南方医科大学学报 ›› 2026, Vol. 46 ›› Issue (4): 939-945.doi: 10.12122/j.issn.1673-4254.2026.04.22
• • 上一篇
谭顺谦1( ), 卓俐1, 曾敏1, 黄方俊2, 朱君1, 蔡光瑶3, 甄鑫1(
), 卓俐1, 曾敏1, 黄方俊2, 朱君1, 蔡光瑶3, 甄鑫1( )
)
收稿日期:2025-09-03
出版日期:2026-04-20
发布日期:2026-04-24
通讯作者:
甄鑫
E-mail:tann66643@gmail.com;xinzhen@smu.edu.cn
作者简介:谭顺谦,在读硕士研究生,E-mail: tann66643@gmail.com
基金资助:
Shunqian TAN1( ), Li ZHUO1, Min ZENG1, Fangjun HUANG2, Jun ZHU1, Guangyao CAI3, Xin ZHEN1(
), Li ZHUO1, Min ZENG1, Fangjun HUANG2, Jun ZHU1, Guangyao CAI3, Xin ZHEN1( )
)
Received:2025-09-03
Online:2026-04-20
Published:2026-04-24
Contact:
Xin ZHEN
E-mail:tann66643@gmail.com;xinzhen@smu.edu.cn
Supported by:摘要:
目的 评估一种基于 Transformer 深度学习模型、结合真实世界实验室检验指标的卵巢癌鉴别诊断性能。 方法 回顾性收集2012年1月1日~2021年4月4日同济妇产科医院收治的卵巢癌患者及良性病变患者的99项实验室检验指标和临床资料。通过基于ANOVA F检验的特征选择算法在训练集上选取20个关键特征,并采用表格数据转换器将每例患者转化为统一的嵌入向量。随后,利用改良的堆叠式Transformer对特征向量进行编码并训练模型。将该模型与多种传统机器学习方法进行比较,评价指标包括受试者工作特征曲线下面积(AUC)、准确率(ACC)、灵敏度(SEN)和特异度(SPE),通过五折交叉验证评估模型的泛化能力与有效性。 结果 五折交叉验证结果显示,基于Transformer深度学习模型在预测卵巢癌患者中表现最佳(AUC=0.931,ACC=0.813,SEN=0.833,SPE=0.865)。 结论 本研究提出的基于Transformer的模型在卵巢癌预测中具有较高的准确性和泛化能力,为卵巢肿瘤的临床辅助诊断提供了新的技术手段。
谭顺谦, 卓俐, 曾敏, 黄方俊, 朱君, 蔡光瑶, 甄鑫. 基于实验室指标的Transformer模型可高效鉴别卵巢癌[J]. 南方医科大学学报, 2026, 46(4): 939-945.
Shunqian TAN, Li ZHUO, Min ZENG, Fangjun HUANG, Jun ZHU, Guangyao CAI, Xin ZHEN. A Transformer-based model using laboratory indicators efficiently differentiates ovarian cancer[J]. Journal of Southern Medical University, 2026, 46(4): 939-945.
