南方医科大学学报 ›› 2026, Vol. 46 ›› Issue (3): 559-569.doi: 10.12122/j.issn.1673-4254.2026.03.10
• 基础研究 • 上一篇
陶红成1( ), 梁富凯1, 黄文波1, 范思奇2(
), 梁富凯1, 黄文波1, 范思奇2( ), 曾平2,3(
), 曾平2,3( )
)
收稿日期:2025-08-09
出版日期:2026-03-20
发布日期:2026-03-26
通讯作者:
范思奇,曾平
E-mail:taohongcheng2021@stu.gxtcmu.edu.cn;fansiqi932828@163.com;zengp@gxtcmu.edu.cn
作者简介:陶红成,在读博士研究生,E-mail: taohongcheng2021@stu.gxtcmu.edu.cn
基金资助:
Hongcheng TAO1( ), Fukai LIANG1, Wenbo HUANG1, Siqi FAN2(
), Fukai LIANG1, Wenbo HUANG1, Siqi FAN2( ), Ping ZENG2,3(
), Ping ZENG2,3( )
)
Received:2025-08-09
Online:2026-03-20
Published:2026-03-26
Contact:
Siqi FAN, Ping ZENG
E-mail:taohongcheng2021@stu.gxtcmu.edu.cn;fansiqi932828@163.com;zengp@gxtcmu.edu.cn
摘要:
目的 运用113种机器学习方法并基于酒精暴露相关基因构建并验证股骨头坏死诊断模型。 方法 从GEO数据库中获取酒精暴露和股骨头坏死相关的转录组学数据,进行系统分析以开发和验证股骨头坏死诊断模型。合并数据集并去除批次效应后,进一步进行差异基因分析,确定酒精暴露和股骨头坏死的差异表达基因(DEGs)并取交集。对取得的交集DEGs 进行功能富集分析。采用单样本基因集富集分析(ssGSEA)对免疫浸润情况进行量化分析。在训练集上探索12种机器学习算法的113种组合,并采用10折交叉验证,构建ONFH的诊断模型,在测试集上进行验证。将MC3T3-E1细胞分为对照组和酒精组,通过qRT-PCR检测两组细胞中构建模型的差异基因表达,以验证构建模型的可靠性。利用“Enrichr”平台来识别ONFH的潜在药物。 结果 总共鉴定出21个酒精暴露和股骨头坏死均密切相关的交集DEGs,基于富集分析,这些基因亦有参与免疫过程。通过免疫浸润分析显示,与健康对照组相比,酒精暴露和股骨头坏死患者体内免疫细胞浸润情况存在明显差异。接着,构建了由8个基因(SOAT1、GMCL1、GMPR、CISD2、ST3GAL6、AHSP、UBL3、PTPN12)组成的可靠的诊断模型,并进一步通过qRT-PCR验证发现,这8个基因在对照组和酒精组MC3T3-E1细胞中表达存在显著差异(P<0.05)。最后,预测了10个可能靶向治疗股骨头坏死的潜在药物。 结论 通过整合生物信息学分析和机器学习方法,并基于酒精暴露和股骨头坏死共有的DEGs,构建出可用于诊断股骨头坏死的可靠的模型。
陶红成, 梁富凯, 黄文波, 范思奇, 曾平. 酒精暴露与股骨头坏死的潜在关联:基于机器学习构建诊断模型[J]. 南方医科大学学报, 2026, 46(3): 559-569.
