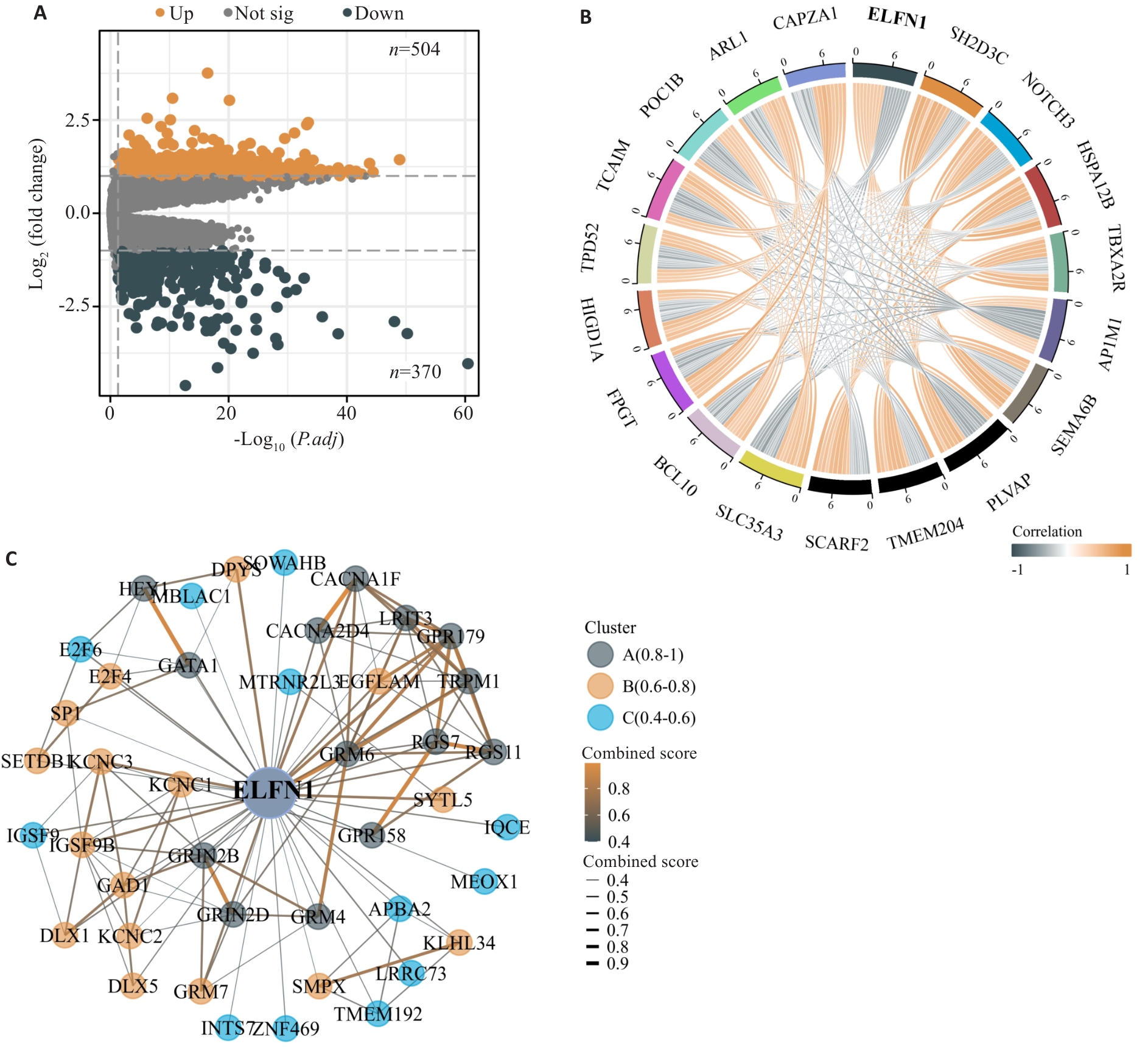
Fig.4 Analysis of differentially expressed genes related to ELFN1 and construction of the PPI network. A: Volcano plot of differentially expressed genes (DEGs, |log2(FC)|>1, P.adj<0.05). B: The co-expression chord diagram showing the top 10 genes positively and negatively correlated with ELFN1. C: Molecular protein interaction network of ELFN1.
