南方医科大学学报 ›› 2026, Vol. 46 ›› Issue (1): 208-218.doi: 10.12122/j.issn.1673-4254.2026.01.23
卢学麒1( ), 陈华元2(
), 陈华元2( ), 吴秋岑1, 温耀棋1, 刘国光3(
), 吴秋岑1, 温耀棋1, 刘国光3( ), 陈超敏1(
), 陈超敏1( )
)
收稿日期:2025-06-24
出版日期:2026-01-20
发布日期:2026-01-16
通讯作者:
刘国光,陈超敏
E-mail:415081161@qq.com;67936668@qq.com;571611621@qq.com
作者简介:陈华元,工程师,E-mail: 415081161@qq.com基金资助:
Xueqi LU1( ), Huayuan CHEN2(
), Huayuan CHEN2( ), Qiucen WU1, Yaoqi WEN1, Guoguang LIU3(
), Qiucen WU1, Yaoqi WEN1, Guoguang LIU3( ), Chaomin CHEN1(
), Chaomin CHEN1( )
)
Received:2025-06-24
Online:2026-01-20
Published:2026-01-16
Contact:
Guoguang LIU, Chaomin CHEN
E-mail:415081161@qq.com;67936668@qq.com;571611621@qq.com
摘要:
目的 提升12导联心电图(ECG)自动诊断的准确性和可信度。 方法 提出了一种基于深度特征融合的12导联ECG自动诊断模型(MRHL-ECGNet)。该模型包含多尺度特征提取前端、ResNet-34、全局特征混合模块及时间序列分析模块,首次将Hyena Hierarchy卷积算子应用于12导联心电图自动诊断任务中,以高效捕捉ECG中的长程依赖关系,并显著降低模型计算复杂度。同时采用基于积分梯度(IG)的可解释性分析技术,实现MRHL-ECGNet决策依据可视化。使用CPSC2018数据集对MRHL-ECGNet进行训练和测试,并采用多项定量评价指标与评估实验对MRHL-ECGNet进行全面评估。 结果 在测试集上对9种类别ECG的分类任务中,MRHL-ECGNet的准确率、AUC值、F1分数、精确率和召回率分别达到0.972、0.983、0.864、0.873和0.857,均优于其他对比模型,且在GPU上对单样本输出诊断结果所需的时间为0.007s,在CPU上也仅需0.156s,内存占用为67.196MB。 结论 本研究提出的MRHL-ECGNet不仅具有卓越的分类性能,还具备轻量化及可解释性的特点,在临床ECG辅助诊断中具有较高的应用价值。
