南方医科大学学报 ›› 2026, Vol. 46 ›› Issue (4): 816-824.doi: 10.12122/j.issn.1673-4254.2026.04.10
• • 上一篇
李梓源1( ), 苏小宇1, 田智涵1, 晁兴亮1, 王咏2, 马度芳2(
), 苏小宇1, 田智涵1, 晁兴亮1, 王咏2, 马度芳2( )
)
收稿日期:2025-10-20
出版日期:2026-04-20
发布日期:2026-04-24
通讯作者:
马度芳
E-mail:liziyuan073721@163.com;madufang@163.com
作者简介:李梓源,在读硕士研究生,E-mail: liziyuan073721@163.com
基金资助:
Ziyuan LI1( ), Xiaoyu SU1, Zhihan TIAN1, Xingliang CHAO1, Yong WANG2, Dufang MA2(
), Xiaoyu SU1, Zhihan TIAN1, Xingliang CHAO1, Yong WANG2, Dufang MA2( )
)
Received:2025-10-20
Online:2026-04-20
Published:2026-04-24
Contact:
Dufang MA
E-mail:liziyuan073721@163.com;madufang@163.com
Supported by:摘要:
目的 联合转录组学和代谢组学分析癌症恶病质中氨基酸(谷氨酰胺)是否通过诱导自噬促进肌肉萎缩。 方法 将20只BALB/c裸鼠随机分为正常组、模型组(n=10)。采用皮下移植CT-26细胞诱导恶病质模型,测量BALB/c裸鼠去瘤体质量、抓力/体质量比值、苏木素-伊红(HE染色)观察肌纤维横径,检测肌肉萎缩相关蛋白(MuRF1/Atrogin-1)。转录组学分析鉴定差异表达基因(DEGs),代谢组学分析差异代谢物(DEMs),绘制代谢通路图谱。透射电镜(TEM)观察自噬小体及腓肠肌形态、Western blotting检测自噬标志物(ULK1、LC3、P62)及信号通路蛋白(AMPK、FOXO3a及mTOR等)。经Gln+AMPK抑制剂处理的C2C12成肌细胞验证了关键实验结果。 结果 与对照组相比,模型组无瘤体质量(P<0.01)、抓力/体质量比值(P<0.0001)下降,肌纤维面积减少(P<0.05)以及Atrogin-1(P<0.01)、MuRF1(P<0.05)水平升高。DEGs主要富集于精氨酸/脯氨酸代谢、AMPK、mTOR、自噬及FOXO信号通路。代谢组学分析显示,恶病质肌肉中谷氨酰胺和谷氨酸水平显著升高。体内实验发现恶病质小鼠肌肉中自噬小体数量增加(P<0.01),肌纤维模糊断裂。AMPK、FOXO3a通路蛋白升高(P<0.05),同时ULK1(P<0.05)、LC3II/I蛋白(P<0.0001)表达水平显著上调,mTOR通路蛋白及P62蛋白下调(P<0.01)。体外实验表明,谷氨酰胺可促进C2C12细胞自噬进程,并促进AMPK、FOXO通路,抑制mTOR通路,而加入AMPK抑制剂则能阻断这些效应。 结论 谷氨酰胺通过激活AMPK/FOXO3a信号通路并抑制mTOR信号通路促进自噬,导致癌症恶病质肌肉萎缩。