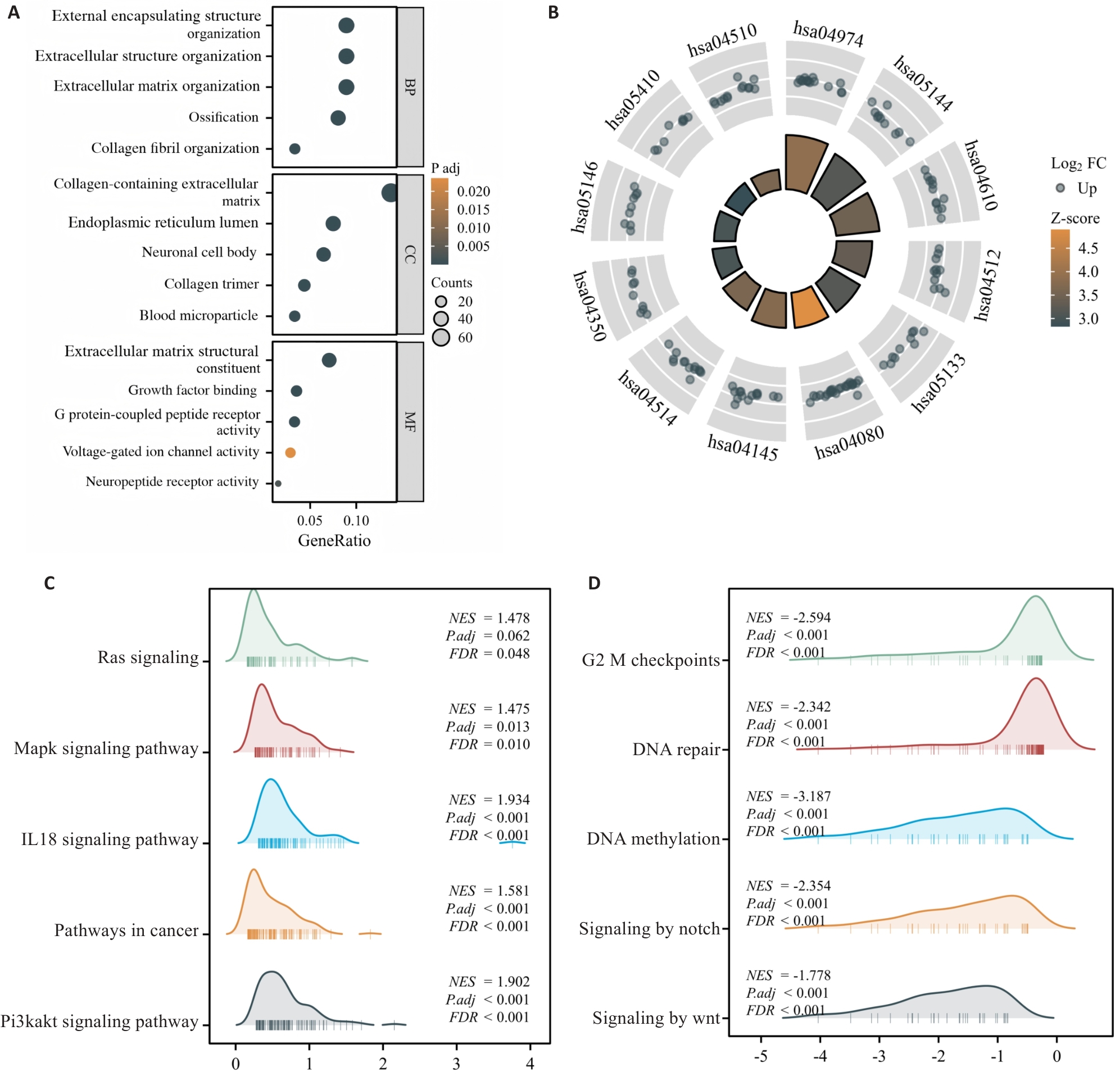
Fig.5 ELFN1 interaction protein identification and gene function annotation analysis. A: GO functional annotation analysis of biological process (BP), cellular component (CC) and molecular function (MF). B: Analysis of 10 KEGG signaling pathways related to ELFN1. C, D: The 10 GSEA pathways associated with ELFN1 in COAD.