卢学麒, 陈华元, 吴秋岑, 温耀棋, 刘国光, 陈超敏. 一种基于深度特征融合的可解释性12导联心电图自动诊断模型研究[J]. 南方医科大学学报, 2026, 46(1): 208-218.
Xueqi LU, Huayuan CHEN, Qiucen WU, Yaoqi WEN, Guoguang LIU, Chaomin CHEN. Evaluation of an interpretable 12-lead ECG automatic diagnosis model based on deep feature fusion[J]. Journal of Southern Medical University, 2026, 46(1): 208-218.
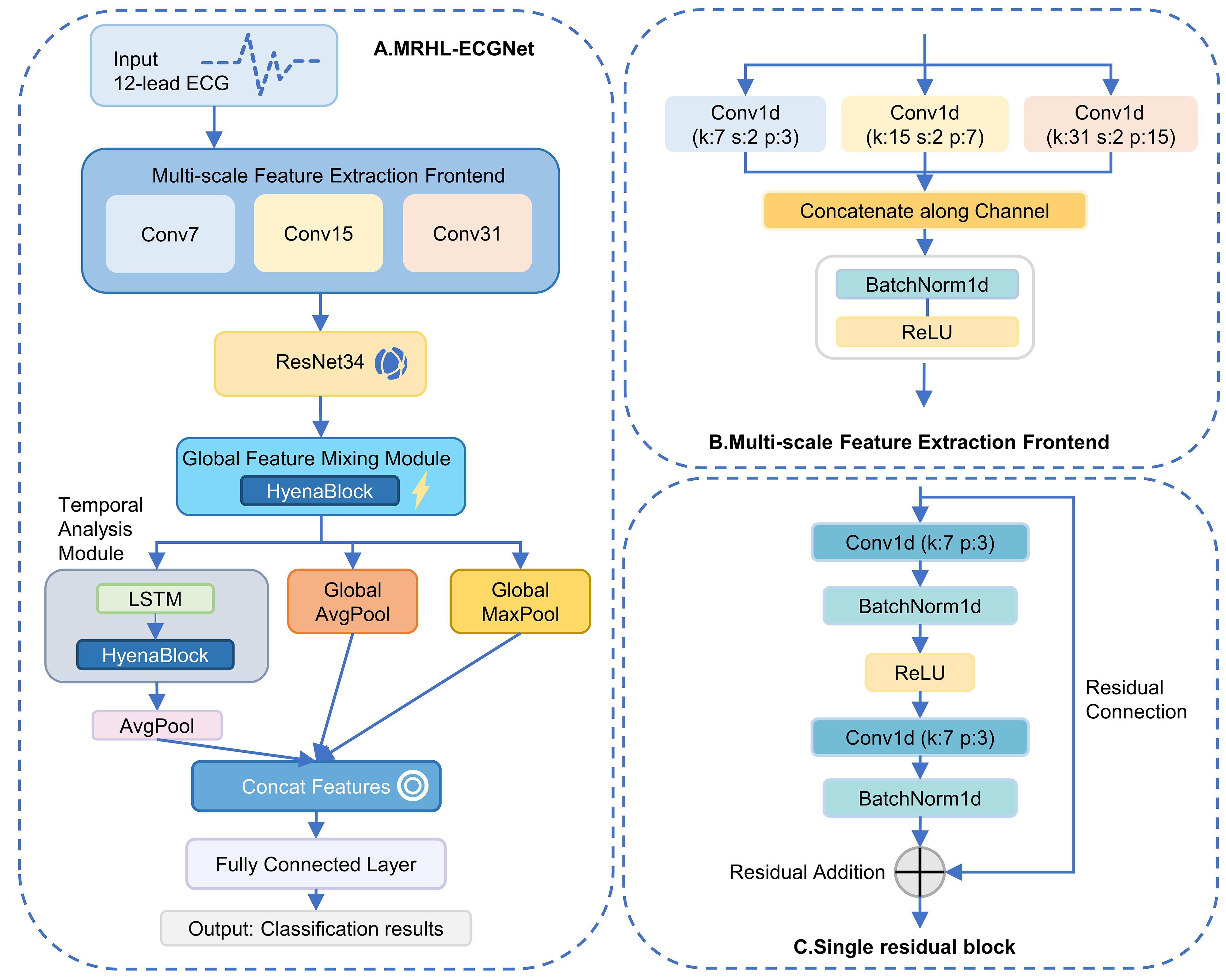
图2 模型结构示意图1
Fig.2 Schematic diagram of the model structure 1. A: Overall structure diagram of MRHL-ECGNet. B: Structure diagram of the multi-scale feature extraction front-end. C: Structure diagram of the single residual block.
| Module | In/Out & K and Other Parameters |
|---|---|
| Input Layer | In: 12×T |
| Multi-scale feature extraction frontend | In: 12×T; Out: 64×T/2; K: 7, 15, 31; Stride: 2 |
| ResNet-34 Layer1 | In: 64; Out: 64; K: 7; BasicBlock1d: 3; Downsample: No |
| ResNet-34 Layer2 | In: 64; Out: 128; K: 7; BasicBlock1d: 4; Downsample: Yes |
| ResNet-34 Layer3 | In: 128; Out: 256; K: 7; BasicBlock1d: 6; Downsample: Yes |
| ResNet-34 Layer4 | In: 256; Out: 512; K: 7; BasicBlock1d: 3; Downsample: Yes |
| Global feature mixing module | In: 512; Out: 512; K: 31 |
| Temporal analysis module (LSTM) | In: 512; Hidden state: 64 |
