南方医科大学学报 ›› 2026, Vol. 46 ›› Issue (4): 929-938.doi: 10.12122/j.issn.1673-4254.2026.04.21
• • 上一篇
陈博湧1( ), 汪新怡1, 赵新新1, 宋婷1(
), 汪新怡1, 赵新新1, 宋婷1( ), 李永宝2(
), 李永宝2( )
)
收稿日期:2025-10-22
出版日期:2026-04-20
发布日期:2026-04-24
通讯作者:
宋婷,李永宝
E-mail:q13729335350@smu.edu.cn;tingsong2015@smu.edu.cn;liyb1@sysucc.org.cn
作者简介:陈博湧,在读硕士研究生,E-mail: q13729335350@smu.edu.cn
基金资助:
Boyong CHEN1( ), Xinyi WANG1, Xinxin ZHAO1, Ting SONG1(
), Xinyi WANG1, Xinxin ZHAO1, Ting SONG1( ), Yongbao LI2(
), Yongbao LI2( )
)
Received:2025-10-22
Online:2026-04-20
Published:2026-04-24
Contact:
Ting SONG, Yongbao LI
E-mail:q13729335350@smu.edu.cn;tingsong2015@smu.edu.cn;liyb1@sysucc.org.cn
Supported by:摘要:
目的 探索基于Swin-ResViT网络从动态cine-MR生成高质量治疗前定位MR(sMR),提升实时影像的信噪比和对比度。 方法 提出一种融合Swin Transformer模块的ResViT模型(Swin-ResViT),通过优化瓶颈层结构以提升特征提取效率。回顾性收集2024年2~7月在中山大学肿瘤防治中心接受治疗的17例肝癌患者数据,其中12例肝癌患者的治疗中cine-MR和治疗前定位MR作为训练集,5例患者为测试集。通过量化sMR与参考定位MR的归一化均方根误差(NRMSE)、峰值信噪比(PSNR)、结构相似性指标(SSIM)、运动标记点误差以及模型推理速度,综合评估图像生成质量和模型性能。 结果 生成图像质量方面,Swin-ResViT生成的sMR相较于原始cine-MR,NRMSE、LPIPS分别下降约90%、82%(P<0.001);PSNR、SSIM、CNR分别提升约157%、79%、181%(P<0.001)。结构准确性方面,动态sMR序列中右肝叶肝膈交界处运动标记点的平均定位误差为0.7695±0.7294 mm(P<0.05)。模型推理速度方面,对于224×224像素的单帧图像,在NVIDIA GeForce RTX 2080 Ti GPU上Swin-ResViT的平均处理时间为15.5 ms,对比标准ResViT为41.4 ms,减少了约62%。 结论 Swin-ResViT模型能从cine-MR合成高质量sMR,该方法兼具高效计算与显著图像增强优势,对实时MRgRT具有重要临床意义。
陈博湧, 汪新怡, 赵新新, 宋婷, 李永宝. 基于Swin-ResViT网络的低质量动态cine-MR至高质量定位MR图像实时生成研究[J]. 南方医科大学学报, 2026, 46(4): 929-938.
Boyong CHEN, Xinyi WANG, Xinxin ZHAO, Ting SONG, Yongbao LI. Swin-ResViT network for real-time generation of high-quality localization MR images from low-quality cine-MR[J]. Journal of Southern Medical University, 2026, 46(4): 929-938.