| Types of laboratory test indicators | Features name |
|---|---|
| Clinical features (n=1) | Age |
| Blood test indicators (n=17) | Albumin (ALB), Globulin (GLB), Albumin to globulin ratio (A/G), Carbohydrate antigen 125 (CA125), Alkaline Phosphatase (ALP), Carbohydrate antigen 724 (CA724), D-Dimer Quantification (D-D), Fibrinogen (FIB), High density lipoprotein (HDL), Total Bilirubin, Lactate dehydrogenase (LDH), Neutrophil Percentage, Lymphocyte Percentage, Plateletcrit,Platelet count (PLT), Bicarbonate, Thrombin Time (TT) |
| Urine test indicators (n=2) | Urine bilirubin (BIL), PH in urine(PH) |
表1 筛选后检验指标特征
Tab.1 Selected laboratory test features
| Types of laboratory test indicators | Features name |
|---|---|
| Clinical features (n=1) | Age |
| Blood test indicators (n=17) | Albumin (ALB), Globulin (GLB), Albumin to globulin ratio (A/G), Carbohydrate antigen 125 (CA125), Alkaline Phosphatase (ALP), Carbohydrate antigen 724 (CA724), D-Dimer Quantification (D-D), Fibrinogen (FIB), High density lipoprotein (HDL), Total Bilirubin, Lactate dehydrogenase (LDH), Neutrophil Percentage, Lymphocyte Percentage, Plateletcrit,Platelet count (PLT), Bicarbonate, Thrombin Time (TT) |
| Urine test indicators (n=2) | Urine bilirubin (BIL), PH in urine(PH) |
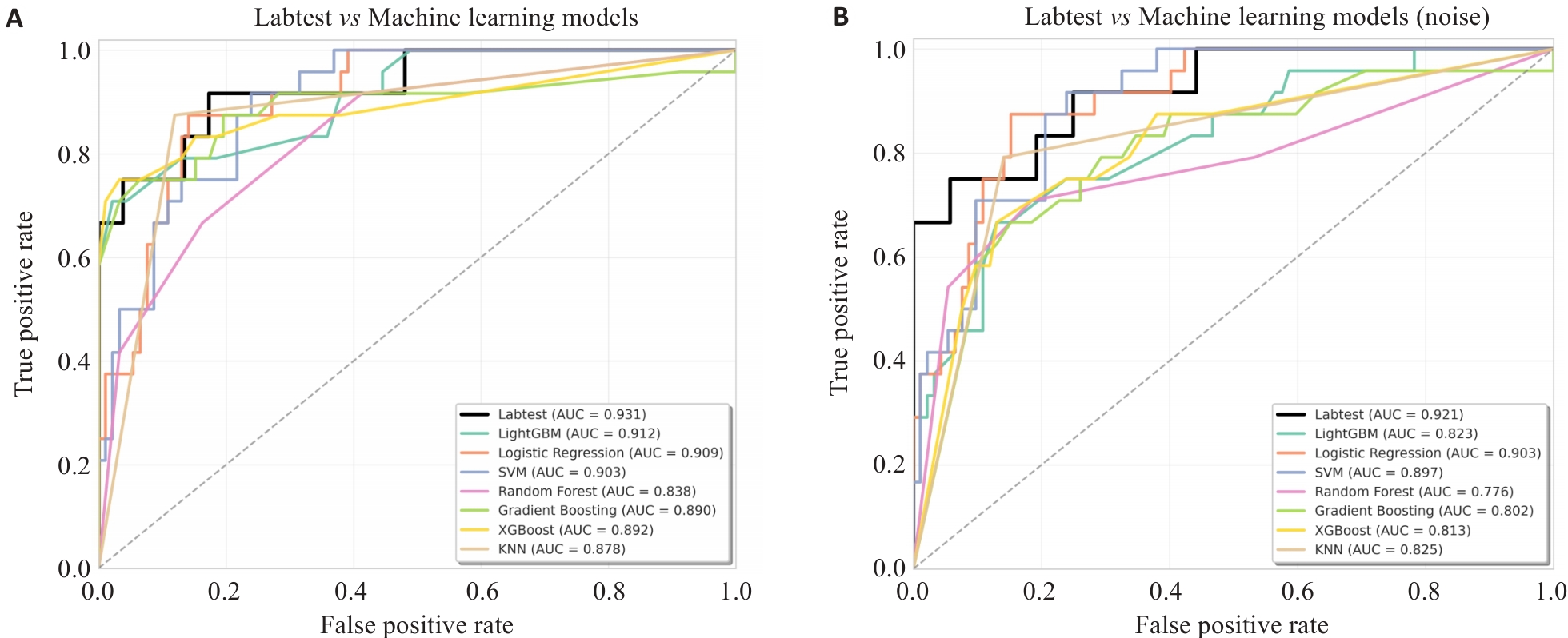
图2 Transformer变体模型与机器学习模型受试者工作特征(ROC)曲线
Fig.2 Receiver operating characteristic (ROC) curves of the Transformer variant model and Machine learning models. A: Clean data. B: Noisy data.