| Temporal analysis module (Hyena) | In: 64; Out: 64; K: 15 |
| Feature fusion and classifier | Concatenated in: 512×2+64; Fully connected layer Output: Classification results |
表1 MRHL-ECGNet参数设置
Tab.1 Parameter settings of the MRHL-ECGNet model
| Module | In/Out & K and Other Parameters |
|---|---|
| Input Layer | In: 12×T |
| Multi-scale feature extraction frontend | In: 12×T; Out: 64×T/2; K: 7, 15, 31; Stride: 2 |
| ResNet-34 Layer1 | In: 64; Out: 64; K: 7; BasicBlock1d: 3; Downsample: No |
| ResNet-34 Layer2 | In: 64; Out: 128; K: 7; BasicBlock1d: 4; Downsample: Yes |
| ResNet-34 Layer3 | In: 128; Out: 256; K: 7; BasicBlock1d: 6; Downsample: Yes |
| ResNet-34 Layer4 | In: 256; Out: 512; K: 7; BasicBlock1d: 3; Downsample: Yes |
| Global feature mixing module | In: 512; Out: 512; K: 31 |
| Temporal analysis module (LSTM) | In: 512; Hidden state: 64 |
| Temporal analysis module (Hyena) | In: 64; Out: 64; K: 15 |
| Feature fusion and classifier | Concatenated in: 512×2+64; Fully connected layer Output: Classification results |
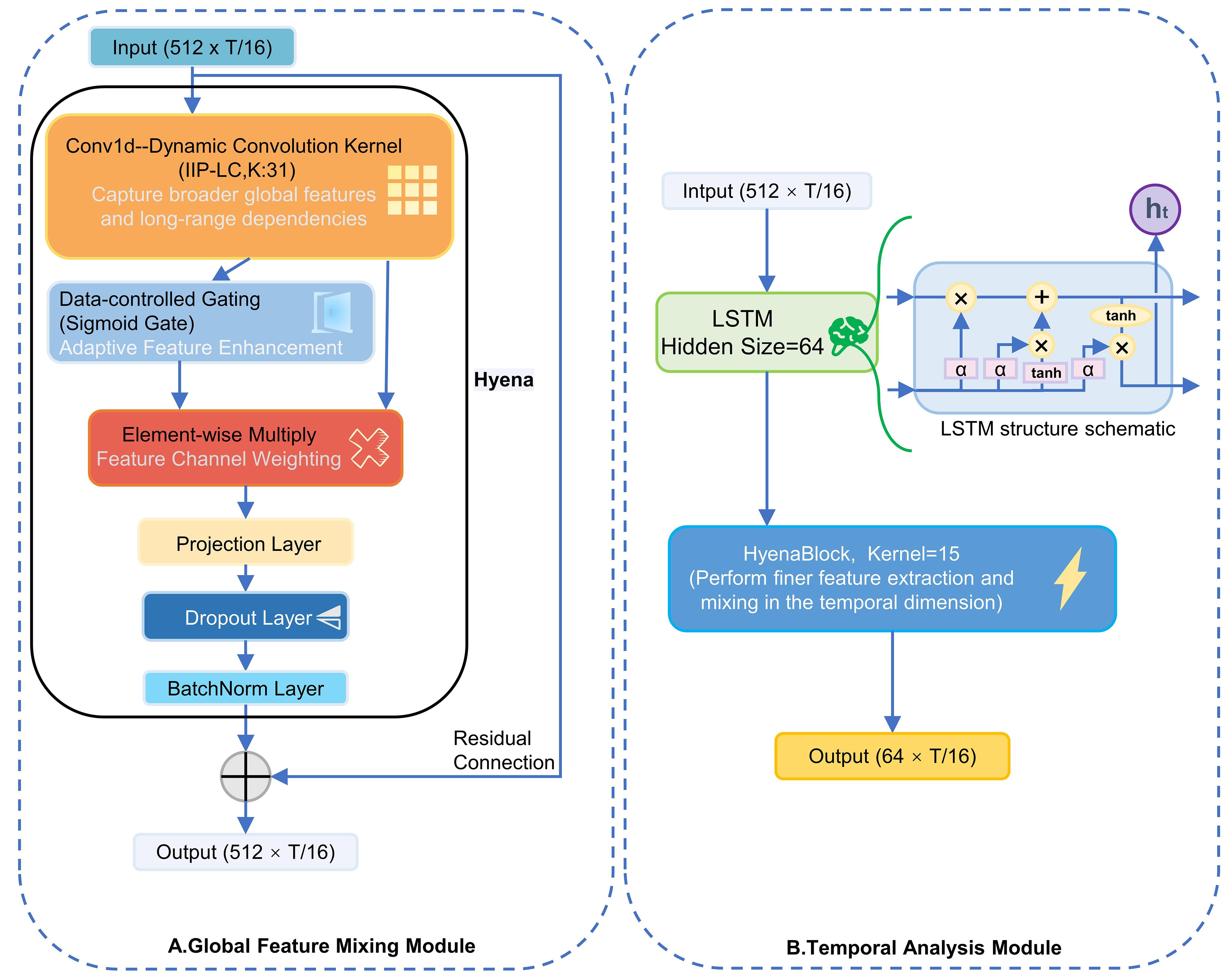
图3 模型结构示意图2
Fig.3 Schematic diagram of model structure 2. A: Global feature merging module structure diagram; B: Temporal analysis module structure diagram.