Hongcheng TAO, Fukai LIANG, Wenbo HUANG, Siqi FAN, Ping ZENG. Analysis of potential association of alcohol exposure with femoral head osteonecrosis and construction of a diagnostic model using machine learning[J]. Journal of Southern Medical University, 2026, 46(3): 559-569.
| Sequence number | GSE number | Source of sample | Type of tissue | Platform |
|---|---|---|---|---|
| 1 | GSE20489 | 29 alcohol and 25 control samples | Blood | GPL570 |
| 2 | GSE44456 | 20 alcohol and 19 control samples | Hippocampus | GPL6244 |
| 3 | GSE56906 | 4 alcohol and 6 control samples | Neural stem cells | GPL570 |
| 4 | GSE123568 | 30 ONFH and 10 control samples | Peripheral serum | GPL15207 |
| 5 | GSE74089 | 4 ONFH and 4 control samples | Cartilage | GPL13497 |
表1 本研究中收集的数据集基本信息
Tab.1 Basic information of the datasets collected in this study
| Sequence number | GSE number | Source of sample | Type of tissue | Platform |
|---|---|---|---|---|
| 1 | GSE20489 | 29 alcohol and 25 control samples | Blood | GPL570 |
| 2 | GSE44456 | 20 alcohol and 19 control samples | Hippocampus | GPL6244 |
| 3 | GSE56906 | 4 alcohol and 6 control samples | Neural stem cells | GPL570 |
| 4 | GSE123568 | 30 ONFH and 10 control samples | Peripheral serum | GPL15207 |
| 5 | GSE74089 | 4 ONFH and 4 control samples | Cartilage | GPL13497 |
| Gene | Forward primers | Reverse primer |
|---|---|---|
| GAPDH | GGTTGTCTCCTGCGACTTCA | TGGTCCAGGGTTTCTTACTCC |
| AHSP | CAGAAGACACACAAACCCCG | ATTGCTCTGAAAAGGGGCCA |