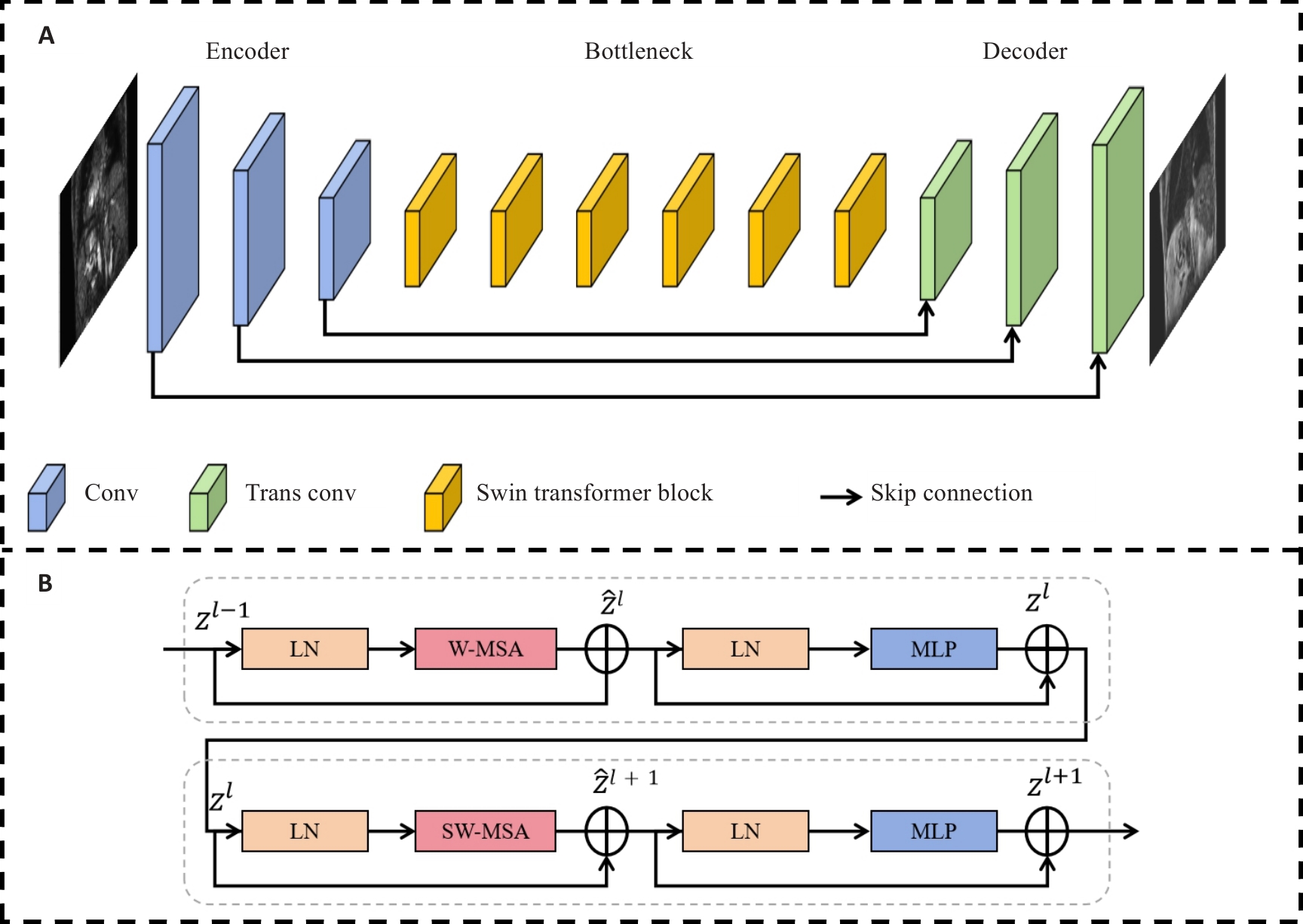
图1 Swin-ResViT的网络结构示意图
Fig.1 Schematic diagram of the structure of the Swin-ResViT network. A: Structure of the network. B: Structure of the Swin Transformer block.
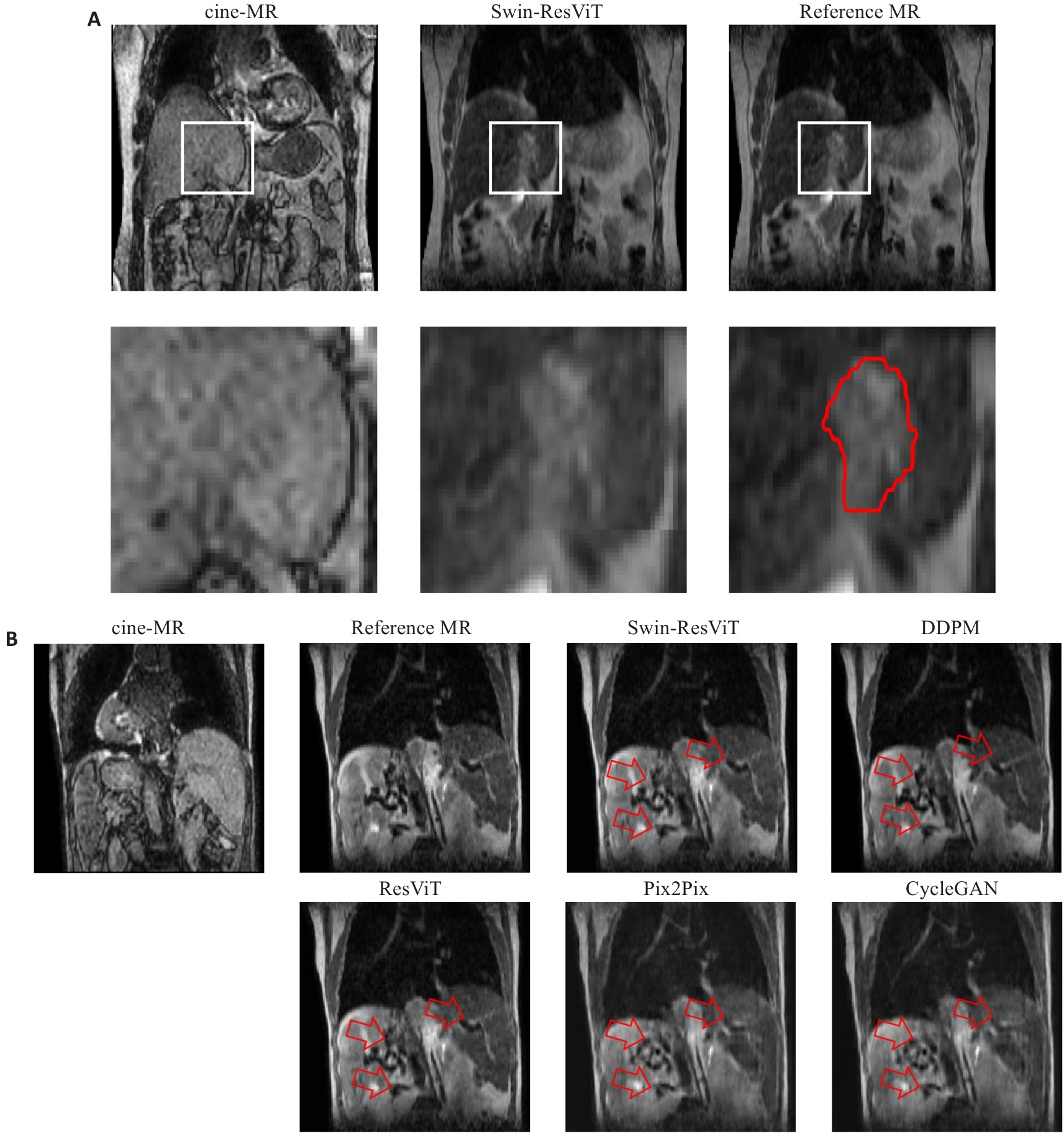
图2 2例肝细胞癌患者的cine-MR、合成sMR和reference MR图像
Fig.2 Cine-MR, synthesized MR (sMR), and localization MR reference slice (reference MR) images from two hepatocellular carcinoma patients. A: Comparison of tumor boundary contrast in images. The white boxes indicate the magnified regions. The red contours outline the gross tumor volume (GTV). B: Comparison of image artifacts. The red arrows highlight regions with anatomical blurring, residual artifacts, or texture distortion in baseline methods, and Swin-ResViT preserved these details more accurately.
| Methods | NRMSE | PSNR | SSIM | CNR | LPIPS | Latency (ms) |
|---|---|---|---|---|---|---|
| cine-MR | 0.1990±0.0305 | 14.2808±1.3035 | 0.5434±0.0436 | 0.3128±0.1939 | 0.3602±0.0076 | - |
| CycleGAN | 0.1335±0.0444 | 18.2640±2.9407 | 0.6918±0.1046 | 0.3322±0.1217 | 0.3471±0.0122 | 15.6 |
| Pix2Pix | 0.1139±0.0239 | 19.6097±2.0893 | 0.7497±0.0372 | 0.3652±0.1428 | 0.2877±0.0109 | 15.9 |
| cDDPM | 0.0350±0.0172 | 29.9420±3.789 | 0.8885±0.0652 | 0.5826±0.1667 | 0.0359±0.0162 | 20026.9 |
| ResViT | 0.0238±0.0172 | 34.4067±3.8455 | 0.9623±0.0308 | 0.8472±0.0513 | 0.0581±0.0154 | 41.4 |
| Swin-ResViT | 0.0199±0.0181 | 36.7590±5.4363 | 0.9737±0.0356 | 0.8795±0.0489 | 0.0656±0.0181 | 15.5 |
表1 sMR、cine-MR与reference MR的图像质量及模型推理速度定量对比
Tab.1 Quantitative comparison of image quality between sMR, cine-MR and reference MR images and latencies of different models (Mean±SD)
| Methods | NRMSE | PSNR | SSIM | CNR | LPIPS | Latency (ms) |
|---|---|---|---|---|---|---|
| cine-MR | 0.1990±0.0305 | 14.2808±1.3035 | 0.5434±0.0436 | 0.3128±0.1939 | 0.3602±0.0076 | - |
| CycleGAN | 0.1335±0.0444 | 18.2640±2.9407 | 0.6918±0.1046 | 0.3322±0.1217 | 0.3471±0.0122 | 15.6 |