李梓源, 苏小宇, 田智涵, 晁兴亮, 王咏, 马度芳. 谷氨酰胺诱导的自噬加剧恶病质裸鼠的肌肉萎缩:基于多组学分析[J]. 南方医科大学学报, 2026, 46(4): 816-824.
Ziyuan LI, Xiaoyu SU, Zhihan TIAN, Xingliang CHAO, Yong WANG, Dufang MA. Glutamine-induced autophagy exacerbates muscle atrophy in cachectic nude mice: a multi-omics analysis[J]. Journal of Southern Medical University, 2026, 46(4): 816-824.
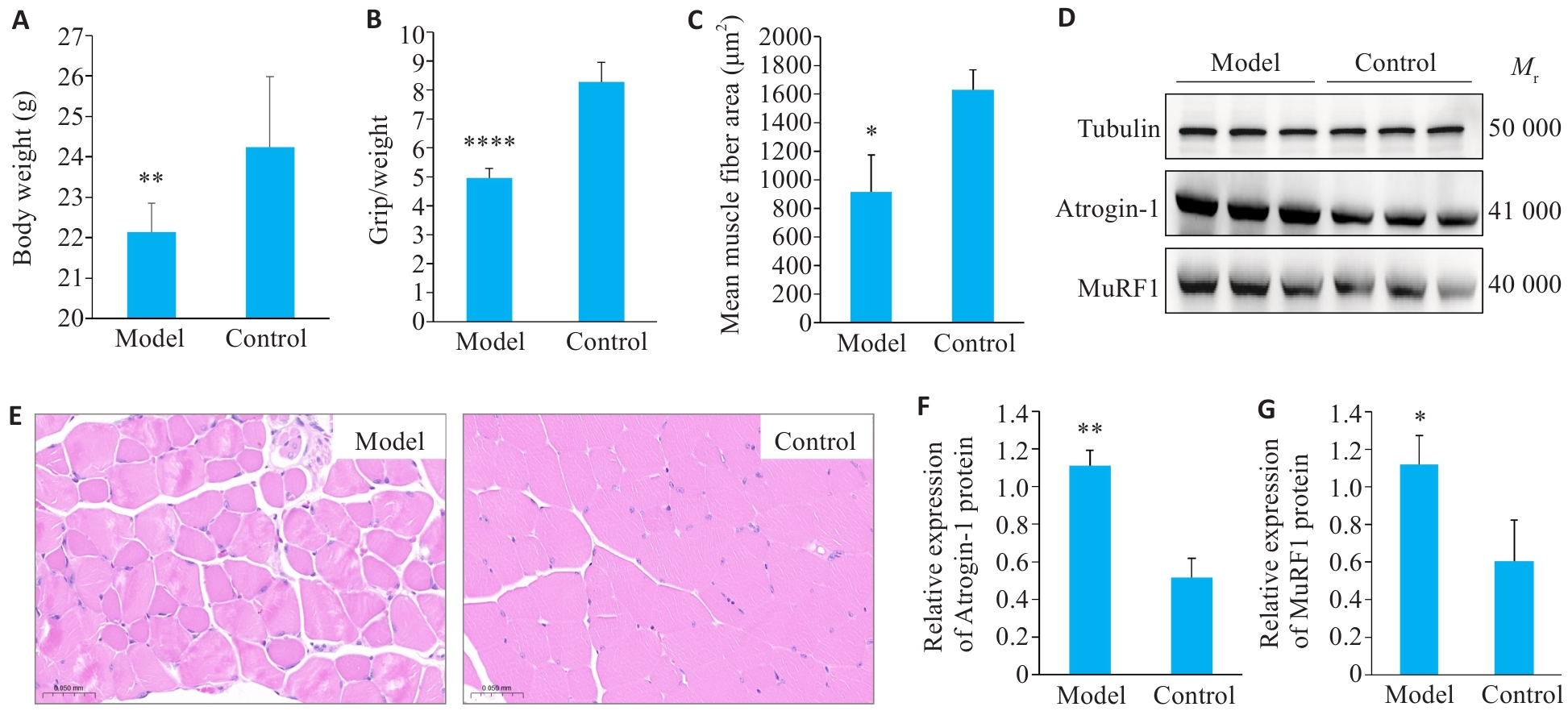
图1 小鼠体质量及肌肉萎缩情况
Fig. 1 Body weight and muscle atrophy in mice. A: Carcass weight of the mice (n=10). B: Ratio of grip strength to body weight (n=10). C: Muscle fiber cross-sectional area. D: Western blotting of atrogin-1 and MuRF1. E: HE staining of the gastrocnemius muscle (Scale bar=50 μm). F, G: Quantification of atrogin-1 and MuRF1 protein expressions (n=3). Data are presented as Mean±SD. *P<0.05, **P<0.01, ****P<0.0001 vs Control group.
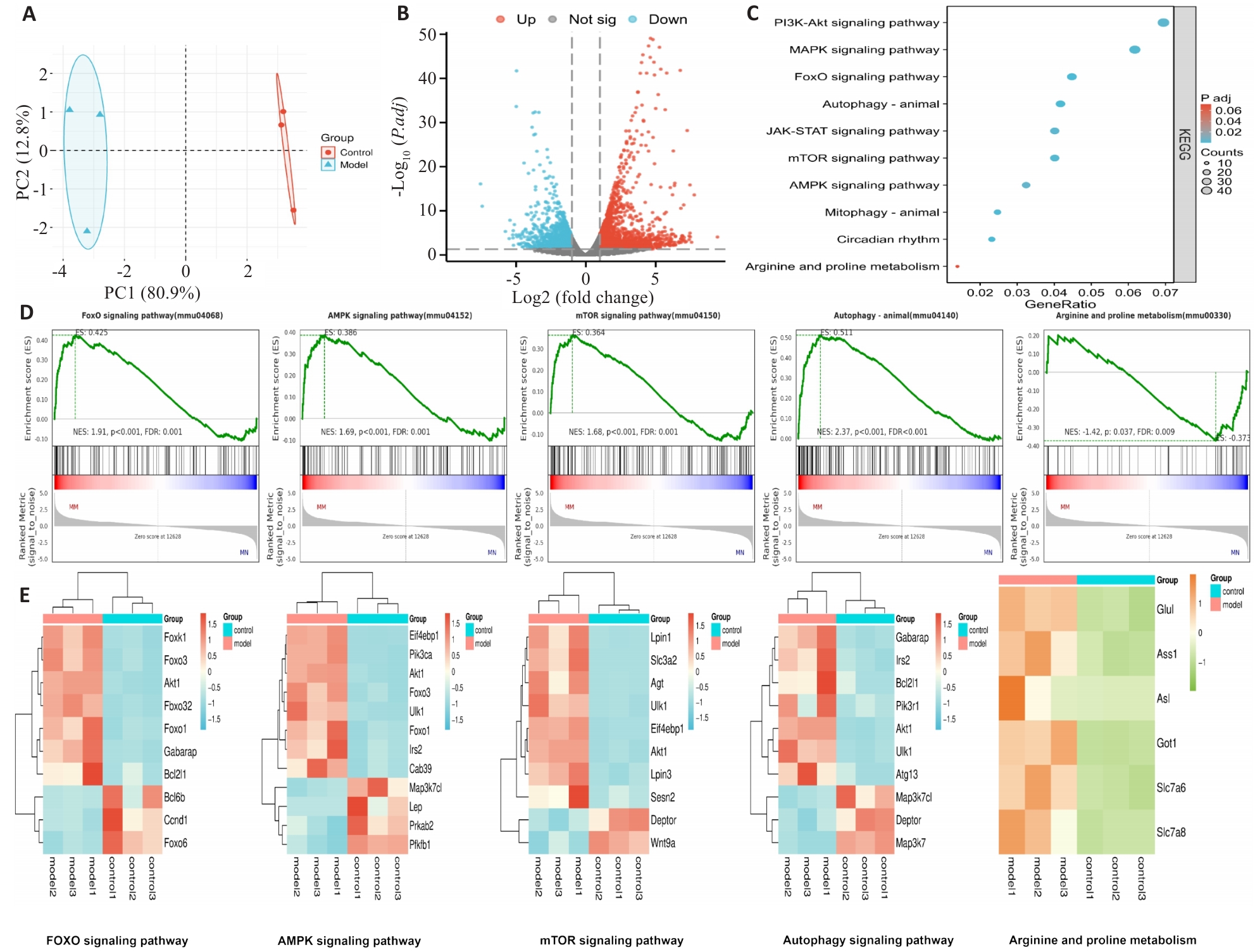
图2 转录组学分析
Fig.2 Transcriptomics analysis. A: Expression distribution of DEGs. B: Volcano plot of the DEGs. C: KEGG pathway enrichment of the DEGs. D: GSEA of FOXO, AMPK, mTOR, autophagy and arginine-proline metabolism pathways (model vs control). E: Heatmap of the DEGs in the indicated pathways.