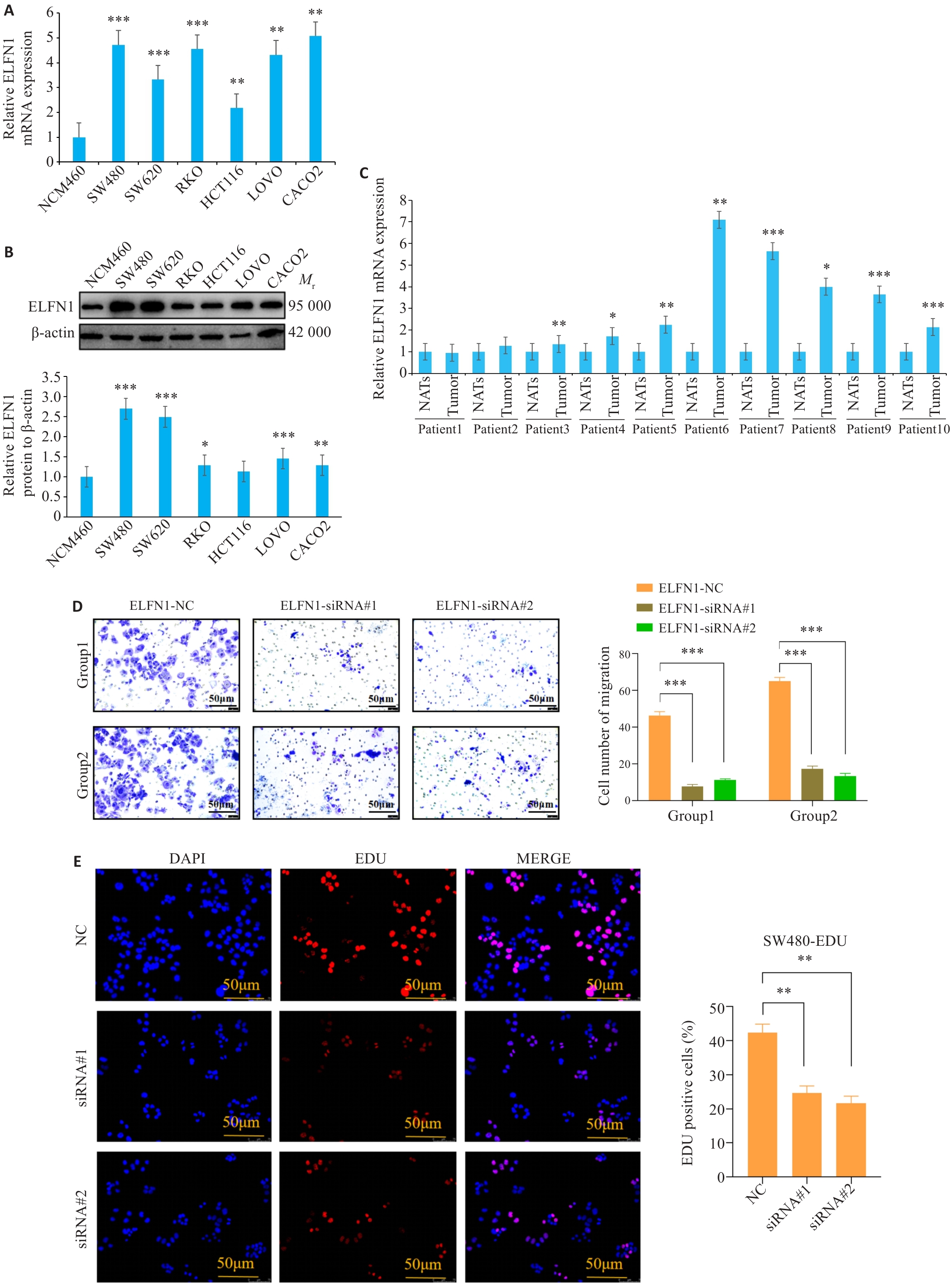
Fig.6 Expression and function of ELFN1 in COAD cell lines. A, B: ELFN1 expression levels in NCM460, SW480, SW620, RKO, HCT116, LOVO, and CACO2 cells detected by qRT-PCR (A) and Western blotting (B). *P<0.05, **P<0.01, ***P<0.001 vs NCM460 cells. C: Expression levels of ELFN1 in 10 pairs of colon cancer tissue samples (*P<0.05, **P<0.01, ***P<0.001 vs NATs). D: Transwell assay of SW480 cells transfected with si-ELFN1. E: EDU proliferation assay of SW480 cells transfected with si-ELFN1. **P<0.01, ***P<0.001.