| CISD2 | AGGGCAGGAAGCATCATCAC | CTACCTTGCCCTTGACGTGT |
| GMCL1 | GCATGCTGCAATTGGATGGTT | GCCACTCGAGGCACTTTTTC |
| GMPR | AGACGCGGACCTTAAACTCG | TTCGCTCAAGATCCACCTCG |
| PTPN12 | GACTCTCCACCTGCTTTCAGT | TTCACTCCCTGCATCATGTCC |
| SOAT1 | TTCGGCCTTGTGCGACTTAT | AAGTCTAACCCGAGGCAAGC |
| ST3GAL6 | ACTGTGGGGAACAAATGGCT | GAAGATTGGCGAAAGCTGGC |
| UBL3 | TGTGTGATCCTGTAGCGTCG | ATTCAGTCCCACGGATGTGC |
表2 构建预测模型基因引物序列
Tab.2 Primer sequences of the genes used to construct the diagnostic model for alcohol-related femoral head osteoporosis
| Gene | Forward primers | Reverse primer |
|---|---|---|
| GAPDH | GGTTGTCTCCTGCGACTTCA | TGGTCCAGGGTTTCTTACTCC |
| AHSP | CAGAAGACACACAAACCCCG | ATTGCTCTGAAAAGGGGCCA |
| CISD2 | AGGGCAGGAAGCATCATCAC | CTACCTTGCCCTTGACGTGT |
| GMCL1 | GCATGCTGCAATTGGATGGTT | GCCACTCGAGGCACTTTTTC |
| GMPR | AGACGCGGACCTTAAACTCG | TTCGCTCAAGATCCACCTCG |
| PTPN12 | GACTCTCCACCTGCTTTCAGT | TTCACTCCCTGCATCATGTCC |
| SOAT1 | TTCGGCCTTGTGCGACTTAT | AAGTCTAACCCGAGGCAAGC |
| ST3GAL6 | ACTGTGGGGAACAAATGGCT | GAAGATTGGCGAAAGCTGGC |
| UBL3 | TGTGTGATCCTGTAGCGTCG | ATTCAGTCCCACGGATGTGC |
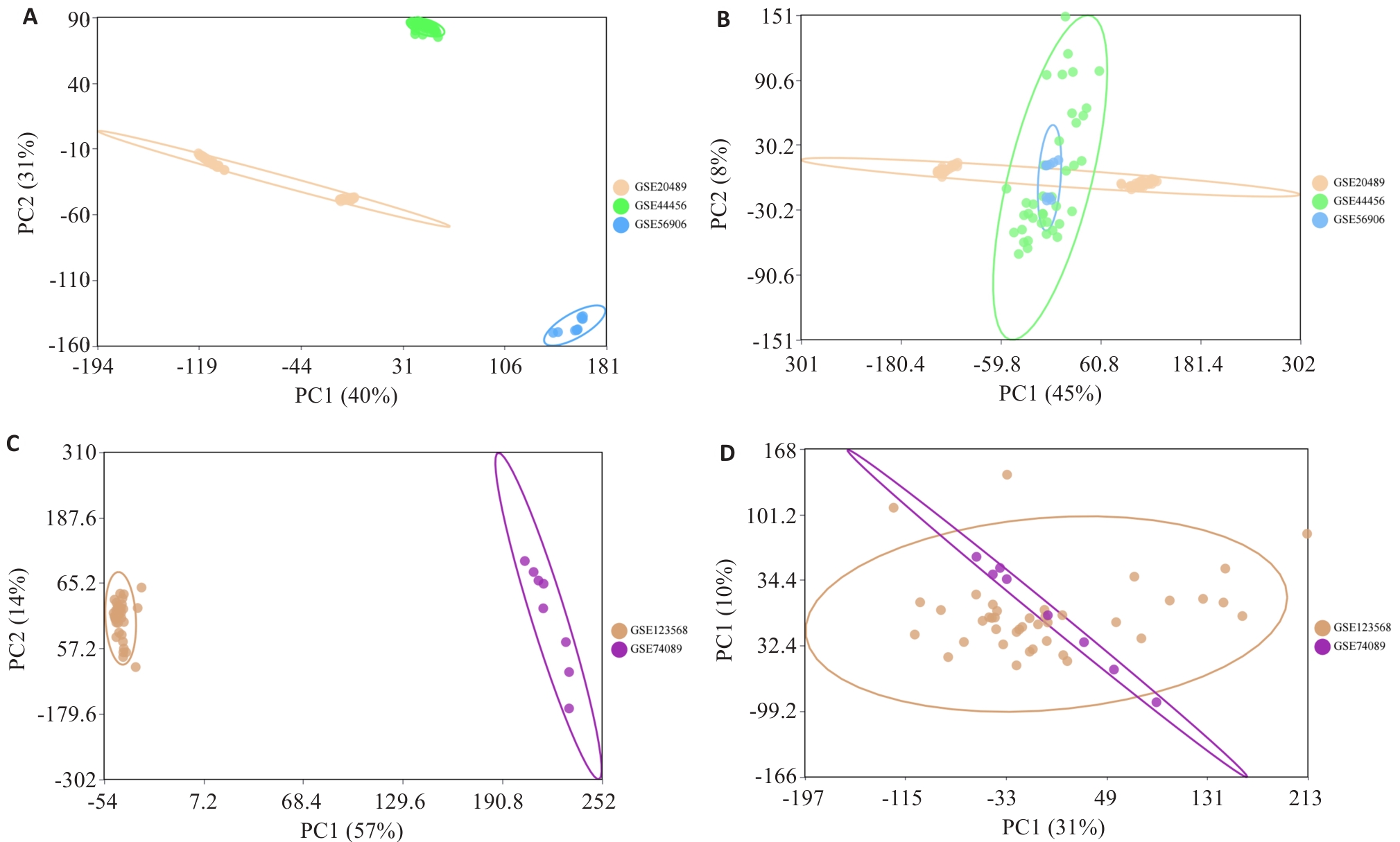
图1 酒精暴露和ONFH数据集整合前后的主成分分析
Fig.1 Principal component analysis before and after integration of alcohol exposure and ONFH datasets. A, B: Principal component analysis of the alcohol exposure datasets before and after correction for batch effects. C, D: Principal component analysis of the ONFH datasets before and after correction for batch effects.
