Journal of Southern Medical University ›› 2025, Vol. 45 ›› Issue (7): 1479-1489.doi: 10.12122/j.issn.1673-4254.2025.07.14
Previous Articles Next Articles
Jinlong PANG1,2( ), Xinli ZHAO1,2(
), Xinli ZHAO1,2( ), Zhen ZHANG1,2, Haojie WANG1,2, Xingqi ZHOU1,2, Yumei YANG1,2, Shanshan LI1,2, Xiaoqiang CHANG1,2, Feng LI1,2(
), Zhen ZHANG1,2, Haojie WANG1,2, Xingqi ZHOU1,2, Yumei YANG1,2, Shanshan LI1,2, Xiaoqiang CHANG1,2, Feng LI1,2( ), Xian LI1,2(
), Xian LI1,2( )
)
Received:2025-03-06
Online:2025-07-20
Published:2025-07-17
Contact:
Feng LI, Xian LI
E-mail:834752819@qq.com;1850466372@qq.com;1583635955@qq.com;Lixian0813@126.com
Jinlong PANG, Xinli ZHAO, Zhen ZHANG, Haojie WANG, Xingqi ZHOU, Yumei YANG, Shanshan LI, Xiaoqiang CHANG, Feng LI, Xian LI. Overexpression of multimerin-2 promotes cutaneous melanoma cell invasion and migration and is associated with poor prognosis[J]. Journal of Southern Medical University, 2025, 45(7): 1479-1489.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.j-smu.com/EN/10.12122/j.issn.1673-4254.2025.07.14
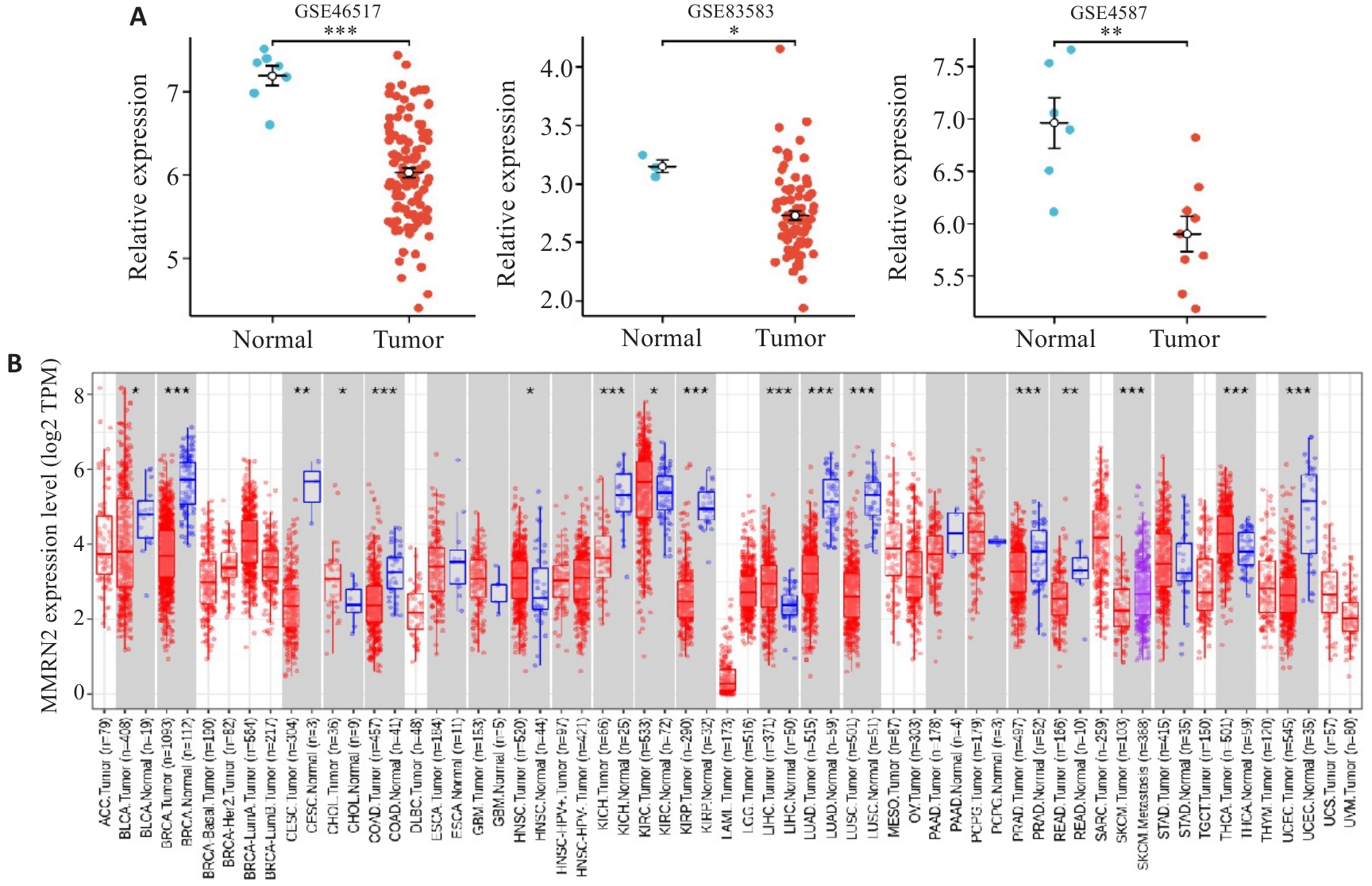
Fig.1 MMRN2 expression in normal and cutaneous melanoma tissues. A: Differential gene expression analysis between tumor and normal tissues using data from GSE46517, GSE83583, and GSE4587 datasets. B: MMRN2 expression levels are significantly higher in metastatic cutaneous melanoma than in normal tissues. *P<0.05, **P<0.01 vs normal tissue; ***P<0.001 vs cutaneous melanoma.