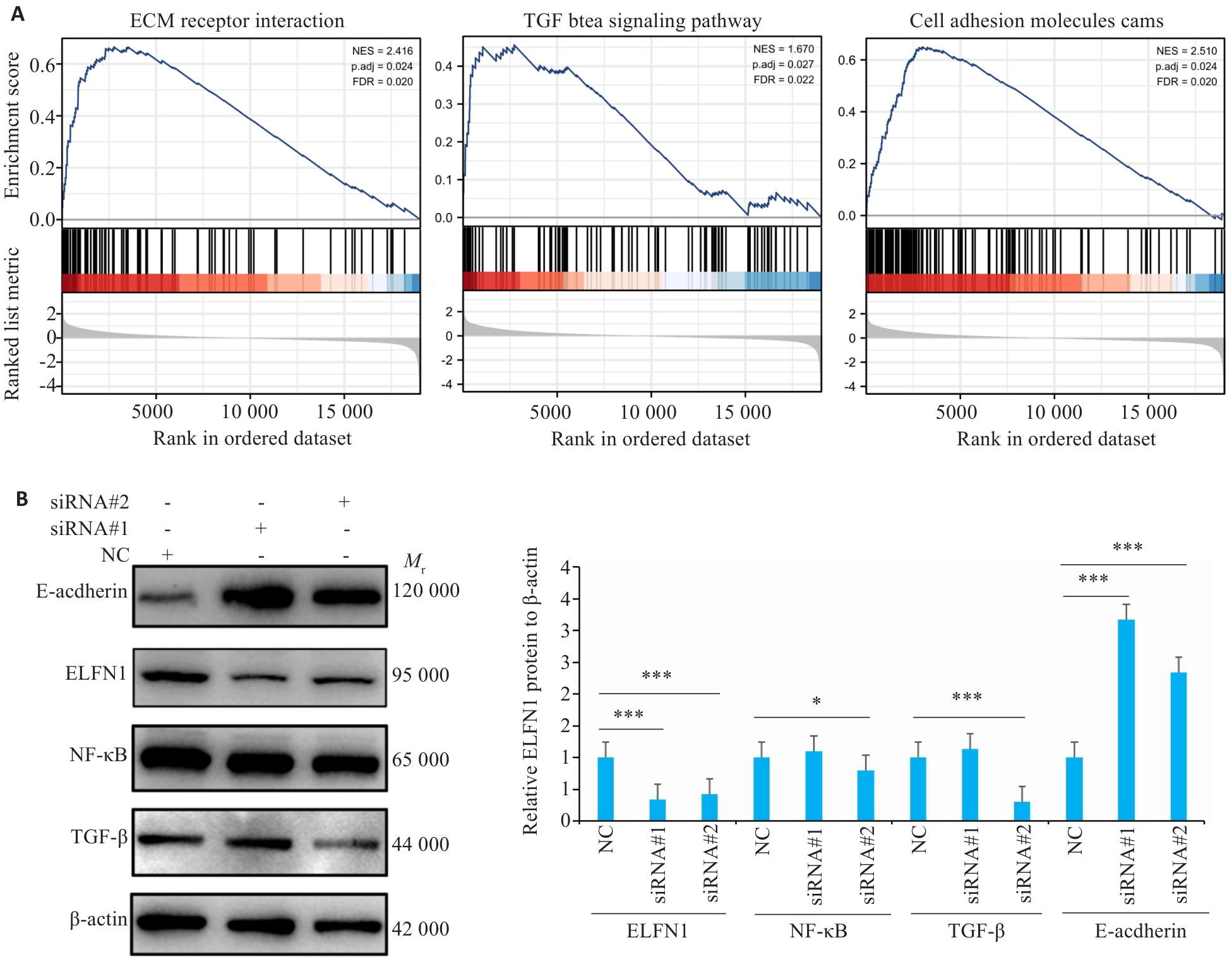
Fig.7 Molecular mechanism of ELFN1 in the oncogenic signaling pathway of colorectal cancer. A: Analysis of relevant carcinogenic pathways of ELFN1 in COAD. B: Western blotting for detecting protein expressions in COAD cells with ELFN1 knockdown. *P<0.05, ***P<0.001.
| [1] | Stachniak TJ, Sylwestrak EL, Scheiffele P, et al. Elfn1-induced constitutive activation of mGluR7 determines frequency-dependent recruitment of somatostatin interneurons[J]. J Neurosci, 2019, 39(23): 4461-74. doi:10.1523/jneurosci.2276-18.2019 |
| [2] | Dunn HA, Patil DN, Cao Y, et al. Synaptic adhesion protein ELFN1 is a selective allosteric modulator of group III metabotropic glutamate receptors in trans [J]. Proc Natl Acad Sci USA, 2018, 115(19): 5022-7. doi:10.1073/pnas.1722498115 |
| [3] | Tomioka NH, Yasuda H, Miyamoto H, et al. Elfn1 recruits presynaptic mGluR7 in trans and its loss results in seizures[J]. Nat Commun, 2014, 5: 4501. doi:10.1038/ncomms5501 |
| [4] | Girgenti MJ, Wang JW, Ji DJ, et al. Transcriptomic organization of the human brain in post-traumatic stress disorder[J]. Nat Neurosci, 2021, 24(1): 24-33. doi:10.1038/s41593-020-00748-7 |
| [5] | Zhang C, Lian H, Xie L, et al. LncRNA ELFN1-AS1 promotes esophageal cancer progression by up-regulating GFPT1 via sponging miR-183-3p[J]. Biol Chem, 2020, 401(9): 1053-61. doi:10.1515/hsz-2019-0430 |
| [6] | Jie YK, Ye L, Chen H, et al. ELFN1-AS1 accelerates cell proliferation, invasion and migration via regulating miR-497-3p/CLDN4 axis in ovarian cancer[J]. Bioengineered, 2020, 11(1): 872-82. doi:10.1080/21655979.2020.1797281 |
| [7] | Feng WG, Zhu RX, Ma JL, et al. LncRNA ELFN1-AS1 promotes retinoblastoma growth and invasion via regulating miR-4270/SBK1 axis[J]. Cancer Manag Res, 2021, 13: 1067-73. doi:10.2147/cmar.s281536 |
| [8] | Ma G, Li GC, Gou AJ, et al. Long non-coding RNA ELFN1-AS1 in the pathogenesis of pancreatic cancer[J]. Ann Transl Med, 2021, 9(10): 877. doi:10.21037/atm-21-2376 |
| [9] | Office FE. Retraction: Long non-coding RNA ELFN1-AS1 promoted colon cancer cell growth and migration via the miR-191-5p/special AT-rich sequence-binding protein 1 axis[J]. Front Oncol, 2023, 13: 1322464. doi:10.3389/fonc.2023.1322464 |