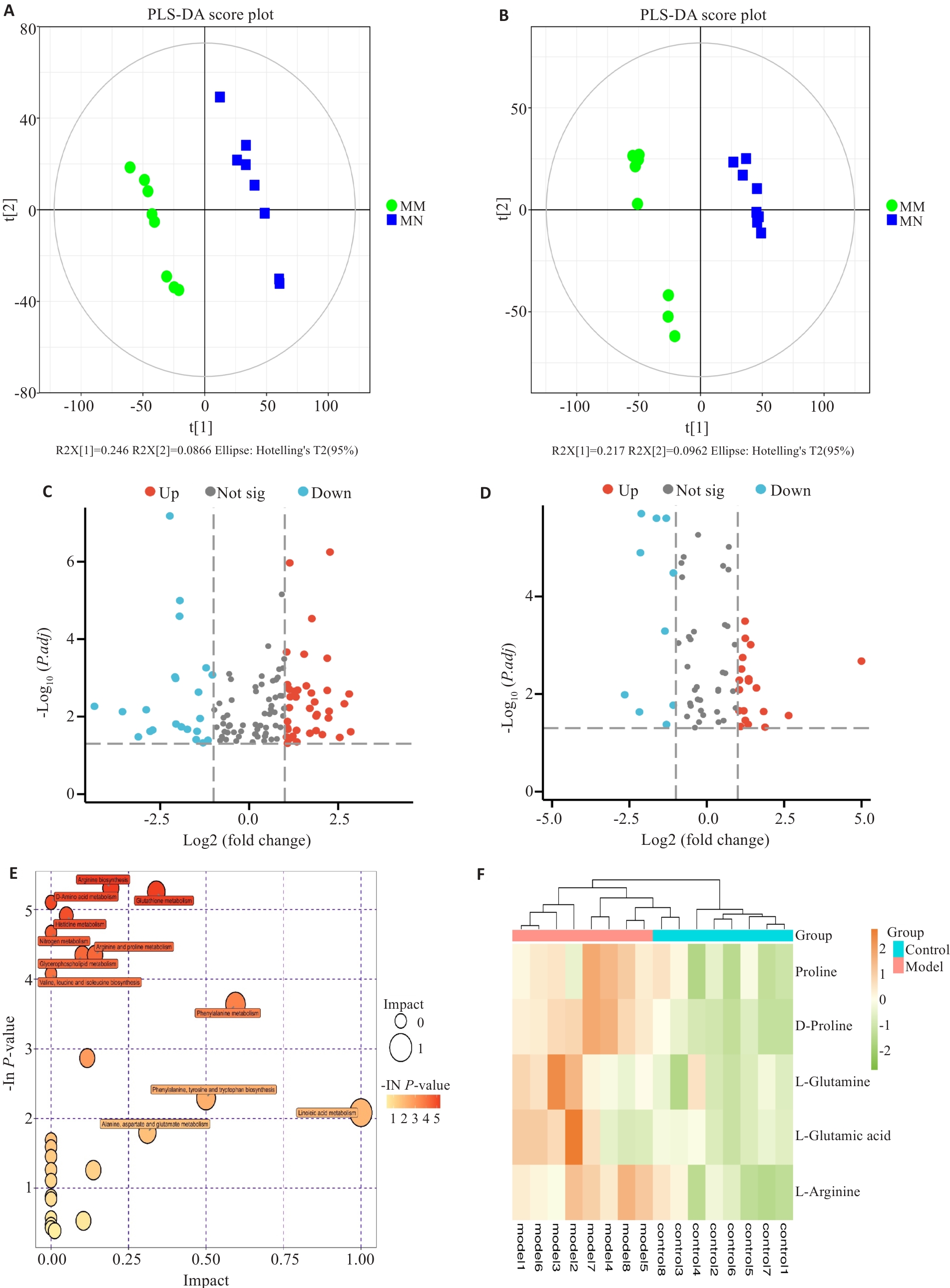
图3 代谢组学分析
Fig.3 Metabolomics analysis. A: PLS-DA scores (negative ion mode). B: PLS-DA scores (positive ion mode). C: Volcano plot of the DEMs (negative ion mode). D: Volcano plot of the DEMs (positive ion mode). E: KEGG pathway enrichment (positive ion mode). F: Heatmap of arginine-proline metabolites (positive ion mode).