| Pix2Pix | 0.1139±0.0239 | 19.6097±2.0893 | 0.7497±0.0372 | 0.3652±0.1428 | 0.2877±0.0109 | 15.9 |
| cDDPM | 0.0350±0.0172 | 29.9420±3.789 | 0.8885±0.0652 | 0.5826±0.1667 | 0.0359±0.0162 | 20026.9 |
| ResViT | 0.0238±0.0172 | 34.4067±3.8455 | 0.9623±0.0308 | 0.8472±0.0513 | 0.0581±0.0154 | 41.4 |
| Swin-ResViT | 0.0199±0.0181 | 36.7590±5.4363 | 0.9737±0.0356 | 0.8795±0.0489 | 0.0656±0.0181 | 15.5 |
| Swin-ResViT | NRMSE | PSNR | SSIM |
|---|---|---|---|
| Fold 1 | 0.0155 | 37.4514 | 0.9865 |
| Fold 2 | 0.0168 | 37.7475 | 0.9840 |
| Fold 3 | 0.0042 | 42.3498 | 0.9985 |
| Fold 4 | 0.0439 | 29.7120 | 0.9274 |
| Average | 0.0201±0.0168 | 36.8151±5.2396 | 0.9741±0.0317 |
表2 Swin-ResViT 4折交叉验证结果
Tab.2 Results of 4-fold cross-validation of Swin-ResViT
| Swin-ResViT | NRMSE | PSNR | SSIM |
|---|---|---|---|
| Fold 1 | 0.0155 | 37.4514 | 0.9865 |
| Fold 2 | 0.0168 | 37.7475 | 0.9840 |
| Fold 3 | 0.0042 | 42.3498 | 0.9985 |
| Fold 4 | 0.0439 | 29.7120 | 0.9274 |
| Average | 0.0201±0.0168 | 36.8151±5.2396 | 0.9741±0.0317 |
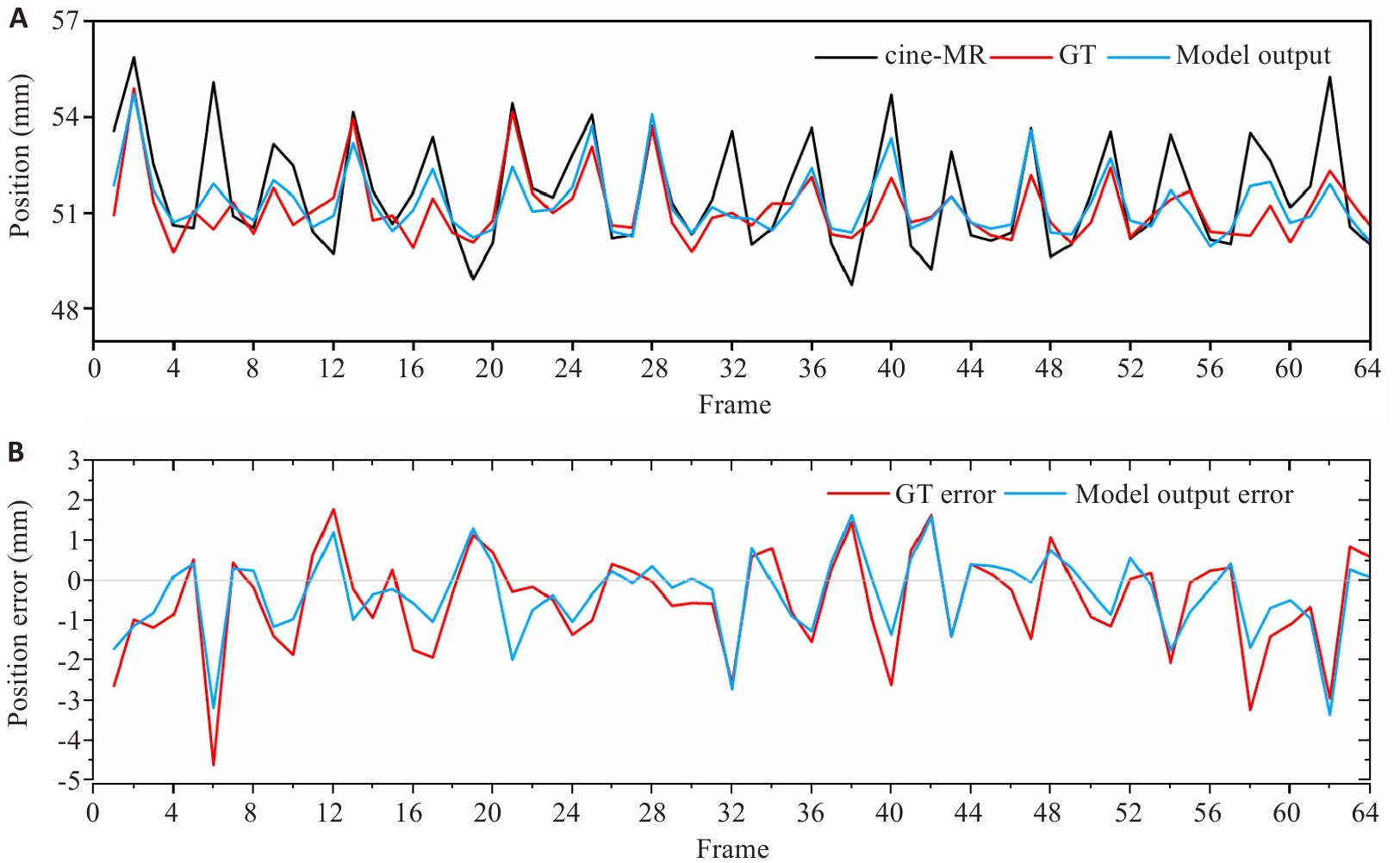
图4 标记点的运动轨迹与误差分析
Fig.4 Motion trajectory and position error of landmarks. A: Motion trajectory of landmarks. B: Position errors of the reference MR and sMR relative to the cine-MR.
| Loss | NRMSE | PSNR | SSIM |
|---|---|---|---|
| MAE | 0.0270±0.0126 | 32.6261±3.8100 | 0.9695±0.0252 |
| MAE+grad loss | 0.0209±0.0135 | 35.5751±5.9679 | 0.9780±0.0244 |
| Proposed loss | 0.0199±0.0181 | 36.7590±5.4363 | 0.9737±0.0356 |
表3 不同损失函数组合下的模型性能对比
Tab.3 Model performance comparison with different loss function combinations (Mean±SD)
| Loss | NRMSE | PSNR | SSIM |
|---|---|---|---|
| MAE | 0.0270±0.0126 | 32.6261±3.8100 | 0.9695±0.0252 |
| MAE+grad loss | 0.0209±0.0135 | 35.5751±5.9679 | 0.9780±0.0244 |
| Proposed loss | 0.0199±0.0181 | 36.7590±5.4363 | 0.9737±0.0356 |
| Skip connection | NRMSE | PSNR | SSIM |