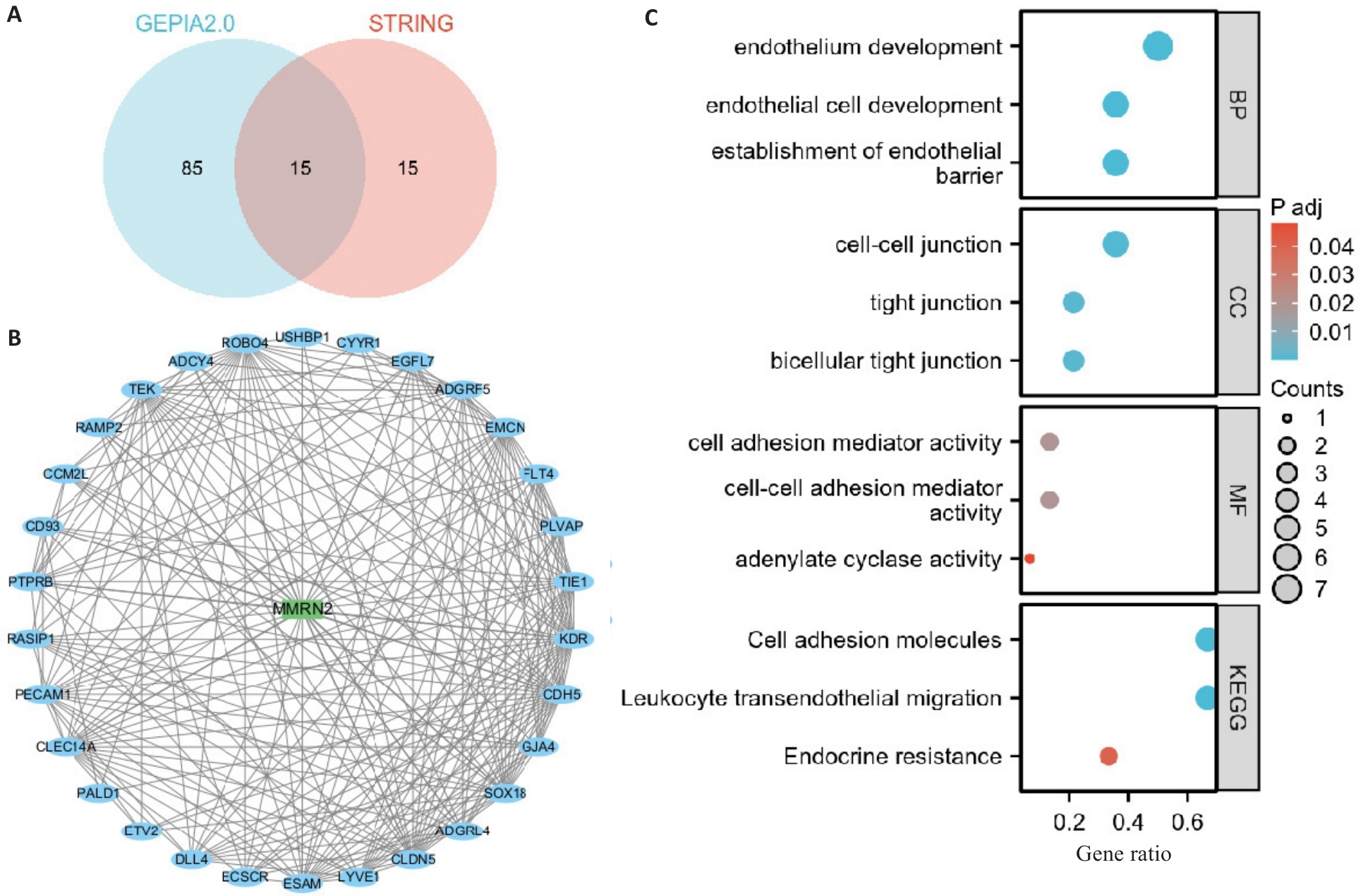
Fig.2 Cutaneous melanoma-related genes and the protein-protein interaction network. A: Intersection analysis of genes associated with cutaneous melanoma using the GEPIA2.0 database. B: Protein-protein interaction (PPI) network constructed by retrieving 30 interacting molecules from the STRING database, resulting in 15 hub genes shared across analyses. C: Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses of the hub genes.
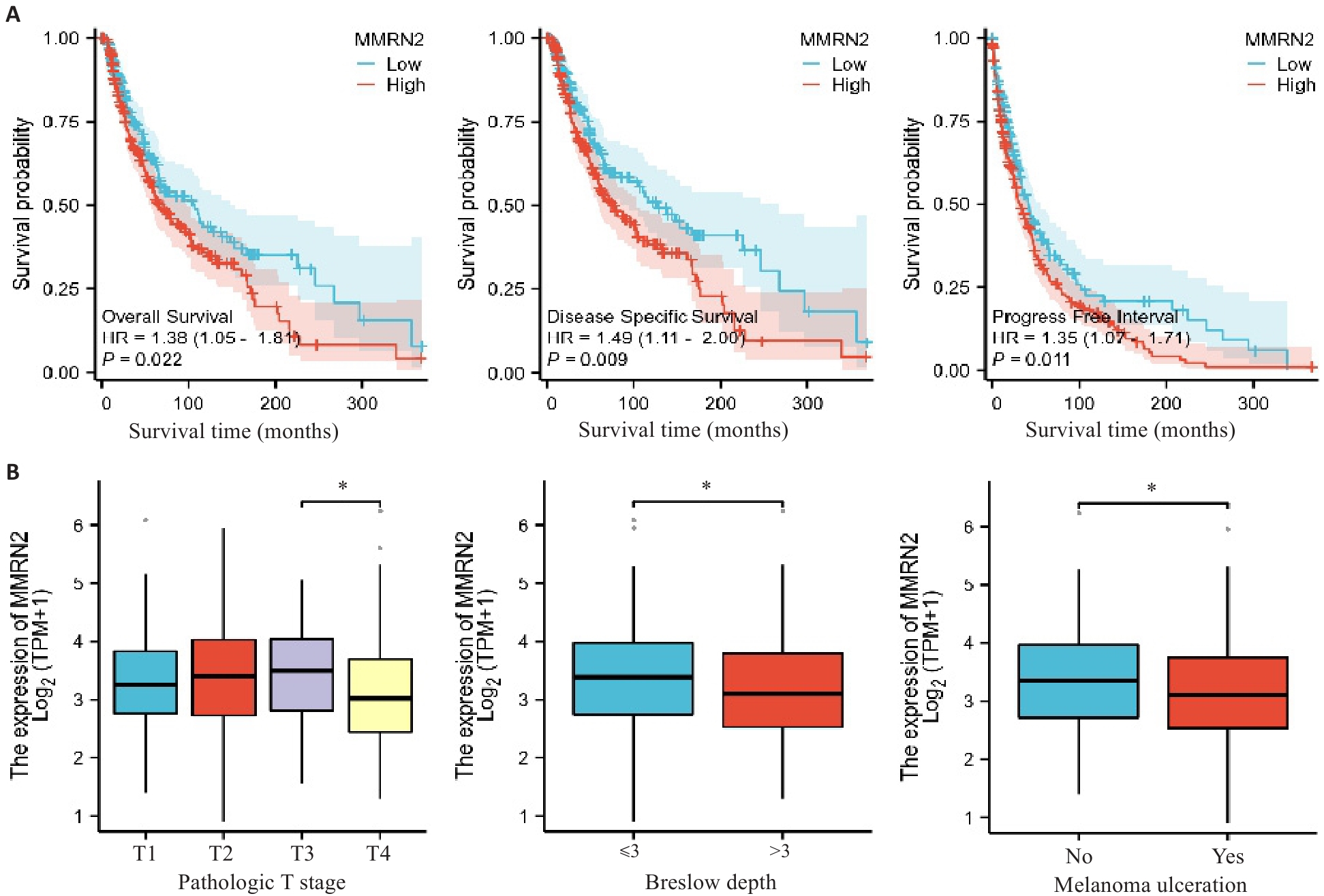
Fig.3 Association of MMRN2 expression levels with clinicopathological features of patients with cutaneous melanoma. A: Kaplan-Meier survival analysis of patients with cutaneous melanoma stratified by MMRN2 expression level. B: Correlations of MMRN2 expression