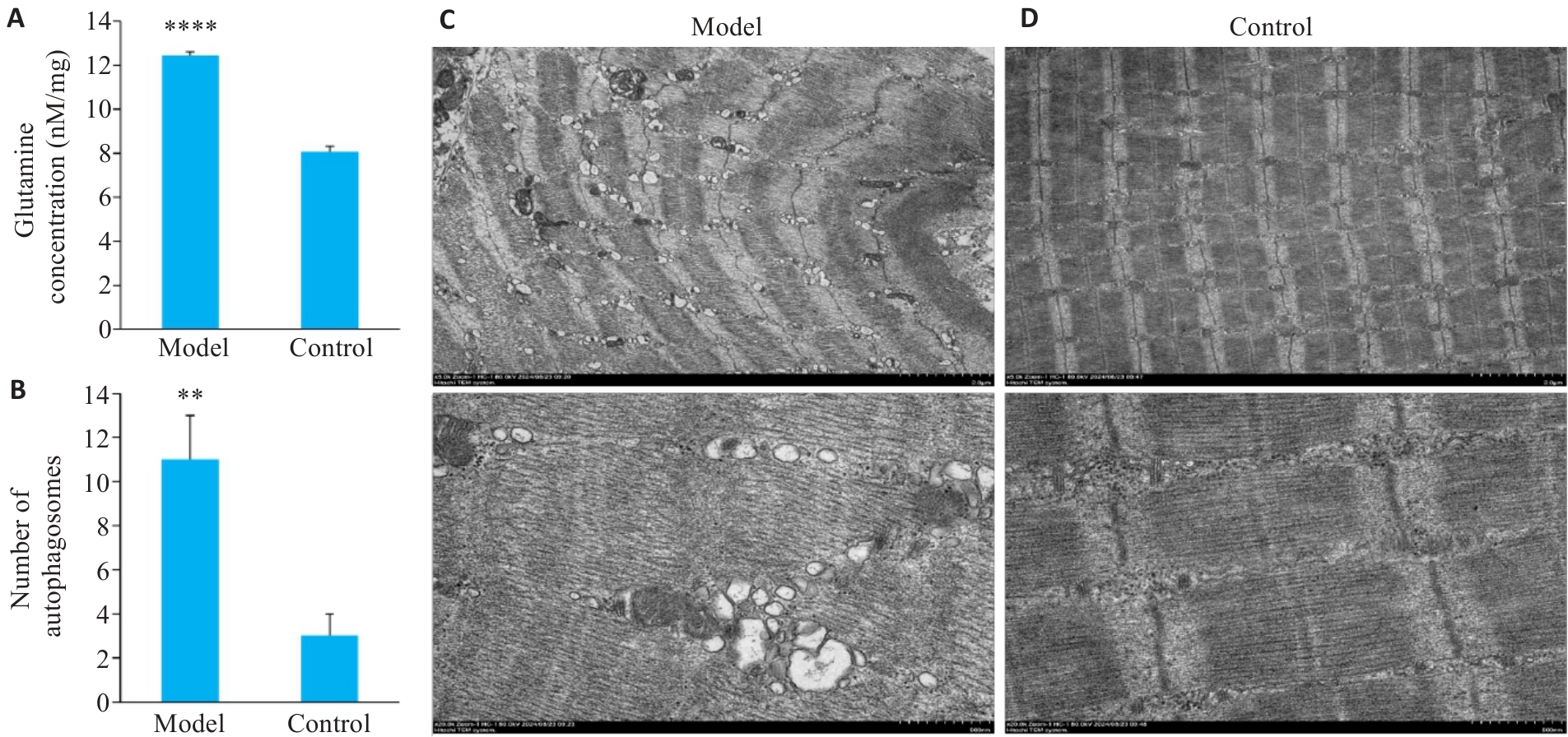
图4 谷氨酰胺浓度及透射电镜检测
Fig.4 Glutamine concentration and TEM detection. A: Comparison of glutamine concentrations between the model and control groups. B: Quantification of autophagosomes. C, D: TEM images in the model and control groups. The upper and lower panels show muscle tissue at ×5 000 and ×20 000 magnification, respectively. Data are presented as Mean±SD (n=3). **P<0.01, ****P<0.0001 vs Control group.
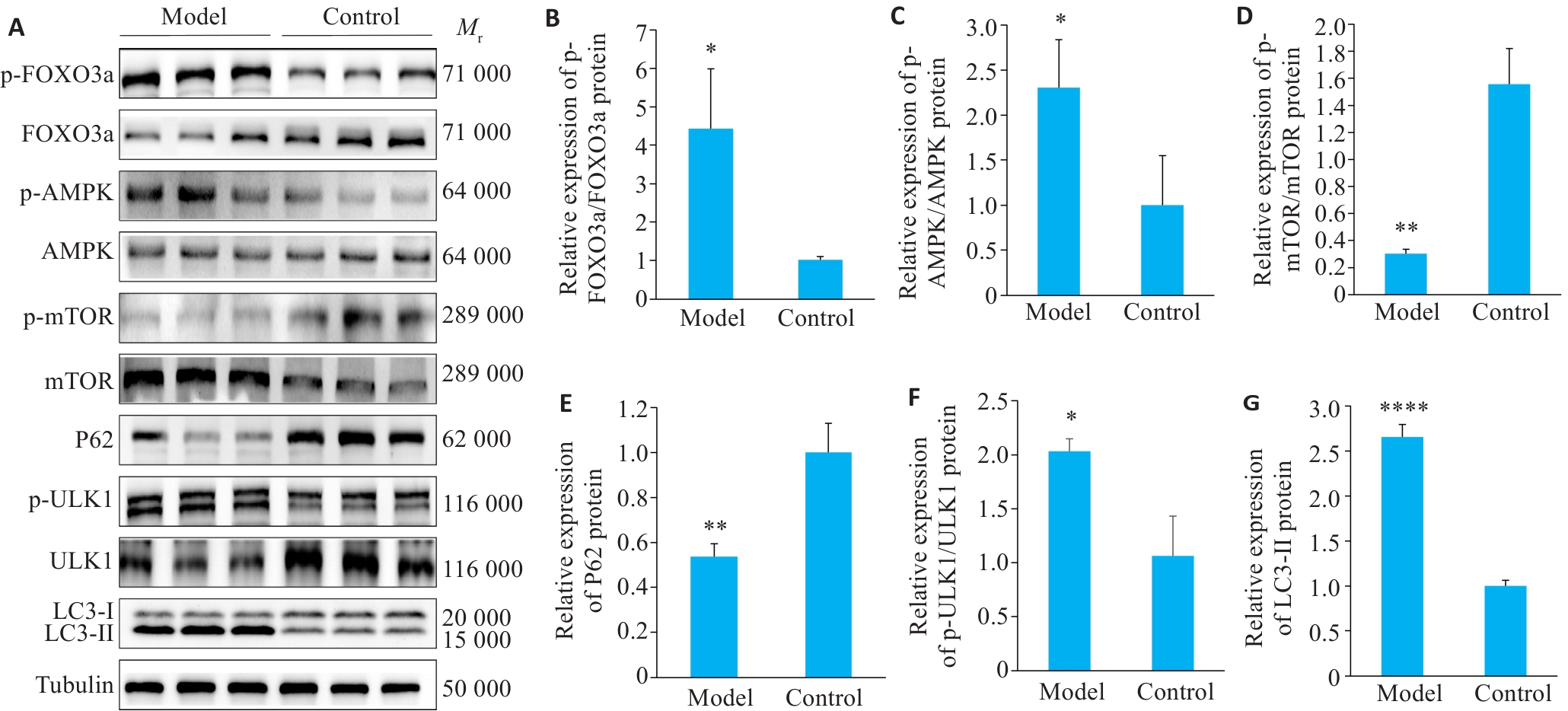
图5 肌肉自噬蛋白表达情况
Fig.5 Autophagy occurrences in skeletal muscle. A: Western blotting of p-FOXO3a, FOXO3a, p-mTOR, mTOR, p-AMPK, AMPK, P62, p-ULK1, ULK1 and LC3-II/I. B-G: Quantification of p-FOXO3a/FOXO3a, p-mTOR/mTOR, p-AMPK/AMPK, p-ULK1/ULK1, P62 and LC3-II/I protein expression. Data are presented as Mean±SD (n=3). *P<0.05, **P<0.01, ****P<0.0001 vs Control group.