|---|---|---|---|
| 0 | 0.0238±0.0128 | 35.8764±5.8769 | 0.9684±0.0242 |
| 1 | 0.0232±0.0126 | 35.9615±5.7804 | 0.9709±0.0241 |
| 1, 2 | 0.0228±0.0127 | 35.9133±5.8754 | 0.9718±0.0236 |
| 1, 2, 3 | 0.0199±0.0181 | 36.7590±5.4363 | 0.9737±0.0356 |
表4 不同数量跳跃连接的模型性能对比
Tab.4 Model performance comparison with different numbers of skip connections (Mean±SD)
| Skip connection | NRMSE | PSNR | SSIM |
|---|---|---|---|
| 0 | 0.0238±0.0128 | 35.8764±5.8769 | 0.9684±0.0242 |
| 1 | 0.0232±0.0126 | 35.9615±5.7804 | 0.9709±0.0241 |
| 1, 2 | 0.0228±0.0127 | 35.9133±5.8754 | 0.9718±0.0236 |
| 1, 2, 3 | 0.0199±0.0181 | 36.7590±5.4363 | 0.9737±0.0356 |
| Model variant | NRMSE | PSNR | SSIM | Latency (ms) |
|---|---|---|---|---|
| Baseline 1 | 0.0238±0.0128 | 35.8764±5.8769 | 0.9684±0.0242 | 10.7 |
| Baseline 2 | 0.0221±0.0163 | 34.7482±3.8342 | 0.9654±0.0238 | 43.2 |
| Swin-ResViT | 0.0199±0.0181 | 36.7590±5.4363 | 0.9737±0.0356 | 15.5 |
表5 不同网络架构的模型性能对比
Tab.5 Model performance comparison with different network architecture (Mean±SD)
| Model variant | NRMSE | PSNR | SSIM | Latency (ms) |
|---|---|---|---|---|
| Baseline 1 | 0.0238±0.0128 | 35.8764±5.8769 | 0.9684±0.0242 | 10.7 |
| Baseline 2 | 0.0221±0.0163 | 34.7482±3.8342 | 0.9654±0.0238 | 43.2 |
| Swin-ResViT | 0.0199±0.0181 | 36.7590±5.4363 | 0.9737±0.0356 | 15.5 |
| [1] | 田 静, 韩 丹, 周 涛. 肿瘤放射治疗技术的发展及应用研究[J]. 中国医刊, 2022, 57(10): 1064-7. doi:10.3969/j.issn.1008-1070.2022.10.006 |
| [2] | Raaymakers BW, Jürgenliemk-Schulz IM, Bol GH, et al. First patients treated with a 1.5 T MRI-Linac: clinical proof of concept of a high-precision, high-field MRI guided radiotherapy treatment[J]. Phys Med Biol, 2017, 62(23): L41-50. doi:10.1088/1361-6560/aa9517 |
| [3] | Raaymakers BW, Lagendijk JJW, Overweg J, et al. Integrating a 1.5 T MRI scanner with a 6 MV accelerator: proof of concept[J]. Phys Med Biol, 2009, 54(12): N229-37. doi:10.1088/0031-9155/54/12/n01 |