| Item | Configuration |
|---|---|
| Operation system | Windows 11 |
| CPU | Intel Core i5-13490F |
| GPU | NVIDIA GeForce RTX 4060Ti |
| RAM | 16GB |
| Cuda version | 12.7 |
| Pytorch version | 2.3.1 |
| Anaconda version | 24.9.2 |
表2 实验环境
Tab.2 Experimental environment for model testing
| Item | Configuration |
|---|---|
| Operation system | Windows 11 |
| CPU | Intel Core i5-13490F |
| GPU | NVIDIA GeForce RTX 4060Ti |
| RAM | 16GB |
| Cuda version | 12.7 |
| Pytorch version | 2.3.1 |
| Anaconda version | 24.9.2 |
| Method | Accuracy | AUC | F1s | Precision | Recall |
|---|---|---|---|---|---|
| Zhang D et al.[ | 0.963 | 0.961 | 0.823 | 0.826 | 0.821 |
| Reddy L et al.[ | 0.895 | 0.913 | 0.810 | 0.812 | 0.809 |
| Hwang S et al.[ | 0.905 | 0.928 | 0.797 | 0.808 | 0.787 |
| Strodthoff N et al.[ | 0.901 | 0.941 | 0.813 | 0.836 | 0.792 |
| MRHL-ECGNet | 0.972 | 0.983 | 0.864 | 0.873 | 0.857 |
表3 12导联ECG自动诊断模型性能比较结果
Tab.3 Comparison results of the performance of the 12-lead ECG automatic diagnosis model
| Method | Accuracy | AUC | F1s | Precision | Recall |
|---|---|---|---|---|---|
| Zhang D et al.[ | 0.963 | 0.961 | 0.823 | 0.826 | 0.821 |
| Reddy L et al.[ | 0.895 | 0.913 | 0.810 | 0.812 | 0.809 |
| Hwang S et al.[ | 0.905 | 0.928 | 0.797 | 0.808 | 0.787 |
| Strodthoff N et al.[ | 0.901 | 0.941 | 0.813 | 0.836 | 0.792 |
| MRHL-ECGNet | 0.972 | 0.983 | 0.864 | 0.873 | 0.857 |
| Diagnosis category | Accuracy | AUC | F1s | Precision | Recall |
|---|---|---|---|---|---|
| Normal | 0.963 | 0.980 | 0.819 | 0.837 | 0.802 |
| AF | 0.977 | 0.995 | 0.927 | 0.944 | 0.911 |
| I-AVB | 0.975 | 0.975 | 0.896 | 0.912 | 0.882 |
| LBBB | 0.994 | 0.998 | 0.920 | 0.986 | 0.862 |
| RBBB | 0.961 | 0.989 | 0.925 | 0.954 | 0.898 |
| PAC | 0.966 | 0.952 | 0.739 | 0.683 | 0.804 |
| PVC | 0.965 | 0.982 | 0.836 | 0.803 | 0.872 |
| STD | 0.951 | 0.976 | 0.793 | 0.844 | 0.747 |
| STE | 0.996 | 0.998 | 0.914 | 0.889 | 0.941 |
| AVG | 0.972 | 0.983 | 0.864 | 0.873 | 0.857 |
表4 MRHL-ECGNet在9种不同诊断类别ECG上的性能表现
Tab.4 Performance of MRHL-ECGNet model on 9 different diagnostic categories of ECG
| Diagnosis category | Accuracy | AUC | F1s | Precision | Recall |
|---|---|---|---|---|---|
| Normal | 0.963 | 0.980 | 0.819 | 0.837 | 0.802 |