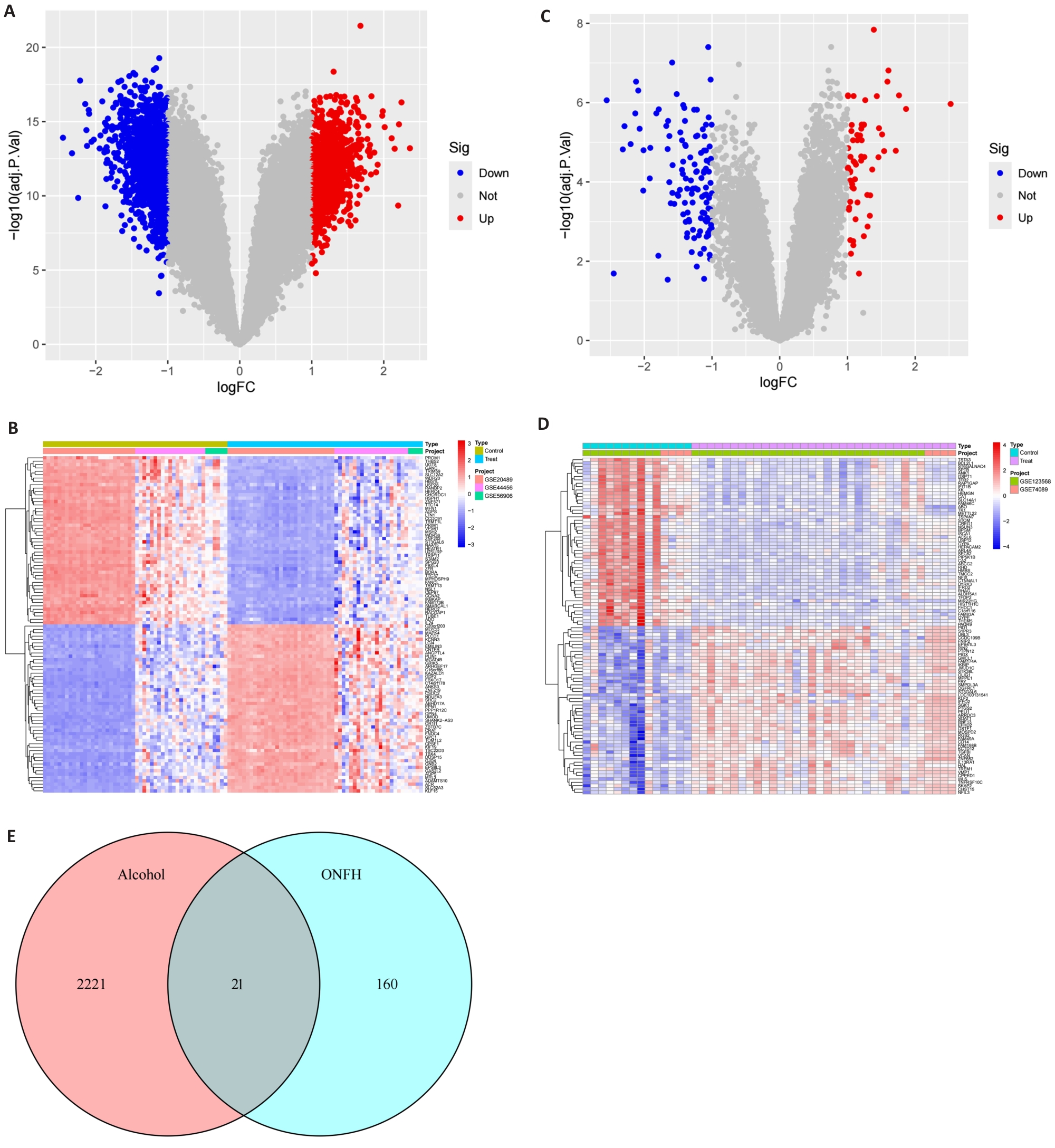
图2 酒精暴露和ONFH的DEGs的鉴定及交集基因
Fig.2 Identification of the differentially expressed genes (DEGs) of alcohol exposure and ONFH and the intersection genes between them. A, B: Volcano and heat maps of the DEGs for alcohol exposure. C, D: Volcano and heat maps of DEGs for ONFH. E: Venn diagram of the 21 DEGs.