level with pathological T stage, Breslow depth and melanoma ulceration in patients with cutaneous melanoma. *P<0.05.
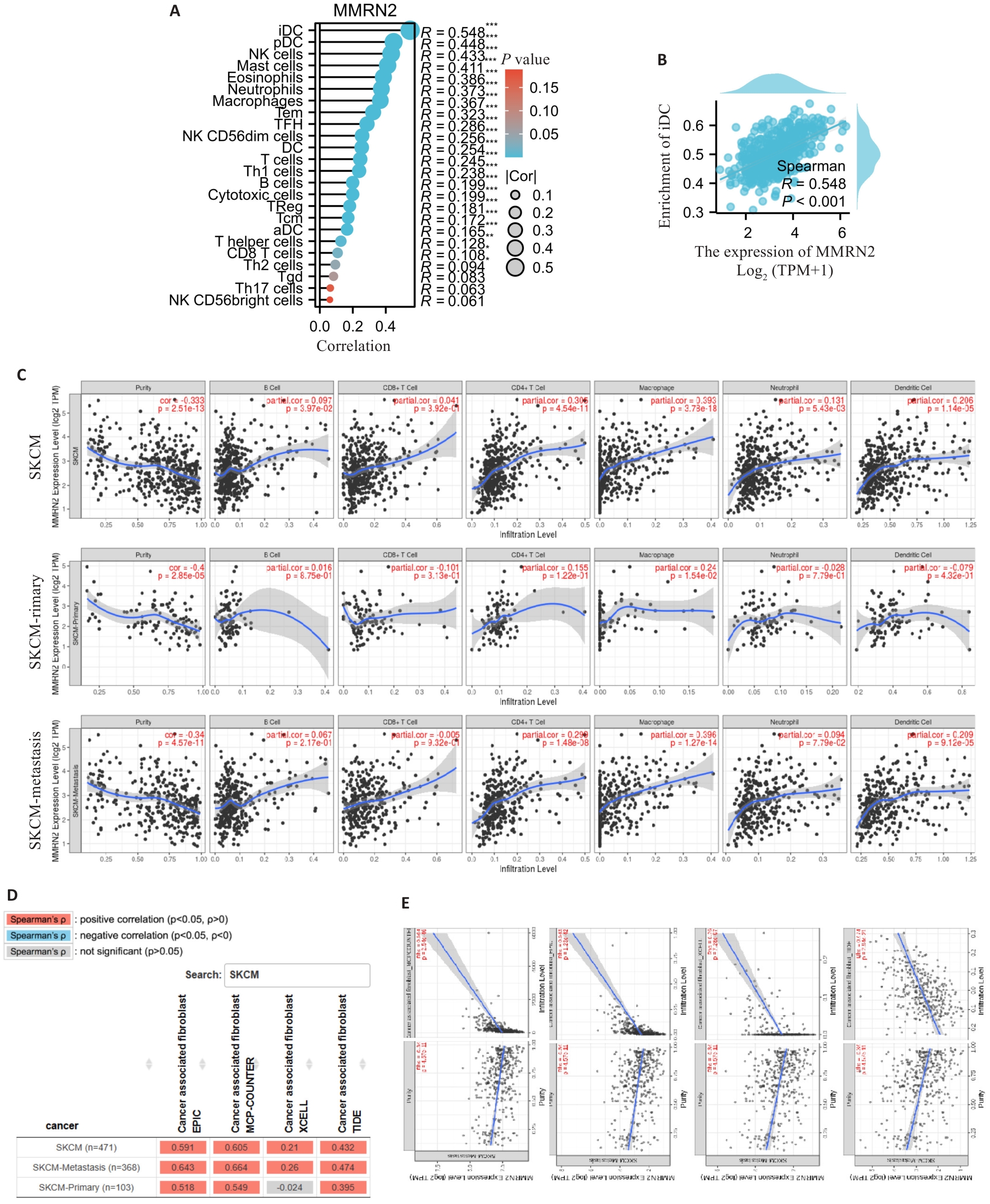
Fig.5 Immune infiltration analysis of MMRN2 in cutaneous melanoma. A: Correlation lollipop chart between MMRN2 and immune infiltration matrix data in TCGA database. The results are visualized using the ggplot2 package. The 24 types of immune cells included aDC (activated DC), B cells, CD8 T cells, cytotoxic cells, DC, eosinophils, iDC (immature DC), macrophages, mast cells, neutrophils, NK CD56bright cells, NK CD56dim cells, NK cells, pDC (plasmacytoid DC), T cells, T helper cells, Tcm (T central memory), Tem (T effector memory), TFH (T follicular helper), Tgd (T gamma delta), Th1 cells, Th17 cells, Th2 