| [10] | Lei R, Feng LC, Hong D. ELFN1-AS1 accelerates the proliferation and migration of colorectal cancer via regulation of miR-4644/TRIM44 axis[J]. Cancer Biomark, 2020, 27(4): 433-43. doi:10.3233/cbm-190559 |
| [11] | Li YM, Gan YQ, Liu JX, et al. Downregulation of MEIS1 mediated by ELFN1-AS1/EZH2/DNMT3a axis promotes tumorigenesis and oxaliplatin resistance in colorectal cancer[J]. Signal Transduct Target Ther, 2022, 7(1): 87. doi:10.1038/s41392-022-00902-6 |
| [12] | Dekker E, Tanis PJ, Vleugels JLA, et al. Colorectal cancer[J]. Lancet, 2019, 394(10207): 1467-80. doi:10.1016/s0140-6736(19)32319-0 |
| [13] | Eng C, Jácome AA, Agarwal R, et al. A comprehensive framework for early-onset colorectal cancer research[J]. Lancet Oncol, 2022, 23(3): e116-28. doi:10.1016/s1470-2045(21)00588-x |
| [14] | Libé-Philippot B, Lejeune A, Wierda K, et al. LRRC37B is a human modifier of voltage-gated sodium channels and axon excitability in cortical neurons[J]. Cell, 2023, 186(26): 5766-83.e25. doi:10.1016/j.cell.2023.11.028 |
| [15] | Huang JX, Yuan WW, Chen BB, et al. lncRNA ELFN1-AS1 upregulates TRIM29 by suppressing miR-211-3p to promote gastric cancer progression[J]. Acta Biochim Biophys Sin (Shanghai), 2023, 55(3): 484-97. doi:10.3724/abbs.2023023 |
| [16] | Deng X, Kong FY, Li S, et al. A KLF4/PiHL/EZH2/HMGA2 regulatory axis and its function in promoting oxaliplatin-resistance of colorectal cancer[J]. Cell Death Dis, 2021, 12(5): 485. doi:10.1038/s41419-021-03753-1 |
| [17] | Huang L, Xiao W, Wang Y, et al. Metabotropic glutamate receptors (mGluRs) in epileptogenesis: an update on abnormal mGluRs signaling and its therapeutic implications[J]. Neural Regen Res, 2024, 19(2): 360-8. doi:10.4103/1673-5374.379018 |
| [18] | Becker JAJ, Pellissier LP, Corde Y, et al. Facilitating mGluR4 activity reverses the long-term deleterious consequences of chronic morphine exposure in male mice[J]. Neuropsychopharmacology, 2021, 46(7): 1373-85. doi:10.1038/s41386-020-00927-x |
| [19] | Qin X, Cardoso Rodriguez F, Sufi J, et al. An oncogenic phenoscape of colonic stem cell polarization[J]. Cell, 2023, 186(25): 5554-68.e18. doi:10.1016/j.cell.2023.11.004 |
| [20] | Chen CK, Ng CS, Wu SM, et al. Regulatory differences in natal down development between altricial Zebra finch and precocial chicken[J]. Mol Biol Evol, 2016, 33(8): 2030-43. doi:10.1093/molbev/msw085 |