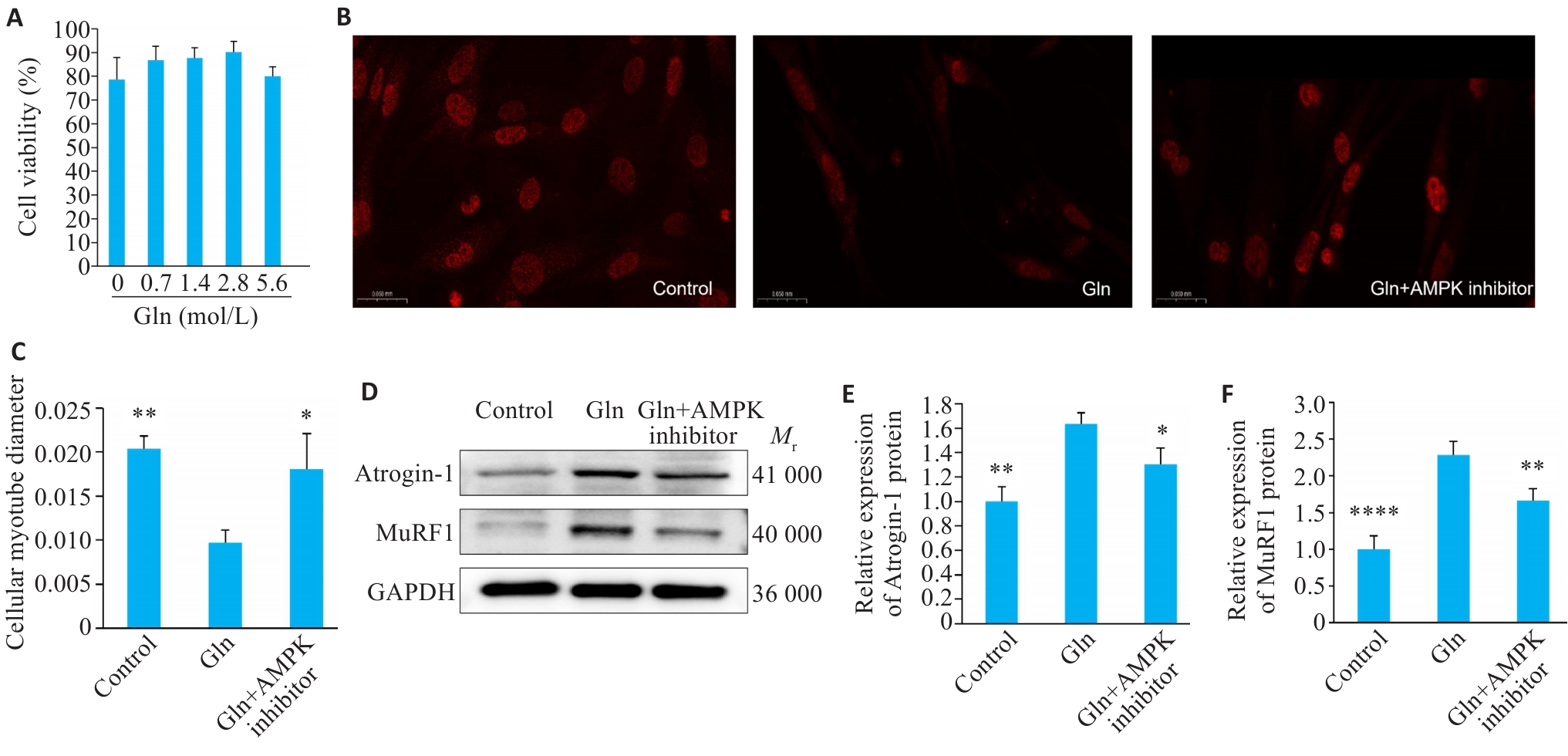
图6 谷氨酰胺浓度的确定及肌管萎缩情况
Fig.6 Determination of glutamine concentration and muscle tube atrophy. A: Cell viability vs Gln concentration. B: Myotube staining. C: Myotube diameter (Scale bar=50 μm). D: Western blotting of MuRF1 and Atrogin-1. E,F: Quantification of MuRF1 and atrogin-1 protein expression. Data are presented as Mean±SD (n=3). *P<0.05, **P<0.01, ****P<0.0001 vs Gln group.