| AF | 0.977 | 0.995 | 0.927 | 0.944 | 0.911 |
| I-AVB | 0.975 | 0.975 | 0.896 | 0.912 | 0.882 |
| LBBB | 0.994 | 0.998 | 0.920 | 0.986 | 0.862 |
| RBBB | 0.961 | 0.989 | 0.925 | 0.954 | 0.898 |
| PAC | 0.966 | 0.952 | 0.739 | 0.683 | 0.804 |
| PVC | 0.965 | 0.982 | 0.836 | 0.803 | 0.872 |
| STD | 0.951 | 0.976 | 0.793 | 0.844 | 0.747 |
| STE | 0.996 | 0.998 | 0.914 | 0.889 | 0.941 |
| AVG | 0.972 | 0.983 | 0.864 | 0.873 | 0.857 |
| Experiment ID | Configuration | Accuracy | AUC | F1s | Precision | Recall |
|---|---|---|---|---|---|---|
| 1 | Remove multi-scale feature extraction frontend | 0.951 | 0.968 | 0.846 | 0.844 | 0.851 |
| 2 | Remove resnet-34 | 0.898 | 0.901 | 0.803 | 0.796 | 0.812 |
| 3 | Remove temporal analysis module | 0.912 | 0.919 | 0.820 | 0.821 | 0.819 |
| 4 | Remove global avgpool | 0.956 | 0.971 | 0.837 | 0.835 | 0.840 |
| 5 | Remove global maxpool | 0.967 | 0.973 | 0.833 | 0.829 | 0.838 |
| 6 | Remove all hyenablock | 0.905 | 0.909 | 0.799 | 0.798 | 0.802 |
| 7 | Replace hyenablock with transformerblock | 0.970 | 0.985 | 0.847 | 0.836 | 0.859 |
| 8 | Full MRHL-ECGNet | 0.972 | 0.983 | 0.864 | 0.873 | 0.857 |
表5 消融实验及机制替换实验结果
Tab.5 Results of the ablation experiment and the mechanism replacement experiment
| Experiment ID | Configuration | Accuracy | AUC | F1s | Precision | Recall |
|---|---|---|---|---|---|---|
| 1 | Remove multi-scale feature extraction frontend | 0.951 | 0.968 | 0.846 | 0.844 | 0.851 |
| 2 | Remove resnet-34 | 0.898 | 0.901 | 0.803 | 0.796 | 0.812 |
| 3 | Remove temporal analysis module | 0.912 | 0.919 | 0.820 | 0.821 | 0.819 |
| 4 | Remove global avgpool | 0.956 | 0.971 | 0.837 | 0.835 | 0.840 |
| 5 | Remove global maxpool | 0.967 | 0.973 | 0.833 | 0.829 | 0.838 |
| 6 | Remove all hyenablock | 0.905 | 0.909 | 0.799 | 0.798 | 0.802 |
| 7 | Replace hyenablock with transformerblock | 0.970 | 0.985 | 0.847 | 0.836 | 0.859 |
| 8 | Full MRHL-ECGNet | 0.972 | 0.983 | 0.864 | 0.873 | 0.857 |
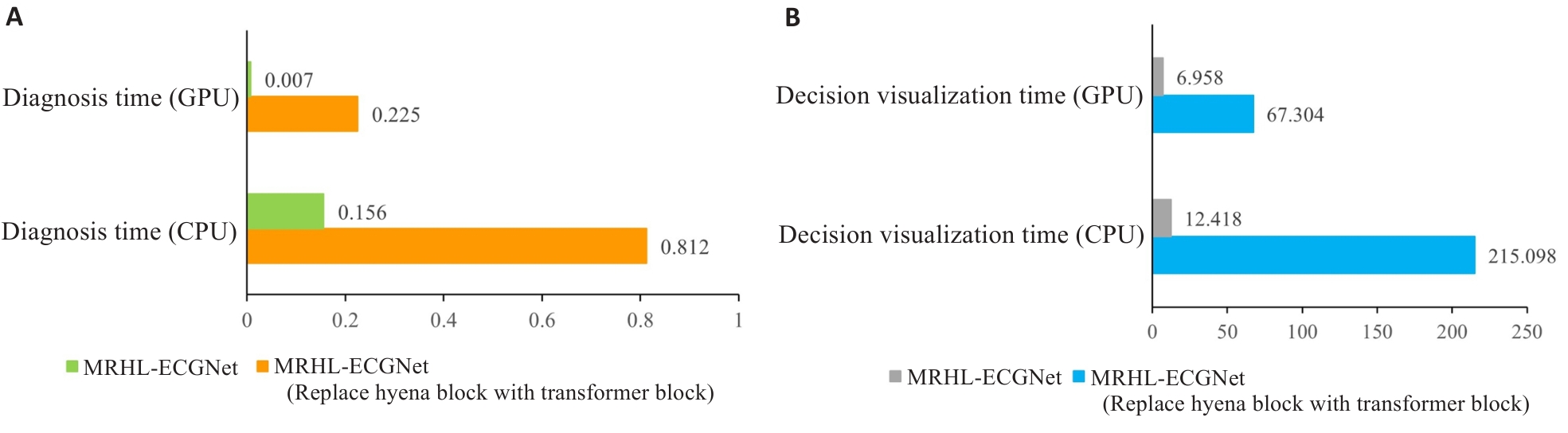
图5 模型运行时间
Fig.5 Model running time. A: Time required for diagnostic results output under different conditions. B: Time for generating the decision-making basis diagram under different conditions.