| [4] | Lombardo E, Dhont J, Page D, et al. Real-time motion management in MRI-guided radiotherapy: Current status and AI-enabled prospects[J]. Radiother Oncol, 2024, 190: 109970. doi:10.1016/j.radonc.2023.109970 |
| [5] | Paganelli C, Whelan B, Peroni M, et al. MRI-guidance for motion management in external beam radiotherapy: current status and future challenges[J]. Phys Med Biol, 2018, 63(22): 22TR03. doi:10.1088/1361-6560/aaebcf |
| [6] | Van Reeth E, Tham IWK, Tan CH, et al. Super-resolution in magnetic resonance imaging: a review[J]. Concepts Magn Reson Part A, 2012, 40A(6): 306-25. doi:10.1002/cmr.a.21249 |
| [7] | Mo YJ, Wu Y, Yang XN, et al. Review the state-of-the-art technologies of semantic segmentation based on deep learning[J]. Neurocomputing, 2022, 493: 626-46. doi:10.1016/j.neucom.2022.01.005 |
| [8] | Lepcha DC, Goyal B, Dogra A, et al. Image super-resolution: a comprehensive review, recent trends, challenges and applications[J]. Inf Fusion, 2023, 91: 230-60. doi:10.1016/j.inffus.2022.10.007 |
| [9] | Dong YY, Yang F, Wen J, et al. Improvement of 2D cine image quality using 3D priors and cycle generative adversarial network for low field MRI-guided radiation therapy[J]. Med Phys, 2024, 51(5): 3495-509. doi:10.1002/mp.16860 |
| [10] | Xie HQ, Lei Y, Wang TH, et al. Synthesizing high-resolution magnetic resonance imaging using parallel cycle-consistent generative adversarial networks for fast magnetic resonance imaging[J]. Med Phys, 2022, 49(1): 357-69. doi:10.1002/mp.15380 |
| [11] | You A, Kim JK, Ryu IH, et al. Application of generative adversarial networks (GAN) for ophthalmology image domains: a survey[J]. Eye Vis, 2022, 9(1): 6. doi:10.1186/s40662-022-00277-3 |
| [12] | Rabbi J, Ray N, Schubert M, et al. Small-object detection in remote sensing images with end-to-end edge-enhanced GAN and object detector network[J]. Remote Sens, 2020, 12(9): 1432. doi:10.3390/rs12091432 |