| [21] | Taylor KR, Barron T, Hui A, et al. Glioma synapses recruit mechanisms of adaptive plasticity[J]. Nature, 2023, 623(7986): 366-74. doi:10.1038/s41586-023-06678-1 |
| [22] | Iijima N, Sato K, Kuranaga E, et al. Differential cell adhesion implemented by Drosophila Toll corrects local distortions of the anterior-posterior compartment boundary[J]. Nat Commun, 2020, 11(1): 6320. doi:10.1038/s41467-020-20118-y |
| [23] | Valle JW, Lamarca A, Goyal L, et al. New horizons for precision medicine in biliary tract cancers[J]. Cancer Discov, 2017, 7(9): 943-62. doi:10.1158/2159-8290.cd-17-0245 |
| [24] | Dai J, Su YZ, Zhong SY, et al. Exosomes: key players in cancer and potential therapeutic strategy[J]. Signal Transduct Target Ther, 2020, 5(1): 145. doi:10.1038/s41392-020-00261-0 |
| [25] | Dunn HA, Dhaliwal SK, Chang CT, et al. Distinct autoregulatory roles of ELFN1 intracellular and extracellular domains on membrane trafficking, synaptic localization, and dimerization[J]. J Biol Chem, 2025, 301(1): 108073. doi:10.1016/j.jbc.2024.108073 |
| [26] | Ayhan S, Dursun A. ELFN1 is a new extracellular matrix (ECM)-associated protein[J]. Life Sci, 2024, 352: 122900. doi:10.1016/j.lfs.2024.122900 |
| [27] | Wishart AL, Conner SJ, Guarin JR, et al. Decellularized extracellular matrix scaffolds identify full-length collagen VI as a driver of breast cancer cell invasion in obesity and metastasis[J]. Sci Adv, 2020, 6(43): eabc3175. doi:10.1126/sciadv.abc3175 |
| [28] | Dunn HA, Orlandi C, Martemyanov KA. Beyond the ligand: extracellular and transcellular G protein-coupled receptor complexes in physiology and pharmacology[J]. Pharmacol Rev, 2019, 71(4): 503-19. doi:10.1124/pr.119.018044 |
| [29] | Kumar US, Natarajan A, Massoud TF, et al. FN3 linked nanobubbles as a targeted contrast agent for US imaging of cancer-associated human PD-L1[J]. J Control Release, 2022, 346: 317-27. doi:10.1016/j.jconrel.2022.04.030 |
| [30] | Winkler J, Abisoye-Ogunniyan A, Metcalf KJ, et al. Concepts of extracellular matrix remodelling in tumour progression and metastasis[J]. Nat Commun, 2020, 11(1): 5120. doi:10.1038/s41467-020-18794-x |
| [31] | Glaviano A, Foo ASC, Lam HY, et al. PI3K/AKT/mTOR signaling transduction pathway and targeted therapies in cancer[J]. Mol Cancer, 2023, 22(1): 138. doi:10.1186/s12943-023-01827-6 |