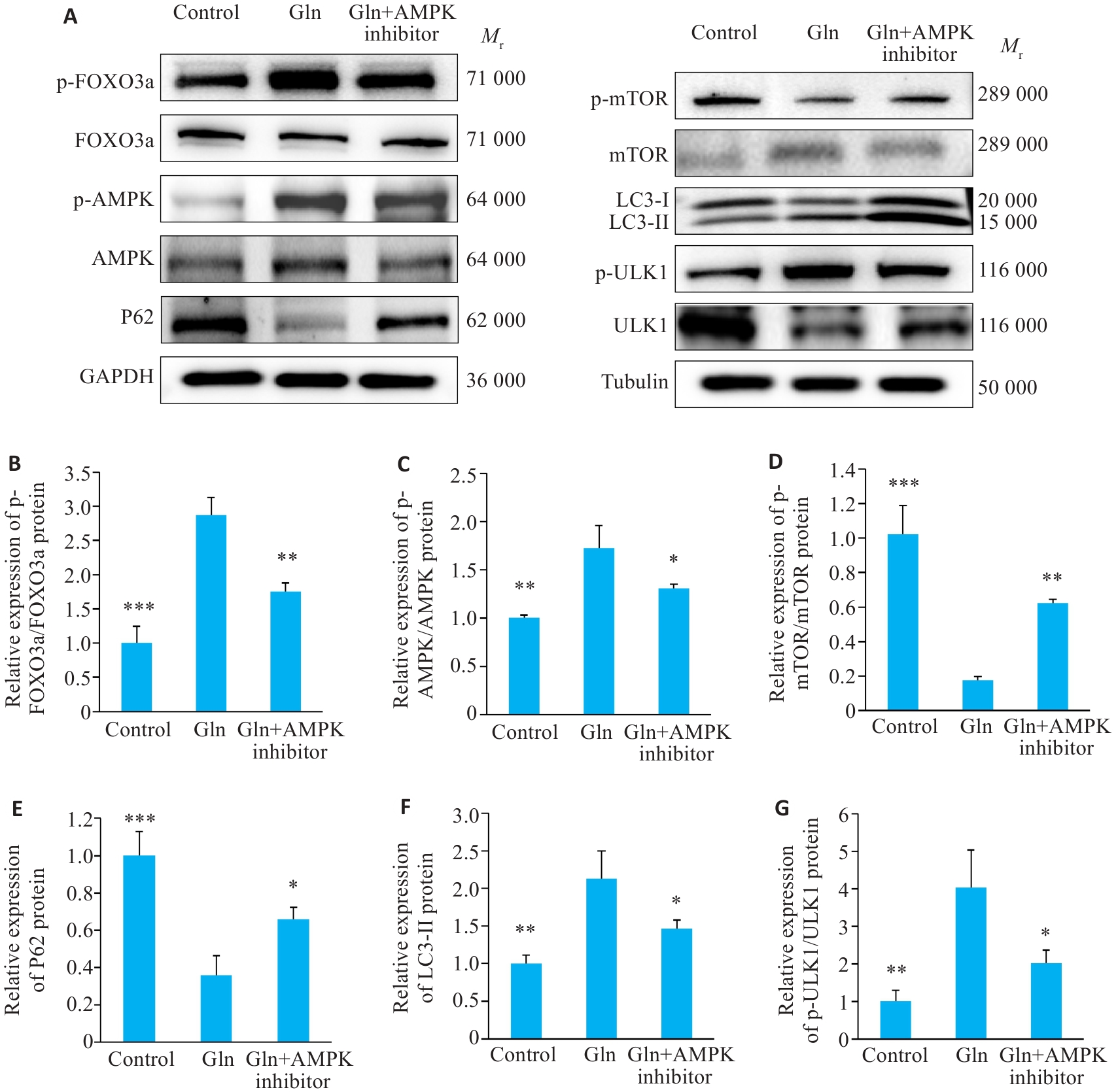
图7 C2C12肌管实验验证
Fig.7 Experimental validation in C2C12 myotubes. A: Western blotting of p-FOXO3a, FOXO3a, p-mTOR, mTOR, p-AMPK, AMPK, P62, p-ULK1, ULK1, LC3-II/I. B-G: Quantification of p-FOXO3a/FOXO3a, p-mTOR/mTOR, p-AMPK/AMPK, p-ULK1/ULK1, P62 and LC3-II/I protein expression. Data are presented as Mean±SD (n=3). *P<0.05, **P<0.01, ***P<0.001 vs Gln group.