| [13] | Radford A, Metz L, Chintala S. Unsupervised representation learning with deep convolutional generative adversarial networks[EB/OL]. 2015: arXiv: 1511.06434. |
| [14] | Zhang H, Goodfellow I, Metaxas D, et al. Self-attention generative adversarial networks[EB/OL]. 2018: arXiv: 1805.08318. . doi:10.48550/arXiv.1805.08318 |
| [15] | Chun J, Zhang H, Gach HM, et al. MRI super-resolution reconstruction for MRI-guided adaptive radiotherapy using cascaded deep learning: in the presence of limited training data and unknown translation model[J]. Med Phys, 2019, 46(9): 4148-64. doi:10.1002/mp.13717 |
| [16] | Huang BY, Xiao HN, Liu WW, et al. MRI super-resolution via realistic downsampling with adversarial learning[J]. Phys Med Biol, 2021, 66(20). DOI:10.1088/1361-6560/ac232e . |
| [17] | Yoon YH, Chun J, Kiser K, et al. Inter-scanner super-resolution of 3D cine MRI using a transfer-learning network for MRgRT[J]. Phys Med Biol, 2024, 69(11). DOI:10.1088/1361-6560/ad43ab . |
| [18] | Ho J, Jain A, Abbeel P. Denoising diffusion probabilistic models[EB/OL]. 2020: arXiv: 2006.11239. |
| [19] | Saharia C, Chan W, Chang HW, et al. Palette: image-to-image diffusion models[C]//Special Interest Group on Computer Graphics and Interactive Techniques Conference Proceedings. Vancouver BC Canada. ACM, 2022: 1-10. doi:10.1145/3528233.3530757 |
| [20] | Chen XQ, Qiu RLJ, Peng JB, et al. CBCT-based synthetic CT image generation using a diffusion model for CBCT-guided lung radiotherapy[J]. Med Phys, 2024, 51(11): 8168-78. doi:10.1002/mp.17328 |
| [21] | Liu Z, Lin YT, Cao Y, et al. Swin transformer: hierarchical vision transformer using shifted windows[C]//2021 IEEE/CVF International Conference on Computer Vision (ICCV). October 10-17, 2021, Montreal, QC, Canada. IEEE, 2022: 9992-10002. doi:10.1109/iccv48922.2021.00986 |