cells, and TReg. B: Scatter plot of the correlation between MMRN2 expression level and iDC abundance. C: Scatter plot of correlations between MMRN2 and immune cells in cutaneous melanoma. D, E: Scatter plots of correlations between MMRN2 and tumor fibroblasts in cutaneous melanoma. F: Scatter plot of correlations between MMRN2 and endothelial cells in cutaneous melanoma. Data source: TIMER database, http://timer.comp-genomics.org/timer/
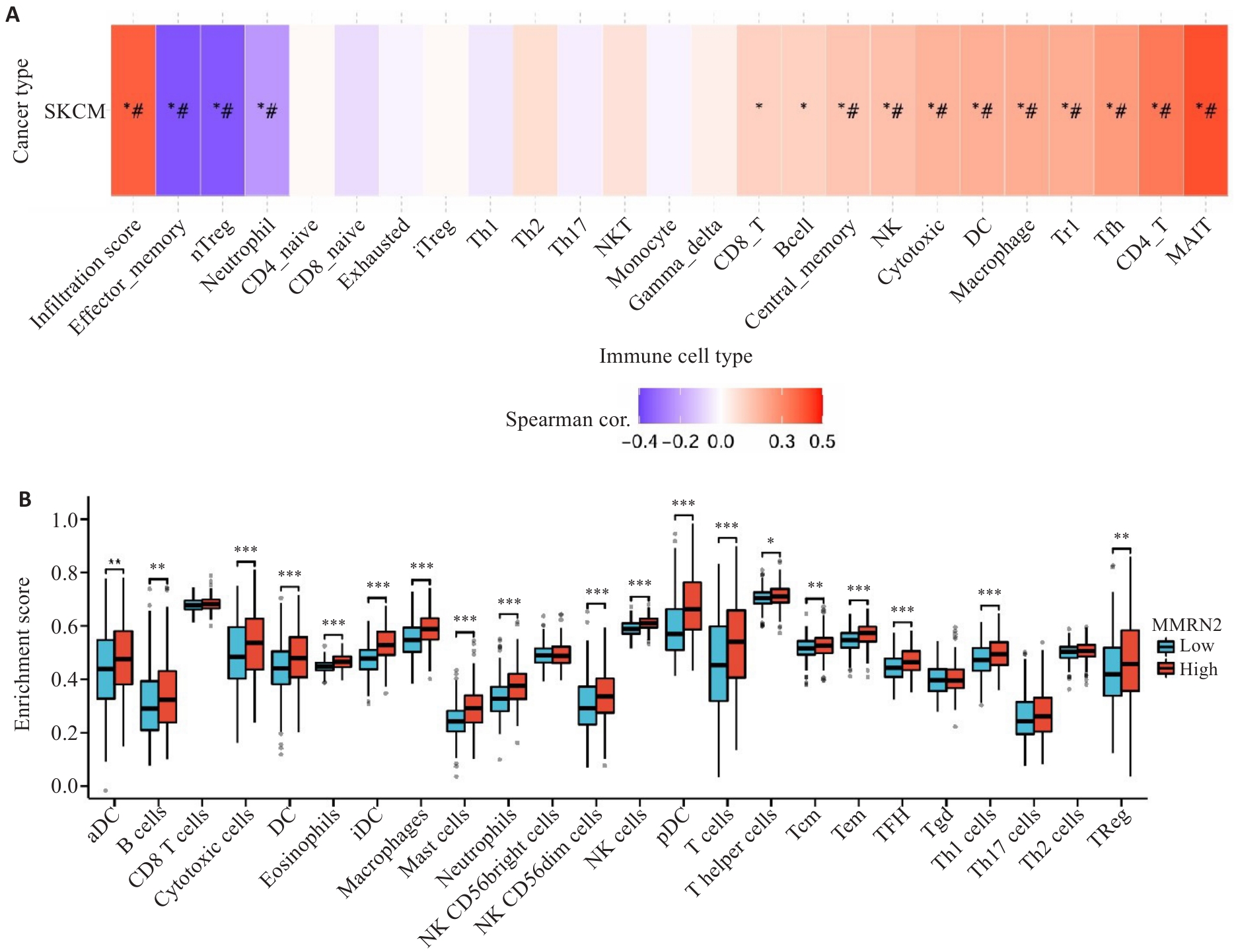
Fig.6 Immune infiltration analysis of MMRN2 in cutaneous melanoma. A: GSVA-based immune cell enrichment scores of MMRN2 in cutaneous melanoma. B: Immune infiltration profiles between high and low MMRN2 expression groups. #: FDR<0.05, *P<0.05, **P<0.01, ***P<0.001 vs low MMRN2 expression group.