| [32] | Deng ZQ, Fan T, Xiao C, et al. TGF-β signaling in health, disease and therapeutics[J]. Sig Transduct Target Ther, 2024, 9: 61. doi:10.1038/s41392-024-01764-w |
| [33] | Rigillo G, Belluti S, Campani V, et al. The NF-Y splicing signature controls hybrid EMT and ECM-related pathways to promote aggressiveness of colon cancer[J]. Cancer Lett, 2023, 567: 216262. doi:10.1016/j.canlet.2023.216262 |
| [34] | Strating E, Verhagen MP, Wensink E, et al. Co-cultures of colon cancer cells and cancer-associated fibroblasts recapitulate the aggressive features of mesenchymal-like colon cancer[J]. Front Immunol, 2023, 14: 1053920. doi:10.3389/fimmu.2023.1053920 |
| [1] | Xinyuan CHEN, Chengting WU, Ruidi LI, Xueqin PAN, Yaodan ZHANG, Junyu TAO, Caizhi LIN. Shuangshu Decoction inhibits growth of gastric cancer cell xenografts by promoting cell ferroptosis via the P53/SLC7A11/GPX4 axis [J]. Journal of Southern Medical University, 2025, 45(7): 1363-1371. |
| [2] | Nuozhou WENG, Bin TAN, Wentao ZENG, Jiayu GU, Lianji WENG, Kehong ZHENG. RGL1 overexpression promotes metastasis of colorectal cancer by upregulating motile focal adhesion assembly via activating the CDC42/RAC1 complex [J]. Journal of Southern Medical University, 2025, 45(5): 1031-1038. |
| [3] | Zhennan MA, Fuquan LIU, Xuefeng ZHAO, Xiaowei ZHANG. High expression of DTX2 promotes proliferation, invasion and epithelial-mesenchymal transition of oxaliplatin-resistant colorectal cancer cells [J]. Journal of Southern Medical University, 2025, 45(4): 829-836. |
| [4] | Zhi GAO, Ao WU, Zhongxiang HU, Peiyang SUN. Bioinformatics analysis of oxidative stress and immune infiltration in rheumatoid arthritis [J]. Journal of Southern Medical University, 2025, 45(4): 862-870. |
| [5] | Shunjie QING, Zhiyong SHEN. High expression of hexokinase 2 promotes proliferation, migration and invasion of colorectal cancer cells by activating the JAK/STAT pathway and regulating tumor immune microenvironment [J]. Journal of Southern Medical University, 2025, 45(3): 542-553. |
| [6] | Huaiwen XU, Li WENG, Hong XUE. CXCL12 is a potential therapeutic target for type 2 diabetes mellitus complicated by chronic obstructive pulmonary disease [J]. Journal of Southern Medical University, 2025, 45(1): 100-109. |
| [7] | Mengnan YE, Hongmei WU, Yan MEI, Qingling ZHANG. High expression of CREM is associated with poor prognosis in gastric cancer patients [J]. Journal of Southern Medical University, 2024, 44(9): 1776-1782. |