| [22] | Dalmaz O, Yurt M, Çukur T. ResViT: residual vision transformers for multimodal medical image synthesis[J]. IEEE Trans Med Imag, 2022, 41(10): 2598-614. doi:10.1109/tmi.2022.3167808 |
| [23] | Vaswani A, Shazeer N, Parmar N, et al. Attention is all you need[J]. Advances in neural information processing systems, 2017, 30. doi:10.3390/rs9080848 |
| [24] | Zhu JY, Park T, Isola P, et al. Unpaired image-to-image translation using cycle-consistent adversarial networks[C]//2017 IEEE International Conference on Computer Vision (ICCV). October 22-29, 2017, Venice, Italy. IEEE, 2017: 2242-51. doi:10.1109/iccv.2017.244 |
| [25] | Ronneberger O, Fischer P, Brox T. U-Net: convolutional networks for biomedical image segmentation[C]//Medical Image Computing and Computer-Assisted Intervention-MICCAI 2015. Cham: Springer, 2015: 234-41. doi:10.1007/978-3-319-24574-4_28 |
| [26] | Hu H, Gu JY, Zhang Z, et al. Relation networks for object detection[C]//2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition. June 18-23, 2018, Salt Lake City, UT, USA. IEEE, 2018: 3588-97. doi:10.1109/cvpr.2018.00378 |
| [27] | Hu H, Zhang Z, Xie ZD, et al. Local relation networks for image recognition[C]//2019 IEEE/CVF International Conference on Computer Vision (ICCV). October 27-November 2, 2019. Seoul, Korea. IEEE, 2019: 3463-72. doi:10.1109/iccv.2019.00356 |
| [28] | Thorwarth D. Functional imaging for radiotherapy treatment planning: current status and future directions-a review[J]. Br J Radiol, 2015, 88(1051): 20150056. doi:10.1259/bjr.20150056 |
| [29] | Galić I, Habijan M, Leventić H, et al. Machine learning empowering personalized medicine: a comprehensive review of medical image analysis methods[J]. Electronics, 2023, 12(21): 4411. doi:10.3390/electronics12214411 |
| [30] | Huynh E, Hosny A, Guthier C, et al. Artificial intelligence in radiation oncology[J]. Nat Rev Clin Oncol, 2020, 17(12): 771-81. doi:10.1038/s41571-020-0417-8 |