| Module | Parameters | Memory (MB) |
|---|---|---|
| Multi-scale feature extraction frontend | 13920 | 0.053 |
| ResNet-34 | 16589312 | 63.283 |
| Global feature mixing module | 295424 | 1.127 |
| Temporal analysis module | 154176 | 0.588 |
| Hyenablock | 8256 | 0.031 |
| Mrhl-ecgnet | 17615017 | 67.196 |
| Mrhl-ecgnet (replace hyenablock with transformerblock) | 20746921 | 80.226 |
表6 模型参数量及内存占用
Tab.6 Model parameter quantity and memory usage
| Module | Parameters | Memory (MB) |
|---|---|---|
| Multi-scale feature extraction frontend | 13920 | 0.053 |
| ResNet-34 | 16589312 | 63.283 |
| Global feature mixing module | 295424 | 1.127 |
| Temporal analysis module | 154176 | 0.588 |
| Hyenablock | 8256 | 0.031 |
| Mrhl-ecgnet | 17615017 | 67.196 |
| Mrhl-ecgnet (replace hyenablock with transformerblock) | 20746921 | 80.226 |
| [1] | Berkaya SK, Uysal AK, Gunal ES, et al. A survey on ECG analysis[J]. Biomed Sig Proc Control, 2018, 43: 216-35. doi:10.1016/j.bspc.2018.03.003 |
| [2] | Chou's Electrocardiography in Clinical Practice[M]. Elsevier Science Health Science div,2008 . doi:10.1016/b978-141603774-3.10027-9 |
| [3] | Hong SD, Zhou YX, Shang JY, et al. Opportunities and challenges of deep learning methods for electrocardiogram data: a systematic review[J]. Comput Biol Med, 2020, 122: 103801. doi:10.1016/j.compbiomed.2020.103801 |
| [4] | Priori SG, Blomström-Lundqvist C, Mazzanti A, et al. 2015 ESC Guidelines for the management of patients with ventricular arrhythmias and the prevention of sudden cardiac death: The Task Force for the Management of Patients with Ventricular Arrhythmias and the Prevention of Sudden Cardiac Death of the European Society of Cardiology (ESC). Endorsed by: Association for European Paediatric and Congenital Cardiology (AEPC)[J]. Eur Heart J, 2015, 36(41): 2793-867. doi:10.1093/eurheartj/ehv316 |
| [5] | Hannun AY, Rajpurkar P, Haghpanahi M, et al. Cardiologist-level arrhythmia detection and classification in ambulatory electro-cardiograms using a deep neural network[J]. Nat Med, 2019, 25(1): 65-9. doi:10.1038/s41591-018-0268-3 |
| [6] | Yang XZ, Zhang XY, Yang MY, et al. 12-Lead ECG arrhythmia classification using cascaded convolutional neural network and expert feature[J]. J Electrocardiol, 2021, 67: 56-62. doi:10.1016/j.jelectrocard.2021.04.016 |
| [7] | Vaswani A, Shazeer NM, Parmar N, et al. Attention is all you need[C]//Neural Information Processing Systems., 2017 |
| [8] | Dosovitskiy A, Beyer L, Kolesnikov A, et al. An image is worth 16x16 words: Transformers for image recognition at scale[J]. arxiv preprint arxiv: 2010. 11929, 2020. |
| [9] | Zhou HY, Zhang SH, Peng JQ, et al. Informer: beyond efficient transformer for long sequence time-series forecasting[J]. Proc AAAI Conf Artif Intell, 2021, 35(12): 11106-15. doi:10.1609/aaai.v35i12.17325 |
| [10] | Lee J, Yoon W, Kim S, et al. BioBERT: a pre-trained biomedical language representation model for biomedical text mining[J]. Bioinformatics, 2020, 36(4): 1234-40. doi:10.1093/bioinformatics/btz682 |