| Models | AUC | ACC | SEN | SPE | |||
|---|---|---|---|---|---|---|---|
| LightGBM | 0.912 | 0.853 | 0.791 | 0.869 | |||
| Logistic regression | 0.909 | 0.801 | 0.875 | 0.783 | |||
| SVM | 0.903 | 0.853 | 0.667 | 0.902 | |||
| Random forest | 0.838 | 0.802 | 0.667 | 0.837 | |||
| Gradient boosting | 0.890 | 0.862 | 0.75 | 0.891 | |||
| XGBoost | 0.892 | 0.853 | 0.792 | 0.870 | |||
| KNN | 0.878 | 0.879 | 0.875 | 0.880 | |||
| Transformer variant | 0.931 | 0.813 | 0.833 | 0.865 |
表2 在干净数据集上机器学习模型和Transformer变体模型的卵巢癌预测性能比较
Tab.2 Comparison of predictive performance for ovarian cancer between the machine learning model and the Transformer variant model on the clean dataset
| Models | AUC | ACC | SEN | SPE | |||
|---|---|---|---|---|---|---|---|
| LightGBM | 0.912 | 0.853 | 0.791 | 0.869 | |||
| Logistic regression | 0.909 | 0.801 | 0.875 | 0.783 | |||
| SVM | 0.903 | 0.853 | 0.667 | 0.902 | |||
| Random forest | 0.838 | 0.802 | 0.667 | 0.837 | |||
| Gradient boosting | 0.890 | 0.862 | 0.75 | 0.891 | |||
| XGBoost | 0.892 | 0.853 | 0.792 | 0.870 | |||
| KNN | 0.878 | 0.879 | 0.875 | 0.880 | |||
| Transformer variant | 0.931 | 0.813 | 0.833 | 0.865 |
| Models | AUC | ACC | SEN | SPE | ||||
|---|---|---|---|---|---|---|---|---|
| LightGBM | 0.823 | 0.759 | 0.750 | 0.761 | ||||
| Logistic regression | 0.903 | 0.793 | 0.875 | 0.772 | ||||
| SVM | 0.897 | 0.853 | 0.667 | 0.902 | ||||
| Random forest | 0.775 | 0.793 | 0.708 | 0.815 | ||||
| Gradient boosting | 0.802 | 0.759 | 0.708 | 0.778 | ||||
| XGBoost | 0.813 | 0.759 | 0.708 | 0.772 | ||||
| KNN | 0.825 | 0.845 | 0.792 | 0.859 | ||||
| Transformer variant | 0.921 | 0.813 | 0.749 | 0.865 |
表3 在带噪声数据集上机器学习模型和Transformer变体模型的卵巢癌预测性能比较
Tab.3 Comparison of predictive performance for ovarian cancer between the machine learning model and the Transformer variant model on the noisy dataset
| Models | AUC | ACC | SEN | SPE | ||||
|---|---|---|---|---|---|---|---|---|
| LightGBM | 0.823 | 0.759 | 0.750 | 0.761 | ||||
| Logistic regression | 0.903 | 0.793 | 0.875 | 0.772 | ||||
| SVM | 0.897 | 0.853 | 0.667 | 0.902 | ||||
| Random forest | 0.775 | 0.793 | 0.708 | 0.815 | ||||
| Gradient boosting | 0.802 | 0.759 | 0.708 | 0.778 | ||||
| XGBoost | 0.813 | 0.759 | 0.708 | 0.772 | ||||
| KNN | 0.825 | 0.845 | 0.792 | 0.859 | ||||
| Transformer variant | 0.921 | 0.813 | 0.749 | 0.865 |
| [1] | Han BF, Zheng RS, Zeng HM, et al. Cancer incidence and mortality in China, 2022[J]. J Natl Cancer Cent, 2024, 4(1): 47-53. doi:10.1016/j.jncc.2024.01.006 |
| [2] | Zeng HM, Zheng RS, Guo YM, et al. Cancer survival in China, 2003-2005: a population-based study[J]. Int J Cancer, 2015, 136(8): 1921-30. doi:10.1002/ijc.29227 |
| [3] | Dochez V, Caillon H, Vaucel E, et al. Biomarkers and algorithms for diagnosis of ovarian cancer: CA125, HE4, RMI and ROMA, a review[J]. J Ovarian Res, 2019, 12(1): 28. doi:10.1186/s13048-019-0503-7 |
| [4] | Guo YY, Jiang TJ, Ouyang LL, et al. A novel diagnostic nomogram based on serological and ultrasound findings for preoperative prediction of malignancy in patients with ovarian masses[J]. Gynecol Oncol, 2021, 160(3): 704-12. doi:10.1016/j.ygyno.2020.12.006 |
| [5] | Madan S, Lentzen M, Brandt J, et al. Transformer models in biomedicine[J]. BMC Med Inform Decis Mak, 2024, 24(1): 214. doi:10.1186/s12911-024-02600-5 |