| [1] | Frontera WR, Ochala J. Skeletal muscle: a brief review of structure and function[J]. Calcif Tissue Int, 2015, 96(3): 183-95. doi:10.1007/s00223-014-9915-y |
| [2] | Porporato PE. Understanding Cachexia as a cancer metabolism syndrome[J]. Oncogenesis, 2016, 5(2): e200. doi:10.1038/oncsis.2016.3 |
| [3] | Wang D, Duan JJ, Guo YF, et al. Targeting the glutamine-arginine-proline metabolism axis in cancer[J]. J Enzyme Inhib Med Chem, 2024, 39: 2367129. doi:10.1080/14756366.2024.2367129 |
| [4] | Bravo-San Pedro JM, Kroemer G, Galluzzi L. Autophagy and mitophagy in cardiovascular disease[J]. Circ Res, 2017, 120(11): 1812-24. doi:10.1161/circresaha.117.311082 |
| [5] | Salazar G, Cullen A, Huang JW, et al. SQSTM1/p62 and PPARGC1A/PGC-1alpha at the interface of autophagy and vascular senescence[J]. Autophagy, 2020, 16(6): 1092-110. doi:10.1080/15548627.2019.1659612 |
| [6] | Aversa Z, Pin F, Lucia S, et al. Autophagy is induced in the skeletal muscle of cachectic cancer patients[J]. Sci Rep, 2016, 6: 30340. doi:10.1038/srep30340 |
| [7] | Dalle Pezze P, Karanasios E, Kandia V, et al. ATG13 dynamics in nonselective autophagy and mitophagy: insights from live imaging studies and mathematical modeling[J]. Autophagy, 2021, 17(5): 1131-41. doi:10.1080/15548627.2020.1749401 |
| [8] | Yang ZB, Chu BZ, Tu Y, et al. Dual inhibitors of DNMT and HDAC remodels the immune microenvironment of colorectal cancer and enhances the efficacy of anti-PD-L1 therapy[J]. Pharmacol Res, 2024, 206: 107271. doi:10.1016/j.phrs.2024.107271 |
| [9] | Cui PF, Shao W, Huang CH, et al. Metabolic derangements of skeletal muscle from a murine model of glioma Cachexia [J]. Skeletal Muscle, 2019, 9(1): 3. doi:10.1186/s13395-018-0188-4 |
| [10] | Zhang SZ, You XH, Zheng YW, et al. The UBE2C/CDH1/DEPTOR axis is an oncogene and tumor suppressor cascade in lung cancer cells[J]. J Clin Investig, 2023, 133(4): e162434. doi:10.1172/jci162434 |
| [11] | Zhou ZH, Wang HY, Zhang XW, et al. Defective autophagy contributes to endometrial epithelial-mesenchymal transition in intrauterine adhesions[J]. Autophagy, 2022, 18(10): 2427-42. doi:10.1080/15548627.2022.2038994 |
| [12] | Chauhan AS, Liu XG, Jing J, et al. STIM2 interacts with AMPK and regulates calcium-induced AMPK activation[J]. FASEB J, 2019, 33(2): 2957-70. doi:10.1096/fj.201801225r |
| [13] | Sakiyama T, Musch MW, Ropeleski MJ, et al. Glutamine increases autophagy under basal and stressed conditions in intestinal epithelial cells[J]. Gastroenterology, 2009, 136(3): 924-32. e2. doi:10.1053/j.gastro.2008.12.002 |
| [14] | Penna F, Baccino FM, Costelli P. Coming back: autophagy in Cachexia [J]. Curr Opin Clin Nutr Metab Care, 2014, 17(3): 241-6. doi:10.1097/mco.0000000000000048 |
| [15] | Penna F, Costamagna D, Pin F, et al. Autophagic degradation contributes to muscle wasting in cancer Cachexia [J]. Am J Pathol, 2013, 182(4): 1367-78. doi:10.1016/j.ajpath.2012.12.023 |
| [16] | Vander Heiden MG, Cantley LC, Thompson CB. Understanding the Warburg effect: the metabolic requirements of cell proliferation[J]. Science, 2009, 324(5930): 1029-33. doi:10.1126/science.1160809 |
| [17] | Do LK, Lee HM, Ha YS, et al. Amino acids in cancer: Understanding metabolic plasticity and divergence for better therapeutic approaches[J]. Cell Rep, 2025, 44(4): 115529. doi:10.1016/j.celrep.2025.115529 |
| [18] | Penna F, Ballarò R, Martinez-Cristobal P, et al. Autophagy exacerbates muscle wasting in cancer Cachexia and impairs mitochondrial function[J]. J Mol Biol, 2019, 431(15): 2674-86. doi:10.1016/j.jmb.2019.05.032 |
| [19] | Feng YY, Chen Y, Wu XY, et al. Interplay of energy metabolism and autophagy[J]. Autophagy, 2024, 20(1): 4-14. doi:10.1080/15548627.2023.2247300 |
| [20] | Li YJ, Chen YY. AMPK and autophagy[M]//Autophagy: Biology and Diseases. Singapore: Springer Singapore, 2019: 85-108. doi:10.1007/978-981-15-0602-4_4 |
| [21] | Bloemberg D, Quadrilatero J. Autophagy, apoptosis, and mito-chondria: molecular integration and physiological relevance in skeletal muscle[J]. Am J Physiol Cell Physiol, 2019, 317(1): C111-30. doi:10.1152/ajpcell.00261.2018 |