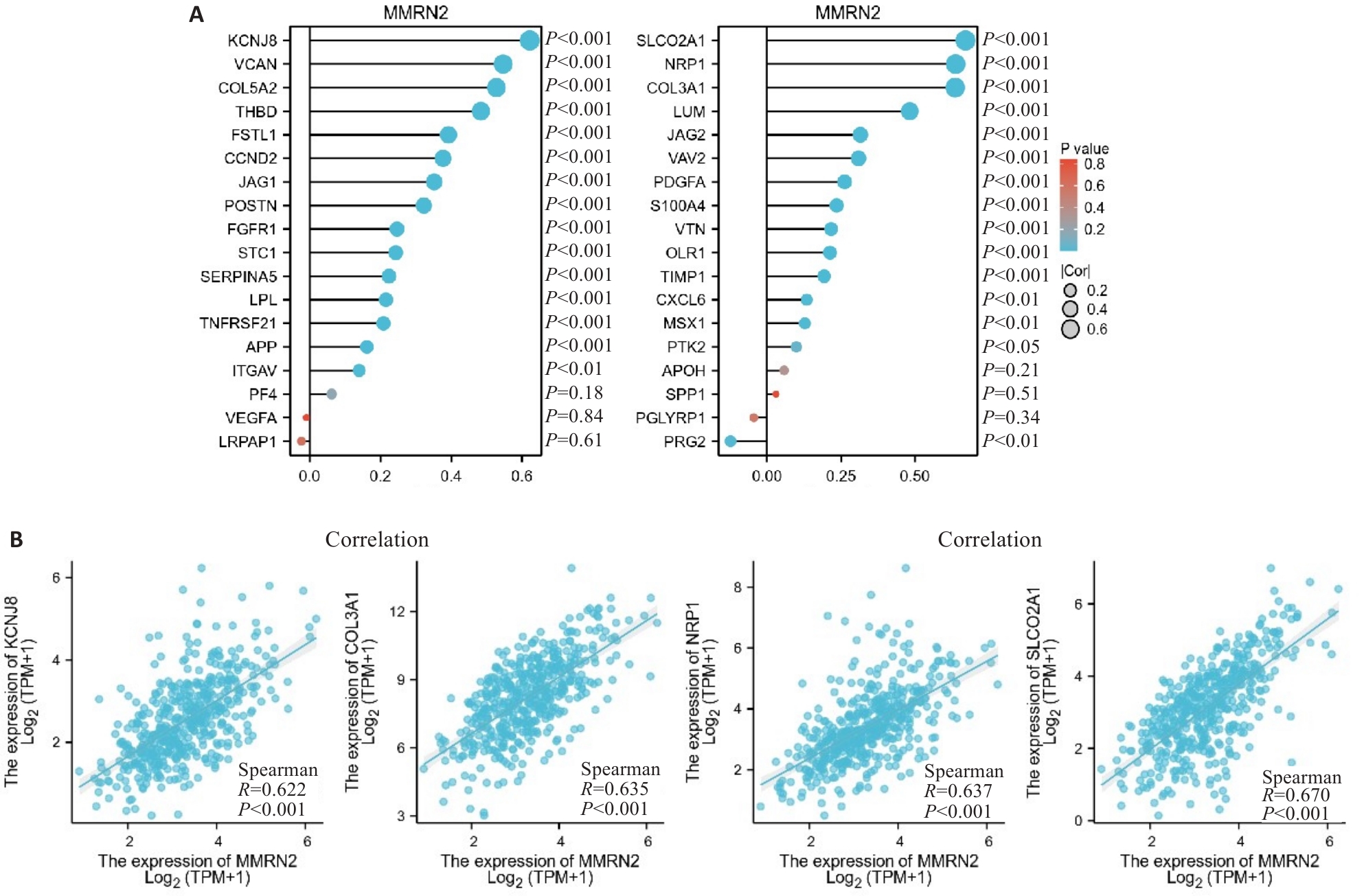
Fig.7 Correlation analysis of MMRN2 and angiogenesis-related genes. A: MMRN2 expression is positively correlated with the majority of angiogenesis-associated genes. B: MMRN2 expression level is strongly correlated with KCNJ8, SLCO2A1, NRP1 and COL3A1.
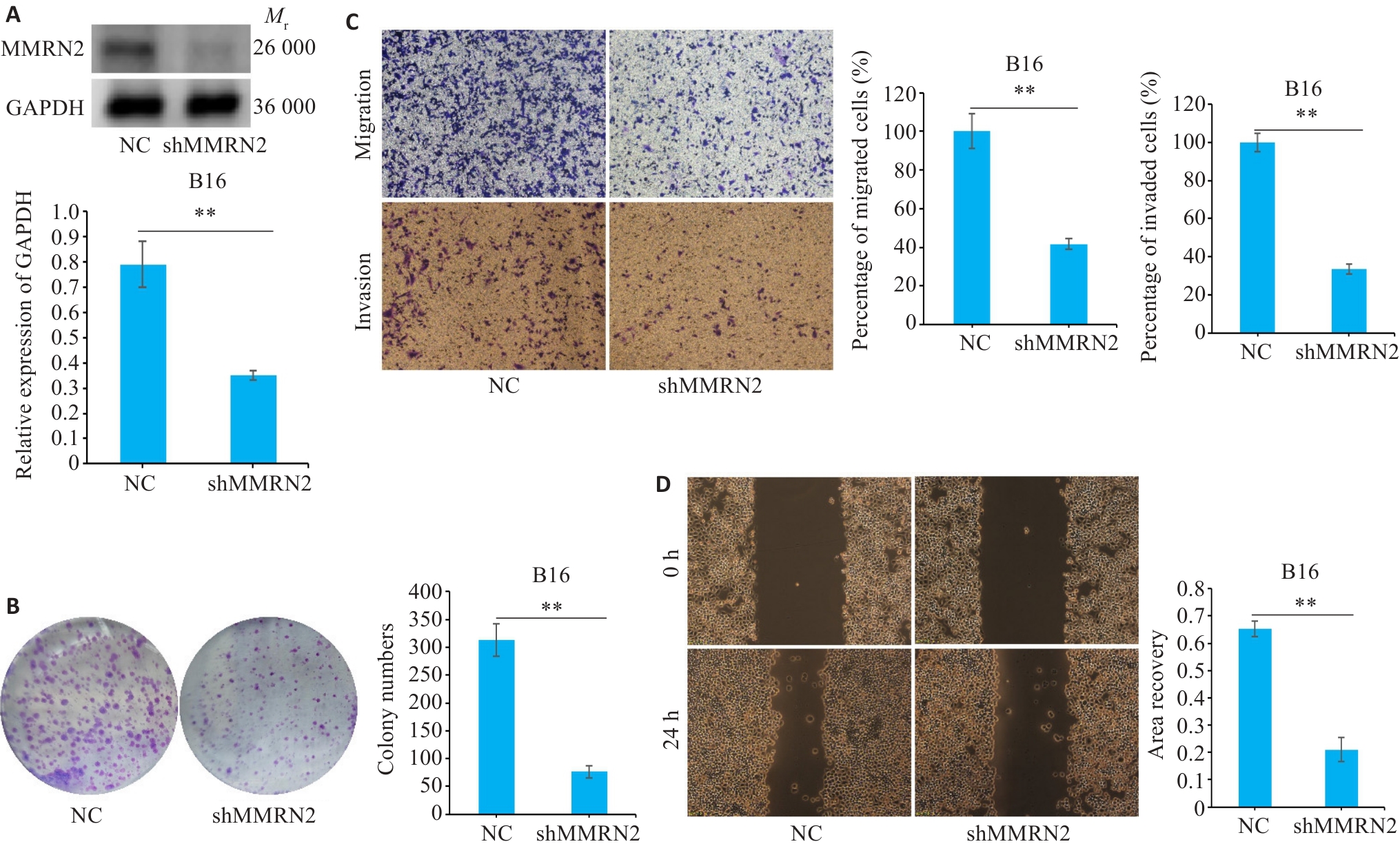
Fig.8 Knockdown of MMRN2 suppresses proliferation, invasion, and migration of cutaneous melanoma cells. A: Western blotting of MMRN2 proteins. B: Colony formation assay for assessing cell proliferation. C: Transwell assay for assessing cell migration and invasion (Original magnification: ×10). D: Knockdown of MMRN2 impairs wound healing in B16 melanoma cells (×200). **P<0.01.