| [8] | Kai JI, Guanyu YU, Leqi ZHOU, Tianshuai ZHANG, Qianlong LING, Wenjiang MAN, Bing ZHU, Wei ZHANG. HNRNPA1 gene is highly expressed in colorectal cancer: its prognostic implications and potential as a therapeutic target [J]. Journal of Southern Medical University, 2024, 44(9): 1685-1695. |
| [9] | Yinliang ZHANG, Zetan LUO, Rui ZHAO, Na ZHAO, Zhidong XU, Di AO, Guyi CONG, Xinyu LIU, Hailun ZHENG. Sanguinarine induces ferroptosis of colorectal cancer cells by upregulating STUB1 and downregulating GPX4 [J]. Journal of Southern Medical University, 2024, 44(8): 1537-1544. |
| [10] | Hongli YANG, Yayun XIANG, Tingting TAN, Yang LEI. ORY-1001 inhibits glioblastoma cell growth by downregulating the Notch/HES1 pathway via suppressing lysine-specific demethylase 1 expression [J]. Journal of Southern Medical University, 2024, 44(8): 1620-1630. |
| [11] | Heping LI, Gaohua LI, Xuehua ZHANG, Yanan WANG. Genetic drivers for inflammatory protein markers in colorectal cancer: a Mendelian randomization approach to clinical prognosis study [J]. Journal of Southern Medical University, 2024, 44(7): 1361-1370. |
| [12] | Wei ZHOU, Jun NIE, Jia HU, Yizhi JIANG, Dafa ZHANG. Differential expressions of endoplasmic reticulum stress-associated genes in aortic dissection and their correlation with immune cell infiltration [J]. Journal of Southern Medical University, 2024, 44(5): 859-866. |
| [13] | Nan WANG, Bin SHI, Xiaolan MAN, Weichao WU, Jia CAO. High expression of fragile X mental retardation protein inhibits ferroptosis of colorectal tumor cells by activating the RAS/MAPK signaling pathway [J]. Journal of Southern Medical University, 2024, 44(5): 885-893. |
| [14] | Bei PEI, Yi ZHANG, Siyuan WEI, Yu MEI, Biao SONG, Gang DONG, Ziang WEN, Xuejun LI. Identification of potential pathogenic genes of intestinal metaplasia based on transcriptomic sequencing and bioinformatics analysis [J]. Journal of Southern Medical University, 2024, 44(5): 941-949. |
| [15] | Qinzhi WANG, Bing SONG, Shirui HAO, Zhiyuan XIAO, Lianhui JIN, Tong ZHENG, Fang CHAI. Bioinformatic analysis of CCND2 expression in papillary thyroid carcinoma and its impact on immune infiltration [J]. Journal of Southern Medical University, 2024, 44(5): 981-988. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||