| [31] | Kazerouni A, Aghdam EK, Heidari M, et al. Diffusion models in medical imaging: a comprehensive survey[J]. Med Image Anal, 2023, 88: 102846. doi:10.1016/j.media.2023.102846 |
| [32] | Wendling M, Morrow A, Hoggarth M. An efficient protocol for radiotherapy quality control with machine learning[J]. Med Phys, 2020, 47(4): 1526-34. |
| [1] | 卢学麒, 陈华元, 吴秋岑, 温耀棋, 刘国光, 陈超敏. 一种基于深度特征融合的可解释性12导联心电图自动诊断模型研究[J]. 南方医科大学学报, 2026, 46(1): 208-218. |
| [2] | 吴秋岑, 卢学麒, 温耀棋, 洪永, 吴煜良, 陈超敏. II导联心电图中心肌梗死检测与定位:基于多尺度残差模块融合改进通道注意力模型[J]. 南方医科大学学报, 2025, 45(8): 1777-1790. |
| [3] | 郑子瑜, 杨夏颖, 吴圣杰, 张诗婕, 吕国荣, 柳培忠, 王珺, 何韶铮. 多特征融合的产时超声胎方位识别模型[J]. 南方医科大学学报, 2025, 45(7): 1563-1570. |
| [4] | 梁业东, 朱雄峰, 黄美燕, 张文聪, 郭翰宇, 冯前进. CRAKUT:融合对比区域注意力机制与临床先验知识的U-Transformer用于放射学报告生成[J]. 南方医科大学学报, 2025, 45(6): 1343-1352. |
| [5] | 渠梦, 傅蓉. ResLSTM-TemporalSE:多导联心电信号的自动分类[J]. 南方医科大学学报, 2025, 45(12): 2708-2717. |
| [6] | 谢辉荣, 胡潮滨, 梁国华, 韩红喆, 黄牧, 冯前进. 大学生心理压力智能评估:基于融合文本与影像的多模态模型的设计及验证[J]. 南方医科大学学报, 2025, 45(11): 2504-2510. |
| [7] | 贺亚迪, 周炫汝, 金锦辉, 宋婷. 基于PE-CycleGAN网络的鼻咽癌自适应放疗CBCT-sCT生成研究[J]. 南方医科大学学报, 2025, 45(1): 179-186. |
| [8] | 方威扬, 肖慧, 王爽, 林晓明, 陈超敏. 基于MRI影像和临床参数特征融合的深度学习模型预测术前肝细胞癌的细胞角蛋白19状态[J]. 南方医科大学学报, 2024, 44(9): 1738-1751. |
| [9] | 欧嘉志, 詹长安, 杨丰. 一维卷积神经网络的自编码癫痫发作异常检测模型[J]. 南方医科大学学报, 2024, 44(9): 1796-1804. |
| [10] | 汪辰, 蒙铭强, 李明强, 王永波, 曾栋, 边兆英, 马建华. 基于双域Transformer耦合特征学习的CT截断数据重建模型[J]. 南方医科大学学报, 2024, 44(5): 950-959. |
| [11] | 龙楷兴, 翁丹仪, 耿 舰, 路艳蒙, 周志涛, 曹 蕾. 基于多模态多示例学习的免疫介导性肾小球疾病自动分类方法[J]. 南方医科大学学报, 2024, 44(3): 585-593. |
| [12] | 肖 慧, 方威扬, 林铭俊, 周振忠, 费洪文, 陈超敏. 基于两阶段分析的多尺度颈动脉斑块检测方法[J]. 南方医科大学学报, 2024, 44(2): 387-396. |
| [13] | 刘操林, 邹青清, 王梦虹, 杨芹枚, 宋丽文, 陆紫箫, 冯前进, 赵英华. 鉴别原发性骨肿瘤骨样和软骨样基质矿化:基于CT和临床特征的深度学习融合模型的多中心回顾性研究[J]. 南方医科大学学报, 2024, 44(12): 2412-2420. |
| [14] | 黄品瑜, 钟丽明, 郑楷宜, 陈泽立, 肖若琳, 全显跃, 阳 维. 多期相CT合成辅助的腹部多器官图像分割[J]. 南方医科大学学报, 2024, 44(1): 83-92. |
| [15] | 弥 佳, 周宇佳, 冯前进. 基于正交视角X线图像重建的3D/2D配准方法[J]. 南方医科大学学报, 2023, 43(9): 1636-1643. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||