| [11] | Zhang SY, Lian C, Xu BR, et al. A token selection-based multi-scale dual-branch CNN-transformer network for 12-lead ECG signal classification[J]. Knowl Based Syst, 2023, 280: 111006. doi:10.1016/j.knosys.2023.111006 |
| [12] | Zheng JW, Zhang JM, Danioko S, et al. A 12-lead electrocardiogram database for arrhythmia research covering more than 10, 000 patients[J]. Sci Data, 2020, 7(1): 48. doi:10.1038/s41597-020-0386-x |
| [13] | Petch J, Di S, Nelson W. Opening the black box: the promise and limitations of explainable machine learning in cardiology[J]. Can J Cardiol, 2022, 38(2): 204-13. doi:10.1016/j.cjca.2021.09.004 |
| [14] | Zhang DD, Yang S, Yuan XH, et al. Interpretable deep learning for automatic diagnosis of 12-lead electrocardiogram[J]. iScience, 2021, 24(4): 102373. doi:10.1016/j.isci.2021.102373 |
| [15] | Lundberg SM, Lee SI. A unified approach to interpreting model predictions[C]//Neural Information Processing Systems., 2017 |
| [16] | Reddy L, Talwar V, Alle S, et al. IMLE-net: an interpretable multi-level multi-channel model for ECG classification[C]//2021 IEEE International Conference on Systems, Man, and Cybernetics (SMC). October 17-20, 2021. Melbourne, Australia. IEEE, 2021: 1068-1074. doi:10.1109/smc52423.2021.9658706 |
| [17] | Liu FF, Liu CY, Zhao LN, et al. An open access database for evaluating the algorithms of electrocardiogram rhythm and morphology abnormality detection[J]. J Med Imaging Hlth Inform, 2018, 8(7): 1368-73. doi:10.1166/jmihi.2018.2442 |
| [18] | Zhang HZ, Zhao W, Liu S. SE-ECGNet: a multi-scale deep residual network with squeeze-and-excitation module for ECG signal classification[C]//2020 IEEE International Conference on Bioinformatics and Biomedicine (BIBM). December 16-19, 2020. Seoul, Korea. IEEE, 2020: 2685-2691. doi:10.1109/bibm49941.2020.9313548 |
| [19] | He KM, Zhang XY, Ren SQ, et al. Deep residual learning for image recognition[C]//2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). June 27-30, 2016, Las Vegas, NV, USA. IEEE, 2016: 770-8. doi:10.1109/cvpr.2016.90 |
| [20] | Poli M, Massaroli S, Nguyen E, et al. Hyena hierarchy: Towards larger convolutional language models[C]//International Conference on Machine Learning. PMLR, 2023: 28043-78. |
| [21] | Yang W, Deo R, Guo WS. Functional feature extraction and validation from twelve-lead electrocardiograms to identify atrial fibrillation[J]. Commun Med (Lond), 2025, 5(1): 32. doi:10.1038/s43856-025-00749-2 |
| [22] | Hochreiter S, Schmidhuber J. Long short-term memory[J]. Neural Comput, 1997, 9(8): 1735-80. doi:10.1162/neco.1997.9.8.1735 |
| [23] | Kingma D P, Ba J. Adam: A method for stochastic optimization[J]. arxiv preprint arxiv:, 2014. |
| [24] | Barredo Arrieta A, Díaz-Rodríguez N, Del Ser J, et al. Explainable artificial intelligence (XAI): concepts, taxonomies, opportunities and challenges toward responsible AI[J]. Inf Fusion, 2020, 58: 82-115. doi:10.1016/j.inffus.2019.12.012 |
| [25] | Sundararajan M, Taly A, Yan Q. Gradients of counterfactuals[J]. arxiv preprint arxiv:, 2016. |