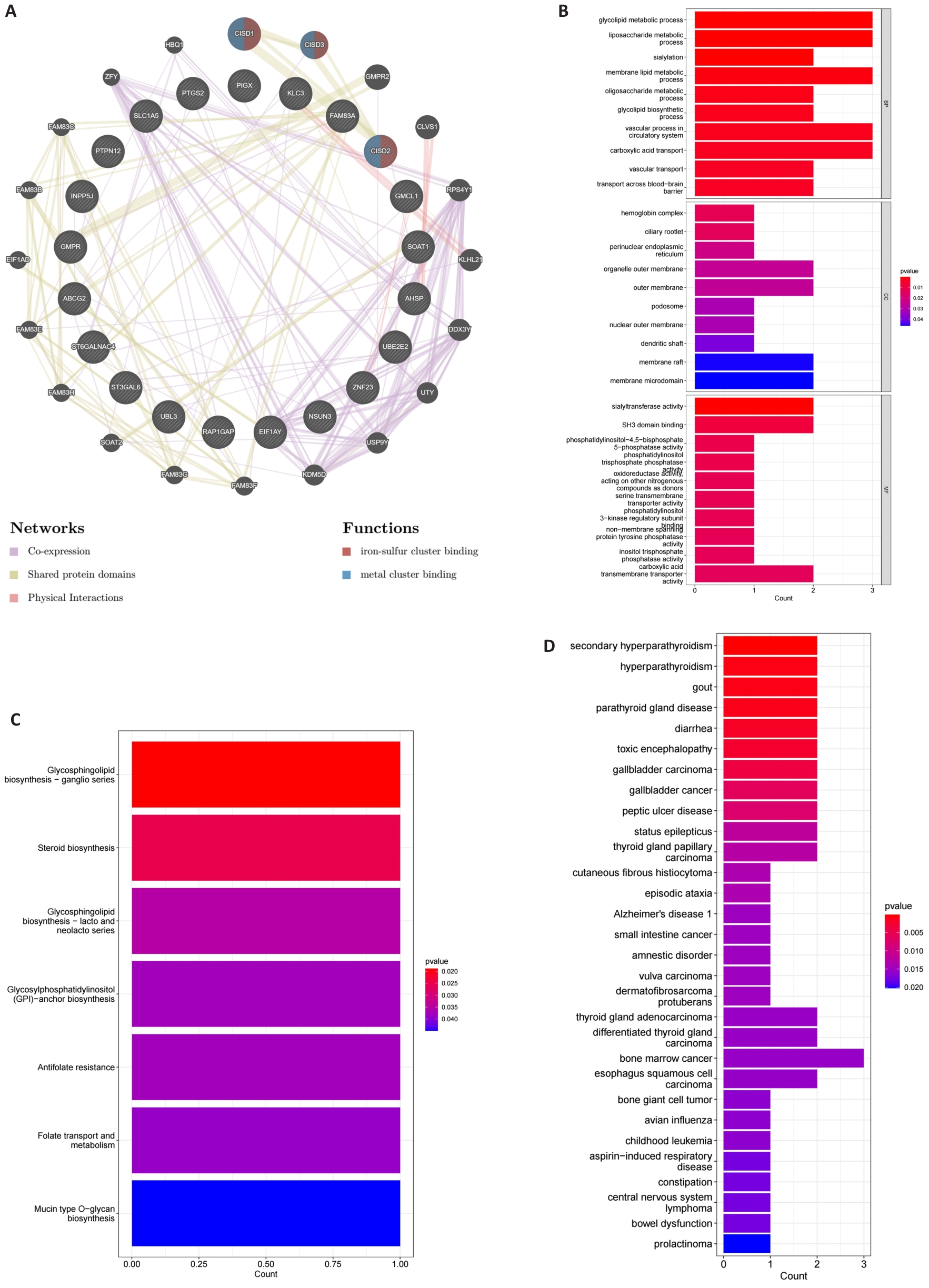
图3 蛋白质互作网络分析及富集分析
Fig.3 Protein interaction network analysis and enrichment analysis. A: Protein interaction network of 21 intersecting DEGs constructed by GeneMANIA. B: Bar graph of GO enrichment analysis results of biological processes, cellular components, and molecular functions. C: Bar graph of KEGG pathway enrichment analysis. D: Bar graph of DO enrichment analysis.