| [6] | Vaswani A, Shazeer N, Parmar N, et al. Attention is all you need[J]. Adv Neural Inf Process Syst, 2017, 30: 6000-10. |
| [7] | Gorishniy Y, Rubachev I, Babenko A. On embeddings for numerical features in tabular deep learning[J]. Adv Neural Inf Process Syst, 2022, 35: 24991-5004. doi:10.52202/068431-1812 |
| [8] | Somepalli G, Goldblum M, Schwarzschild A, et al. SAINT: improved neural networks for tabular data via row attention and contrastive pre-training[J]. arXiv preprint arXiv: 2106. 01342, 2021. |
| [9] | Wang Z, Sun J. Transtab: learning transferable tabular transformers across tables[J]. Adv Neural Inf Process Syst, 2022, 35: 2902-15. doi:10.52202/068431-0210 |
| [10] | Guyon I, Elisseeff A. An introduction to variable and feature selection[J]. J Mach Learn Res, 2003, 3: 1157-82. |
| [11] | Azur MJ, Stuart EA, Frangakis C, et al. Multiple imputation by chained equations: what is it and how does it work[J]. Int J Methods Psych Res, 2011, 20(1): 40-9. doi:10.1002/mpr.329 |
| [12] | Ioffe S, Szegedy C. Batch normalization: accelerating deep network training by reducing internal covariate shift[J]. Proc Mach Learn Res, 2015, 37: 448-56. |
| [13] | Lin BY, Lee S, Khanna R, et al. Birds have four legs? NumerSense: probing numerical commonsense knowledge of pre-trained language models[C]//Proceedings of the 2020 Conference on Empirical Methods in Natural Language Processing (EMNLP). Online. Stroudsburg, PA, USA: ACL, 2020. DOI:10.18653/v1/2020.emnlp-main.557 . |
| [14] | Ba JL, Kiros JR, Hinton GE. Layer normalization[J]. arXiv preprint arXiv:, 2016. doi:10.48550/arXiv.1607.06450 |
| [15] | Chai YK, Jin S, Hou XW. Highway transformer: self-gating enhanced self-attentive networks[C]//Proceedings of the 58th Annual Meeting of the Association for Computational Linguistics. Online. Stroudsburg, PA, USA: ACL, 2020. DOI:10.18653/v1/2020.acl-main.616 . |
| [16] | Lin TY, Wang YX, Liu XY, et al. A survey of transformers[J]. AI Open, 2022, 3: 111-32. doi:10.1016/j.aiopen.2022.10.001 |
| [17] | Devlin J, Chang MW, Lee K, et al. BERT: pre-training of deep bidirectional transformers for language understanding[C] //Proceedings of the Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, 2019. doi:10.18653/v1/n19-1423 |
| [18] | Zhou ZW, Rahman Siddiquee MM, Tajbakhsh N, et al. UNet++: a nested U-Net architecture for medical image segmentation[C]//Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support. Cham: Springer, 2018: 3-11. doi:10.1007/978-3-030-00889-5_1 |
| [19] | Campbell TW, Wilson MP, Roder H, et al. Predicting prognosis in COVID-19 patients using machine learning and readily available clinical data[J]. Int J Med Inform, 2021, 155: 104594. doi:10.1016/j.ijmedinf.2021.104594 |
| [20] | Ke G, Meng Q, Finley T, et al. LightGBM: a highly efficient gradient boosting decision tree[C] // Advances in Neural Information Processing Systems, 2017:30. |
| [21] | Cox DR. The regression analysis of binary sequences[J]. J R Stat Soc Ser B Stat Methodol, 1958, 20(2): 215-32. doi:10.1111/j.2517-6161.1958.tb00292.x |
| [22] | Cortes C, Vapnik V. Support-vector networks[J]. Mach Learn, 1995, 20(3): 273-97. doi:10.1007/bf00994018 |