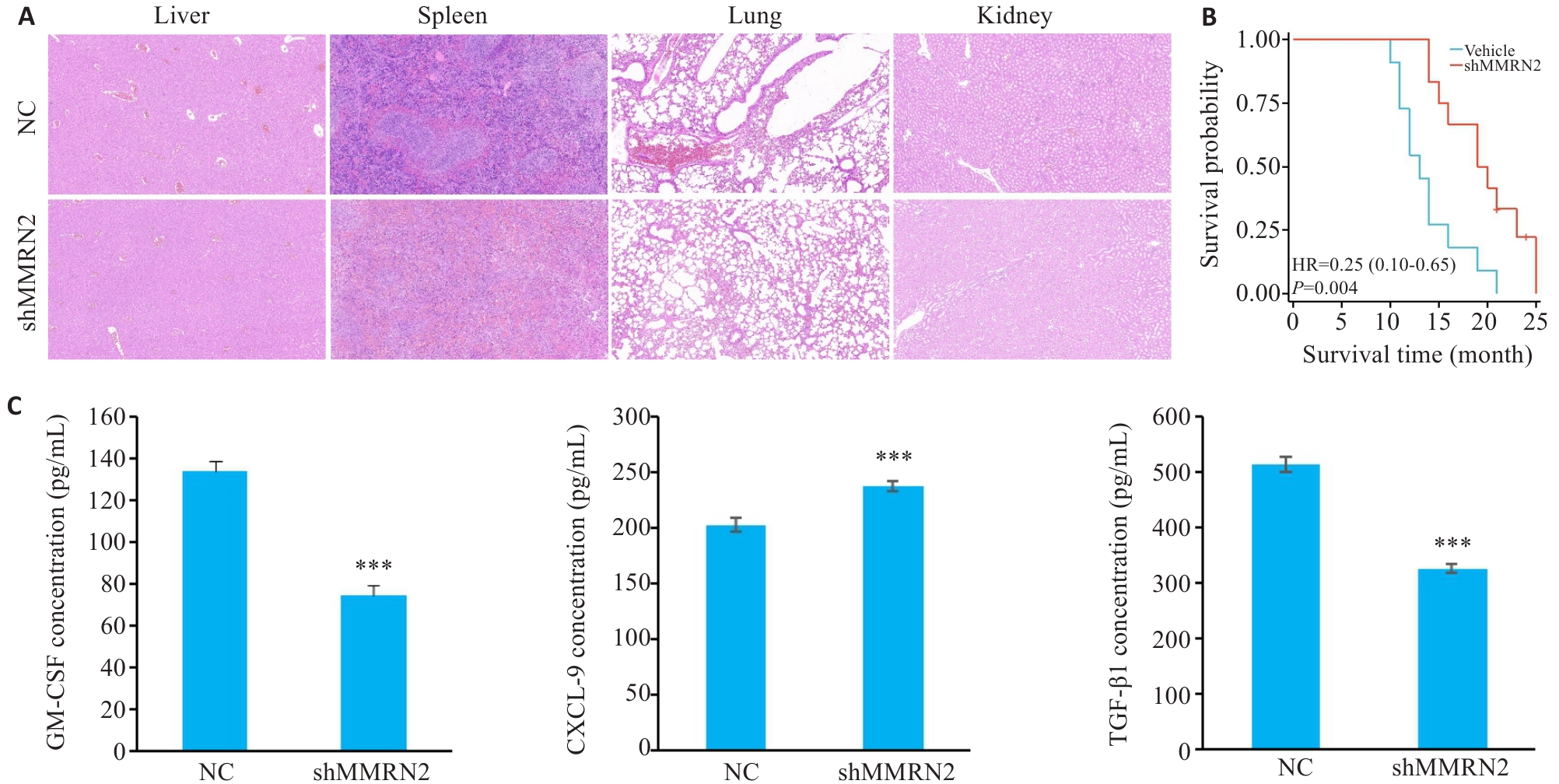
Fig.9 Knockdown of MMRN2 attenuates pathological manifestations and extends survival of mice bearing cutaneous melanoma xenograft. A: HE staining of the liver, spleen, lung, and kidney of mice in MMRN2 knockdown group (×10). B: Kaplan-Meier curve. C: GM-CSF, CXCL9, and TGF-β1 expression levels measured by ELISA. ***P<0.001 vs NC group.