| [22] | Zhao JH, Brault JJ, Schild A, et al. FoxO3 coordinately activates protein degradation by the autophagic/lysosomal and proteasomal pathways in atrophying muscle cells[J]. Cell Metab, 2007, 6(6): 472-83. doi:10.1016/j.cmet.2007.11.004 |
| [23] | White JP, Gao S, Puppa MJ, et al. Testosterone regulation of Akt/mTORC1/FoxO3a signaling in skeletal muscle[J]. Mol Cell Endocrinol, 2013, 365(2): 174-86. doi:10.1016/j.mce.2012.10.019 |
| [24] | Peixoto da Silva S, Santos JMO, Costa e Silva MP, et al. Cancer Cachexia and its pathophysiology: links with sarcopenia, anorexia and Asthenia [J]. J Cachexia Sarcopenia Muscle, 2020, 11(3): 619-35. doi:10.1002/jcsm.12528 |
| [25] | Salabei JK, Lorkiewicz PK, Holden CR, et al. Glutamine regulates cardiac progenitor cell metabolism and proliferation[J]. Stem Cells, 2015, 33(8): 2613-27. doi:10.1002/stem.2047 |
| [26] | Leone RD, Zhao L, Englert JM, et al. Glutamine blockade induces divergent metabolic programs to overcome tumor immune evasion[J]. Science, 2019, 366(6468): 1013-21. doi:10.1126/science.aav2588 |
| [27] | Shen YM, Zhang YH, Li WD, et al. Glutamine metabolism: from proliferating cells to cardiomyocytes[J]. Metabolism, 2021, 121: 154778. doi:10.1016/j.metabol.2021.154778 |
| [28] | Yang LF, Venneti S, Nagrath D. Glutaminolysis: a hallmark of cancer metabolism[J]. Annu Rev Biomed Eng, 2017, 19: 163-94. doi:10.1146/annurev-bioeng-071516-044546 |
| [29] | van der Vos KE, Eliasson P, Proikas-Cezanne T, et al. Modulation of glutamine metabolism by the PI(3)K-PKB-FOXO network regulates autophagy[J]. Nat Cell Biol, 2012, 14(8): 829-37. doi:10.1038/ncb2536 |
| [30] | van der Vos KE, Coffer PJ. Glutamine metabolism links growth factor signaling to the regulation of autophagy[J]. Autophagy, 2012, 8(12): 1862-4. doi:10.4161/auto.22152 |
| [31] | Zhang YY, Huang ZQ, Han WT, et al. Glutamine suppresses senescence and promotes autophagy through glycolysis inhibition-mediated AMPKα lactylation in intervertebral disc degeneration[J]. Commun Biol, 2024, 7: 325. doi:10.1038/s42003-024-06000-3 |
| [1] | 雷艳萍, 宋嘉晟, 徐乐吾, 刘睿, 赵岳. 麦冬皂苷D通过激活β-catenin/FUNDC1/线粒体自噬轴减轻阿霉素诱导的小鼠心肌肥厚[J]. 南方医科大学学报, 2026, 46(4): 803-815. |
| [2] | 李彩霞, 崔立华, 曹婕, 樊宇星, 周雪颖, 张淑坤, 左艳洁. 活血清解灵通过调控Sirt1-自噬信号通路减轻胆管结扎诱导的大鼠肝纤维化[J]. 南方医科大学学报, 2026, 46(3): 513-522. |
| [3] | 张语洋, 申颖, 佟欣雨, 段宇魁, 罗云娜, 郭文奇. 水飞蓟宾通过诱导细胞自噬及凋亡双重机制抑制兔青光眼术后纤维化[J]. 南方医科大学学报, 2026, 46(3): 666-674. |
| [4] | 乔通, 尹林, 张可妮, 牛民主, 黄菊, 耿志军, 李静, 胡建国. 茯苓新酸A通过调节AMPK/mTOR介导的自噬来减轻葡聚糖硫酸钠诱导的小鼠结肠炎[J]. 南方医科大学学报, 2026, 46(1): 131-140. |
| [5] | 徐嘉艺, 杨迪, 臧开来, 褚孟恩, 赵庆瑶, 李晴, 鲁森, 陈修丽, 李宁. EVA1A过表达通过调节脂质代谢和促进脂滴自噬改善非酒精性脂肪肝[J]. 南方医科大学学报, 2026, 46(1): 150-158. |
| [6] | 林心君, 何昱霖, 施红, 刘佳绣, 胡海霞. 石斛合剂通过调控Sirt3介导的线粒体自噬通路缓解大鼠糖尿病心肌病[J]. 南方医科大学学报, 2026, 46(1): 47-54. |
| [7] | 许嘉蓉, 黄傲, 丁祉恺, 鲍宇, 赵仓焕, 蔡文智. 子午流注针法对失眠大鼠Glu/GABA-Gln代谢环路异常的调节作用[J]. 南方医科大学学报, 2025, 45(8): 1616-1624. |
| [8] | 李玮怡, 江露, 张宗星, 陈丹, 包卓玛, 黄丽, 袁林. 强骨康疏方通过抑制HIF-1α/BNIP3自噬信号通路减少类风湿性关节炎大鼠的破骨细胞分化[J]. 南方医科大学学报, 2025, 45(7): 1389-1396. |
| [9] | 王心恒, 邵小涵, 李童童, 张璐, 杨勤军, 叶卫东, 童佳兵, 李泽庚, 方向明. 平喘宁方通过调控HMGB1/Beclin-1轴介导的自噬改善患寒哮证大鼠的气道炎症[J]. 南方医科大学学报, 2025, 45(6): 1153-1162. |
| [10] | 董妍妍, 张可敬, 储俊, 储全根. 抵当汤含药血清通过PI3K/Akt/mTOR信号通路增强高糖诱导的大鼠肾小球内皮细胞自噬[J]. 南方医科大学学报, 2025, 45(3): 461-469. |
| [11] | 廖茗, 钟文华, 张冉, 梁娟, 徐文陶睿, 万文珺, 吴超, 李曙. 源自蛇毒的蛋白C激活剂通过调控HIF-1α抑制BNIP3活性氧生成保护人脐静脉内皮细胞免受缺氧-复氧损伤[J]. 南方医科大学学报, 2025, 45(3): 614-621. |
| [12] | 花代平, 宣巧玉, 孙兰婷, 于庆生, 王琴, 王涛, 马麒颜, 杨文明, 汪瀚. Wilson 病患者 LncRNA Meg3 表达水平与肝纤维化指标的相关性分析[J]. 南方医科大学学报, 2025, 45(11): 2365-2374. |
| [13] | 易明, 罗烨, 吴露, 吴泽衡, 蒋翠平, 陈史钰, 柯晓. 黄芪甲苷通过抑制PINK1/Parkin通路调控细胞线粒体自噬减轻D-半乳糖诱导的内皮细胞衰老[J]. 南方医科大学学报, 2025, 45(11): 2427-2436. |
| [14] | 郭克磊, 李颖利, 宣晨光, 侯紫君, 叶松山, 李林运, 陈丽平, 韩立, 卞华. 益气养阴化浊通络方通过调控miR-21a-5p/FoxO1/PINK1介导的线粒体自噬减轻糖尿病肾病小鼠的足细胞损伤[J]. 南方医科大学学报, 2025, 45(1): 27-34. |
| [15] | 展俊平, 黄硕, 孟庆良, 范围, 谷慧敏, 崔家康, 王慧莲. 缺氧微环境下补阳还五汤通过抑制BNIP3-PI3K/Akt通路抑制类风湿关节炎滑膜成纤维细胞的线粒体自噬[J]. 南方医科大学学报, 2025, 45(1): 35-42. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||