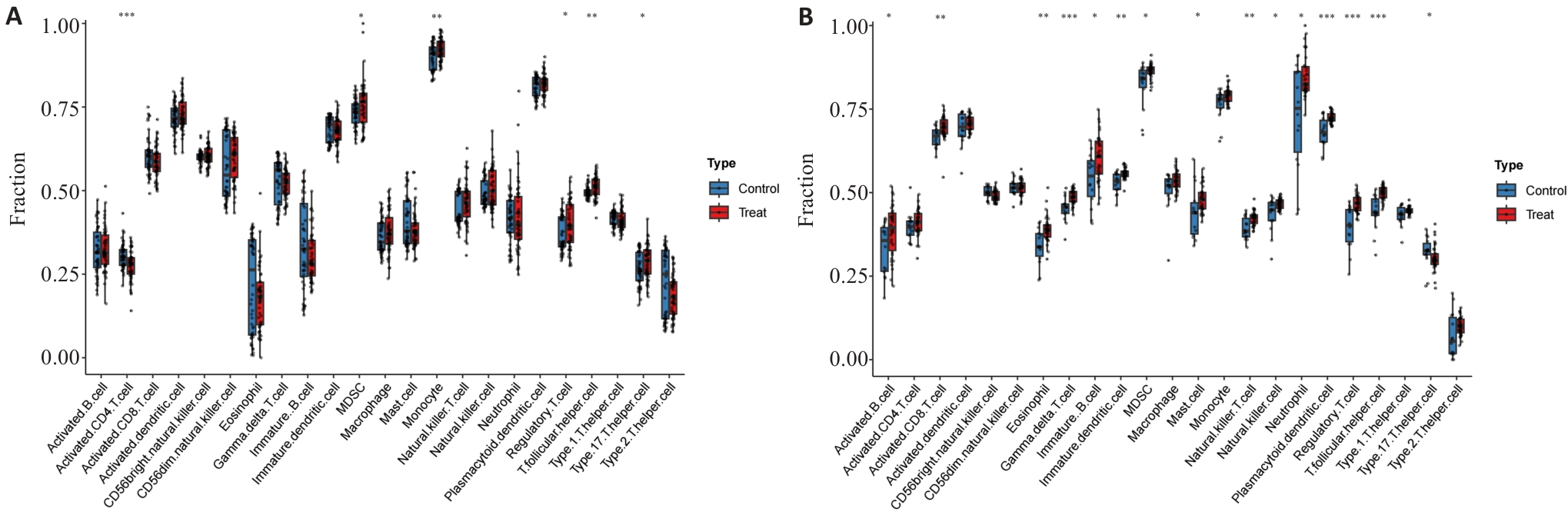
图4 酒精暴露和ONFH的免疫学特征
Fig.4 Immunological features of alcohol exposure and ONFH in their respective datasets. A: Boxplot comparison of immune cell abundance between alcohol exposed and control groups. B: Boxplot comparison of immune cell abundance between ONFH and control groups. *P<0.05, **P<0.01, ***P<0.001.