| [23] | Breiman L. Random forests[J]. Mach Learn, 2001, 45(1): 5-32. doi:10.1023/a:1010933404324 |
| [24] | Friedman JH. Greedy function approximation: a gradient boosting machine[J]. Ann Statist, 2001, 29(5): 1189-232. doi:10.1214/aos/1013203451 |
| [25] | Chen TQ, Guestrin C. XGBoost: a scalable tree boosting system[C]//Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. San Francisco California USA. ACM, 2016: 785-94. DOI:10.1145/2939672.2939785 . |
| [26] | Cover T, Hart P. Nearest neighbor pattern classification[J]. IEEE Trans Inform Theory, 1967, 13(1): 21-7. doi:10.1109/tit.1967.1053964 |
| [27] | Lundberg SM, Lee SI. A unified approach to interpreting model predictions[J]. Adv Neural Inf Process Syst, 2017, 30: 4768-77. |
| [28] | Charkhchi P, Cybulski C, Gronwald J, et al. CA125 and ovarian cancer: a comprehensive review[J]. Cancers, 2020, 12(12): 3730. doi:10.3390/cancers12123730 |
| [29] | Yang JN, Jin Y, Cheng SS, et al. Clinical significance for combined coagulation indexes in epithelial ovarian cancer prognosis[J]. J Ovarian Res, 2021, 14(1): 106. doi:10.1186/s13048-021-00858-1 |
| [30] | Bishara S, Griffin M, Cargill A, et al. Pre-treatment white blood cell subtypes as prognostic indicators in ovarian cancer[J]. Eur J Obstet Gynecol Reprod Biol, 2008, 138(1): 71-5. doi:10.1016/j.ejogrb.2007.05.012 |
| [31] | Mu J, Wu Y, Jiang C, et al. Progress in applicability of scoring systems based on nutritional and inflammatory parameters for ovarian cancer[J]. Front Nutr, 2022, 9: 809091. doi:10.3389/fnut.2022.809091 |
| [1] | 陈博湧, 汪新怡, 赵新新, 宋婷, 李永宝. 基于Swin-ResViT网络的低质量动态cine-MR至高质量定位MR图像实时生成研究[J]. 南方医科大学学报, 2026, 46(4): 929-938. |
| [2] | 卢学麒, 陈华元, 吴秋岑, 温耀棋, 刘国光, 陈超敏. 一种基于深度特征融合的可解释性12导联心电图自动诊断模型研究[J]. 南方医科大学学报, 2026, 46(1): 208-218. |
| [3] | 吴秋岑, 卢学麒, 温耀棋, 洪永, 吴煜良, 陈超敏. II导联心电图中心肌梗死检测与定位:基于多尺度残差模块融合改进通道注意力模型[J]. 南方医科大学学报, 2025, 45(8): 1777-1790. |
| [4] | 郑子瑜, 杨夏颖, 吴圣杰, 张诗婕, 吕国荣, 柳培忠, 王珺, 何韶铮. 多特征融合的产时超声胎方位识别模型[J]. 南方医科大学学报, 2025, 45(7): 1563-1570. |
| [5] | 梁业东, 朱雄峰, 黄美燕, 张文聪, 郭翰宇, 冯前进. CRAKUT:融合对比区域注意力机制与临床先验知识的U-Transformer用于放射学报告生成[J]. 南方医科大学学报, 2025, 45(6): 1343-1352. |
| [6] | 郭晓娟, 杜瑞娟, 陈丽平, 郭克磊, 周彪, 卞华, 韩立. WW结构域E3泛素连接酶1调控卵巢癌肿瘤微环境中的免疫浸润[J]. 南方医科大学学报, 2025, 45(5): 1063-1073. |
| [7] | 渠梦, 傅蓉. ResLSTM-TemporalSE:多导联心电信号的自动分类[J]. 南方医科大学学报, 2025, 45(12): 2708-2717. |
| [8] | 陈一镠, 马民, 苏燃, 朱寅宾, 冯晴, 罗嘉丽, 冯伟峰, 颜显欣. 理冲消癥颗粒通过上调ANT3介导的线粒体凋亡增强小鼠卵巢癌移植瘤对顺铂的敏感性[J]. 南方医科大学学报, 2025, 45(11): 2309-2319. |
| [9] | 谢辉荣, 胡潮滨, 梁国华, 韩红喆, 黄牧, 冯前进. 大学生心理压力智能评估:基于融合文本与影像的多模态模型的设计及验证[J]. 南方医科大学学报, 2025, 45(11): 2504-2510. |
| [10] | 卢梓涵, 黄方俊, 蔡光瑶, 刘继红, 甄鑫. 针对缺失实验室指标多约束表征学习的卵巢癌鉴别方法[J]. 南方医科大学学报, 2025, 45(1): 170-178. |
| [11] | 贺亚迪, 周炫汝, 金锦辉, 宋婷. 基于PE-CycleGAN网络的鼻咽癌自适应放疗CBCT-sCT生成研究[J]. 南方医科大学学报, 2025, 45(1): 179-186. |
| [12] | 方威扬, 肖慧, 王爽, 林晓明, 陈超敏. 基于MRI影像和临床参数特征融合的深度学习模型预测术前肝细胞癌的细胞角蛋白19状态[J]. 南方医科大学学报, 2024, 44(9): 1738-1751. |
| [13] | 欧嘉志, 詹长安, 杨丰. 一维卷积神经网络的自编码癫痫发作异常检测模型[J]. 南方医科大学学报, 2024, 44(9): 1796-1804. |
| [14] | 汪辰, 蒙铭强, 李明强, 王永波, 曾栋, 边兆英, 马建华. 基于双域Transformer耦合特征学习的CT截断数据重建模型[J]. 南方医科大学学报, 2024, 44(5): 950-959. |
| [15] | 龙楷兴, 翁丹仪, 耿 舰, 路艳蒙, 周志涛, 曹 蕾. 基于多模态多示例学习的免疫介导性肾小球疾病自动分类方法[J]. 南方医科大学学报, 2024, 44(3): 585-593. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||