| [1] | Gosman LM, \u 021 aăpoi DA, Costache M. Cutaneous melanoma: a review of multifactorial pathogenesis, immunohistochemistry, and emerging biomarkers for early detection and management[J]. Int J Mol Sci, 2023, 24(21): 15881. doi:10.3390/ijms242115881 |
| [2] | Leonardi GC, Falzone L, Salemi R, et al. Cutaneous melanoma: from pathogenesis to therapy (review)[J]. Int J Oncol, 2018, 52(4): 1071-80. |
| [3] | Davis LE, Shalin SC, Tackett AJ. Current state of melanoma diagnosis and treatment[J]. Cancer Biol Ther, 2019, 20(11): 1366-79. doi:10.1080/15384047.2019.1640032 |
| [4] | Andreuzzi E, Colladel R, Pellicani R, et al. The angiostatic molecule Multimerin 2 is processed by MMP-9 to allow sprouting angiogenesis[J]. Matrix Biol, 2017, 64: 40-53. doi:10.1016/j.matbio.2017.04.002 |
| [5] | Khan KA, Naylor AJ, Khan A, et al. Multimerin-2 is a ligand for group 14 family C-type lectins CLEC14A, CD93 and CD248 spanning the endothelial pericyte interface[J]. Oncogene, 2017, 36(44): 6097-108. doi:10.1038/onc.2017.214 |
| [6] | Lorenzon E, Colladel R, Andreuzzi E, et al. MULTIMERIN2 impairs tumor angiogenesis and growth by interfering with VEGF-A/VEGFR2 pathway[J]. Oncogene, 2012, 31(26): 3136-47. doi:10.1038/onc.2011.487 |
| [7] | Galvagni F, Nardi F, Spiga O, et al. Dissecting the CD93-Multimerin 2 interaction involved in cell adhesion and migration of the activated endothelium[J]. Matrix Biol, 2017, 64: 112-27. doi:10.1016/j.matbio.2017.08.003 |
| [8] | Chen CJ, Kajita H, Aramaki-Hattori N, et al. Screening of autophagy-related prognostic genes in metastatic skin melanoma[J]. Dis Markers, 2022, 2022: 8556593. doi:10.1155/2022/8556593 |
| [9] | Liu JF, Lichtenberg T, Hoadley KA, et al. An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics[J]. Cell, 2018, 173(2): 400-16.e11. |
| [10] | Bindea G, Mlecnik B, Tosolini M, et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer[J]. Immunity, 2013, 39(4): 782-95. doi:10.1016/j.immuni.2013.10.003 |
| [11] | Qing X, Xu WJ, Liu SL, et al. Molecular characteristics, clinical significance, and cancer immune interactions of angiogenesis-associated genes in gastric cancer[J]. Front Immunol, 2022, 13: 843077. doi:10.3389/fimmu.2022.843077 |
| [12] | Yin QL, Shi XM, Lan SJ, et al. Effect of melanoma stem cells on melanoma metastasis[J]. Oncol Lett, 2021, 22(1): 566. doi:10.3892/ol.2021.12827 |
| [13] | Holderfield M, Deuker MM, McCormick F, et al. Targeting RAF kinases for cancer therapy: BRAF-mutated melanoma and beyond[J]. Nat Rev Cancer, 2014, 14(7): 455-67. doi:10.1038/nrc3760 |
| [14] | Cicaloni V, Karmakar M, Frusciante L, et al. Bioinformatics approaches to predict mutation effects in the binding site of the proangiogenic molecule CD93[J]. Front Bioinform, 2022, 2: 891553. doi:10.3389/fbinf.2022.891553 |
| [15] | Pellicani R, Poletto E, Andreuzzi E, et al. Multimerin-2 maintains vascular stability and permeability[J]. Matrix Biol, 2020, 87: 11-25. doi:10.1016/j.matbio.2019.08.002 |
| [16] | Wang DY, Xing CY, Liang Y, et al. Ultrasound imaging of tumor vascular CD93 with MMRN2 modified microbubbles for immune microenvironment prediction[J]. Adv Mater, 2024, 36(18): e2310421. doi:10.1002/adma.202470134 |
| [17] | Liu H, Zhang JH, Zhao YJ, et al. CD93 regulates breast cancer growth and vasculogenic mimicry through the PI3K/AKT/SP2 signaling pathway activated by integrin β1[J]. J Biochem Mol Toxicol, 2024, 38(4): e23688. doi:10.1002/jbt.23688 |
| [18] | Li Y, Fu L, Wu BK, et al. Angiogenesis modulated by CD93 and its natural ligands IGFBP7 and MMRN2: a new target to facilitate solid tumor therapy by vasculature normalization[J]. Cancer Cell Int, 2023, 23(1): 189. doi:10.1186/s12935-023-03044-z |
| [19] | Cao Z, Zhang ZJ, Tang XY, et al. Comprehensive analysis of tissue proteomics in patients with papillary thyroid microcarcinoma uncovers the underlying mechanism of lymph node metastasis and its significant sex disparities[J]. Front Oncol, 2022, 12: 887977. doi:10.3389/fonc.2022.887977 |
| [20] | Zheng WZ, Zhang SQ, Guo H, et al. Multi-omics analysis of tumor angiogenesis characteristics and potential epigenetic regulation mechanisms in renal clear cell carcinoma[J]. Cell Commun Signal, 2021, 19(1): 39. doi:10.1186/s12964-021-00728-9 |
| [21] | Zhao YH, Zhang X, Yao JC, et al. Expression patterns and the prognostic value of the EMILIN/Multimerin family members in low-grade glioma[J]. PeerJ, 2020, 8: e8696. doi:10.7717/peerj.8696 |
| [22] | Zheng Y, Fang MJ, Sanan S, et al. Investigating angiogenesis-related biomarkers in osteoarthritis patients through transcriptomic profiling[J]. J Inflamm Res, 2024, 17: 10681-97. doi:10.2147/jir.s493889 |
| [23] | Simiczyjew A, Dratkiewicz E, Mazurkiewicz J, et al. The influence of tumor microenvironment on immune escape of melanoma[J]. Int J Mol Sci, 2020, 21(21): 8359. doi:10.3390/ijms21218359 |
| [24] | Tordesillas L, Berin MC. Mechanisms of oral tolerance[J]. Clin Rev Allergy Immunol, 2018, 55(2): 107-17. doi:10.1007/s12016-018-8680-5 |
| [25] | Marzagalli M, Ebelt ND, Manuel ER. Unraveling the crosstalk between melanoma and immune cells in the tumor microenvironment[J]. Semin Cancer Biol, 2019, 59: 236-50. doi:10.1016/j.semcancer.2019.08.002 |