| [26] | Sokolova M, Lapalme G. A systematic analysis of performance measures for classification tasks[J]. Inf Process Manag, 2009, 45(4): 427-37. doi:10.1016/j.ipm.2009.03.002 |
| [27] | Hwang S, Cha J, Heo J, et al. Multi-label abnormality classification from 12-lead ECG using a 2D residual U-net[C]//ICASSP 2024 - 2024 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP). April 14-19, 2024, Seoul, Korea, Republic of. IEEE, 2024: 2265-9. doi:10.1109/icassp48485.2024.10448259 |
| [28] | Strodthoff N, Wagner P, Schaeffter T, et al. Deep learning for ECG analysis: benchmarks and insights from PTB-XL[J]. IEEE J Biomed Health Inform, 2021, 25(5): 1519-28. doi:10.1109/jbhi.2020.3022989 |
| [29] | Conen D, Adam M, Roche F, et al. Premature atrial contractions in the general population: frequency and risk factors[J]. Circulation, 2012, 126(19): 2302-8. doi:10.1161/circulationaha.112.112300 |
| [30] | Joglar JA, Chung MK, Armbruster AL, et al. 2023 ACC/AHA/ACCP/HRS guideline for the diagnosis and management of atrial fibrillation: a report of the American college of cardiology/American heart association joint committee on clinical practice guidelines[J]. Circulation, 2024, 149(1): e1-156. doi:10.1161/cir.0000000000001207 |
| [31] | Ikeda T. Right bundle branch block: current considerations[J]. Curr Cardiol Rev, 2021, 17(1): 24-30. doi:10.2174/1573403x16666200708111553 |
| [1] | 吴秋岑, 卢学麒, 温耀棋, 洪永, 吴煜良, 陈超敏. II导联心电图中心肌梗死检测与定位:基于多尺度残差模块融合改进通道注意力模型[J]. 南方医科大学学报, 2025, 45(8): 1777-1790. |
| [2] | 郑子瑜, 杨夏颖, 吴圣杰, 张诗婕, 吕国荣, 柳培忠, 王珺, 何韶铮. 多特征融合的产时超声胎方位识别模型[J]. 南方医科大学学报, 2025, 45(7): 1563-1570. |
| [3] | 渠梦, 傅蓉. ResLSTM-TemporalSE:多导联心电信号的自动分类[J]. 南方医科大学学报, 2025, 45(12): 2708-2717. |
| [4] | 谢辉荣, 胡潮滨, 梁国华, 韩红喆, 黄牧, 冯前进. 大学生心理压力智能评估:基于融合文本与影像的多模态模型的设计及验证[J]. 南方医科大学学报, 2025, 45(11): 2504-2510. |
| [5] | 贺亚迪, 周炫汝, 金锦辉, 宋婷. 基于PE-CycleGAN网络的鼻咽癌自适应放疗CBCT-sCT生成研究[J]. 南方医科大学学报, 2025, 45(1): 179-186. |
| [6] | 方威扬, 肖慧, 王爽, 林晓明, 陈超敏. 基于MRI影像和临床参数特征融合的深度学习模型预测术前肝细胞癌的细胞角蛋白19状态[J]. 南方医科大学学报, 2024, 44(9): 1738-1751. |
| [7] | 欧嘉志, 詹长安, 杨丰. 一维卷积神经网络的自编码癫痫发作异常检测模型[J]. 南方医科大学学报, 2024, 44(9): 1796-1804. |
| [8] | 汪辰, 蒙铭强, 李明强, 王永波, 曾栋, 边兆英, 马建华. 基于双域Transformer耦合特征学习的CT截断数据重建模型[J]. 南方医科大学学报, 2024, 44(5): 950-959. |
| [9] | 龙楷兴, 翁丹仪, 耿 舰, 路艳蒙, 周志涛, 曹 蕾. 基于多模态多示例学习的免疫介导性肾小球疾病自动分类方法[J]. 南方医科大学学报, 2024, 44(3): 585-593. |
| [10] | 肖 慧, 方威扬, 林铭俊, 周振忠, 费洪文, 陈超敏. 基于两阶段分析的多尺度颈动脉斑块检测方法[J]. 南方医科大学学报, 2024, 44(2): 387-396. |
| [11] | 刘操林, 邹青清, 王梦虹, 杨芹枚, 宋丽文, 陆紫箫, 冯前进, 赵英华. 鉴别原发性骨肿瘤骨样和软骨样基质矿化:基于CT和临床特征的深度学习融合模型的多中心回顾性研究[J]. 南方医科大学学报, 2024, 44(12): 2412-2420. |
| [12] | 弥 佳, 周宇佳, 冯前进. 基于正交视角X线图像重建的3D/2D配准方法[J]. 南方医科大学学报, 2023, 43(9): 1636-1643. |
| [13] | 楚智钦, 屈耀铭, 钟 涛, 梁淑君, 温志波, 张 煜. 磁共振酰胺质子转移模态的胶质瘤IDH基因分型识别:基于深度学习的Dual-Aware框架[J]. 南方医科大学学报, 2023, 43(8): 1379-1387. |
| [14] | 于佳弘, 张昆鹏, 靳 爽, 苏 哲, 徐晓桐, 张 华. 弦图插值结合UNIT网络图像转换的CT金属伪影校正[J]. 南方医科大学学报, 2023, 43(7): 1214-1223. |
| [15] | 周 昊, 曾 栋, 边兆英, 马建华. 基于半监督网络的组织感知CT图像对比度的增强方法[J]. 南方医科大学学报, 2023, 43(6): 985-993. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||