| [26] | Ellis LM, Hicklin DJ. VEGF-targeted therapy: mechanisms of anti-tumour activity[J]. Nat Rev Cancer, 2008, 8(8): 579-91. doi:10.1038/nrc2403 |
| [27] | Thomas DA, Massagué J. TGF-beta directly targets cytotoxic T cell functions during tumor evasion of immune surveillance[J]. Cancer Cell, 2005, 8(5): 369-80. doi:10.1016/j.ccr.2005.10.012 |
| [28] | Seeger P, Musso T, Sozzani S. The TGF-β superfamily in dendritic cell biology[J]. Cytokine Growth Factor Rev, 2015, 26(6): 647-57. doi:10.1016/j.cytogfr.2015.06.002 |
| [29] | Wang X, Cui ZD, Chen X, et al. The CXCR4/miR-1910-5p/MMRN2 axis is involved in corneal neovascularization by affecting vascular permeability[J]. Invest Ophthalmol Vis Sci, 2023, 64(4): 10. doi:10.1167/iovs.64.4.10 |
| [30] | Janka EA, Ványai B, Dajnoki Z, et al. Regional variability of melanoma incidence and prevalence in Hungary. Epidemiological impact of ambient UV radiation and socioeconomic factors[J]. Eur J Cancer Prev, 2022, 31(4): 377-84. doi:10.1097/cej.0000000000000716 |
| [1] | Xuan WU, Jiamin FANG, Weiwei HAN, Lin CHEN, Jing SUN, Qili JIN. High PRELID1 expression promotes epithelial-mesenchymal transition in gastric cancer cells and is associated with poor prognosis [J]. Journal of Southern Medical University, 2025, 45(7): 1535-1542. |
| [2] | Kang WANG, Haibin LI, Jing YU, Yuan MENG, Hongli ZHANG. High expression of ELFN1 is a prognostic biomarker and promotes proliferation and metastasis of colorectal cancer cells [J]. Journal of Southern Medical University, 2025, 45(7): 1543-1553. |
| [3] | Huanyu JI, Rui WANG, Shengxiang GAO, Wengang CHE. SG-UNet: a melanoma segmentation model enhanced with global attention and self-calibrated convolution [J]. Journal of Southern Medical University, 2025, 45(6): 1317-1326. |
| [4] | Xiaojuan GUO, Ruijuan DU, Liping CHEN, Kelei GUO, Biao ZHOU, Hua BIAN, Li HAN. WW domain-containing ubiquitin E3 ligase 1 regulates immune infiltration in tumor microenvironment of ovarian cancer [J]. Journal of Southern Medical University, 2025, 45(5): 1063-1073. |
| [5] | Yi ZHANG, Yu SHEN, Zhiqiang WAN, Song TAO, Yakui LIU, Shuanhu WANG. High expression of CDKN3 promotes migration and invasion of gastric cancer cells by regulating the p53/NF-κB signaling pathway and inhibiting cell apoptosis [J]. Journal of Southern Medical University, 2025, 45(4): 853-861. |
| [6] | Zhi GAO, Ao WU, Zhongxiang HU, Peiyang SUN. Bioinformatics analysis of oxidative stress and immune infiltration in rheumatoid arthritis [J]. Journal of Southern Medical University, 2025, 45(4): 862-870. |
| [7] | Qingqing HUANG, Wenjing ZHANG, Xiaofeng ZHANG, Lian WANG, Xue SONG, Zhijun GENG, Lugen ZUO, Yueyue WANG, Jing LI, Jianguo HU. High MYO1B expression promotes proliferation, migration and invasion of gastric cancer cells and is associated with poor patient prognosis [J]. Journal of Southern Medical University, 2025, 45(3): 622-631. |
| [8] | Huali LI, Ting SONG, Jiawen LIU, Yongbao LI, Zhaojing JIANG, Wen DOU, Linghong ZHOU. Prognosis-guided optimization of intensity-modulated radiation therapy plans for lung cancer [J]. Journal of Southern Medical University, 2025, 45(3): 643-649. |
| [9] | Xue SONG, Yue CHEN, Min ZHANG, Nuo ZHANG, Lugen ZUO, Jing LI, Zhijun GENG, Xiaofeng ZHANG, Yueyue WANG, Lian WANG, Jianguo HU. GPSM2 is highly expressed in gastric cancer to affect patient prognosis by promoting tumor cell proliferation [J]. Journal of Southern Medical University, 2025, 45(2): 229-238. |
| [10] | Tianwei TANG, Luan LI, Yuanhan CHEN, Li ZHANG, Lixia XU, Zhilian LI, Zhonglin FENG, Huilin ZHANG, Ruifang HUA, Zhiming YE, Xinling LIANG, Ruizhao LI. High serum cystatin C is an independent risk factor for poor renal prognosis in IgA nephropathy [J]. Journal of Southern Medical University, 2025, 45(2): 379-386. |
| [11] | Xiaorui CHEN, Qingzheng WEI, Zongliang ZHANG, Jiangshui YUAN, Weiqing SONG. Overexpression of CHMP2B suppresses proliferation of renal clear cell carcinoma cells [J]. Journal of Southern Medical University, 2025, 45(1): 126-136. |
| [12] | Yaobin WANG, Liuyan CHEN, Yiling LUO, Jiqing SHEN, Sufang ZHOU. Predictive value of NUF2 for prognosis and immunotherapy responses in pan-cancer [J]. Journal of Southern Medical University, 2025, 45(1): 137-149. |
| [13] | Chao ZHOU, Jingjing ZHANG, Qiao TANG, Shuangnan FU, Ning ZHANG, Zhaoyun HE, Jin ZHANG, Tianyi ZHANG, Pengcheng LIU, Man GONG. Value of serum tryptophan in stratified management of 90-day mortality risk in patients with hepatitis B virus-related acute-on-chronic liver failure: a multicenter retrospective study [J]. Journal of Southern Medical University, 2025, 45(1): 59-64. |
| [14] | Xiaohua CHEN, Hui LU, Ziliang WANG, Lian WANG, Yongsheng XIA, Zhijun GENG, Xiaofeng ZHANG, Xue SONG, Yueyue WANG, Jing LI, Jianguo HU, Lugen ZUO. Role of Abelson interactor 2 in progression and prognosis of gastric cancer and its regulatory mechanisms [J]. Journal of Southern Medical University, 2024, 44(9): 1653-1661. |
| [15] | Mengnan YE, Hongmei WU, Yan MEI, Qingling ZHANG. High expression of CREM is associated with poor prognosis in gastric cancer patients [J]. Journal of Southern Medical University, 2024, 44(9): 1776-1782. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||