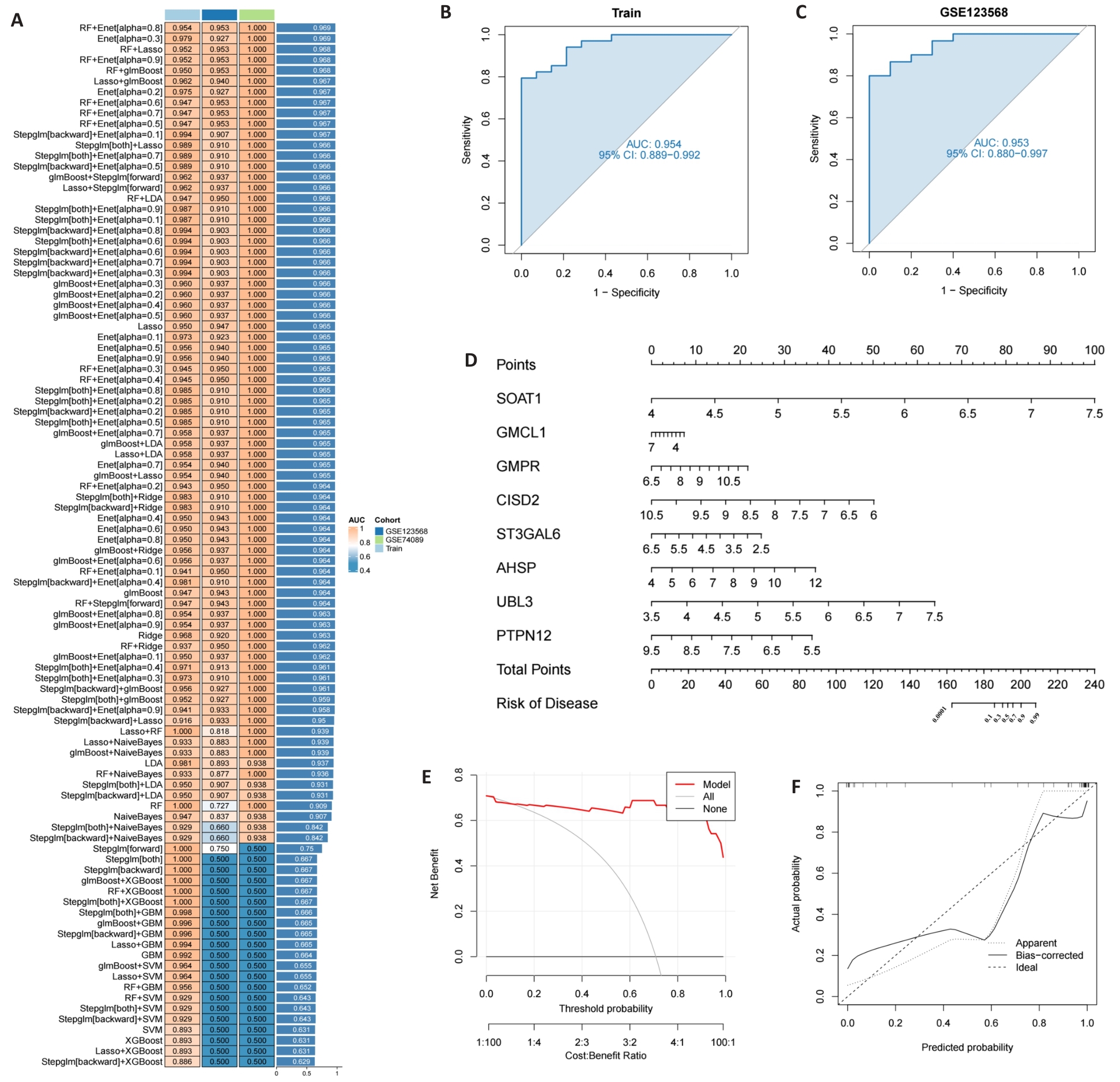
图5 构建ONFH模型及其性能
Fig.5 The diagnostic model of ONFH and assessment of its performance. A: 113 combinations of machine learning algorithms evaluated by 10-fold cross-validation. B, C: ROC curves for the training and test cohorts; D: Model nomogram based on eight genes. E, F: Decision curve and calibration curve of ONFH diagnostic model.
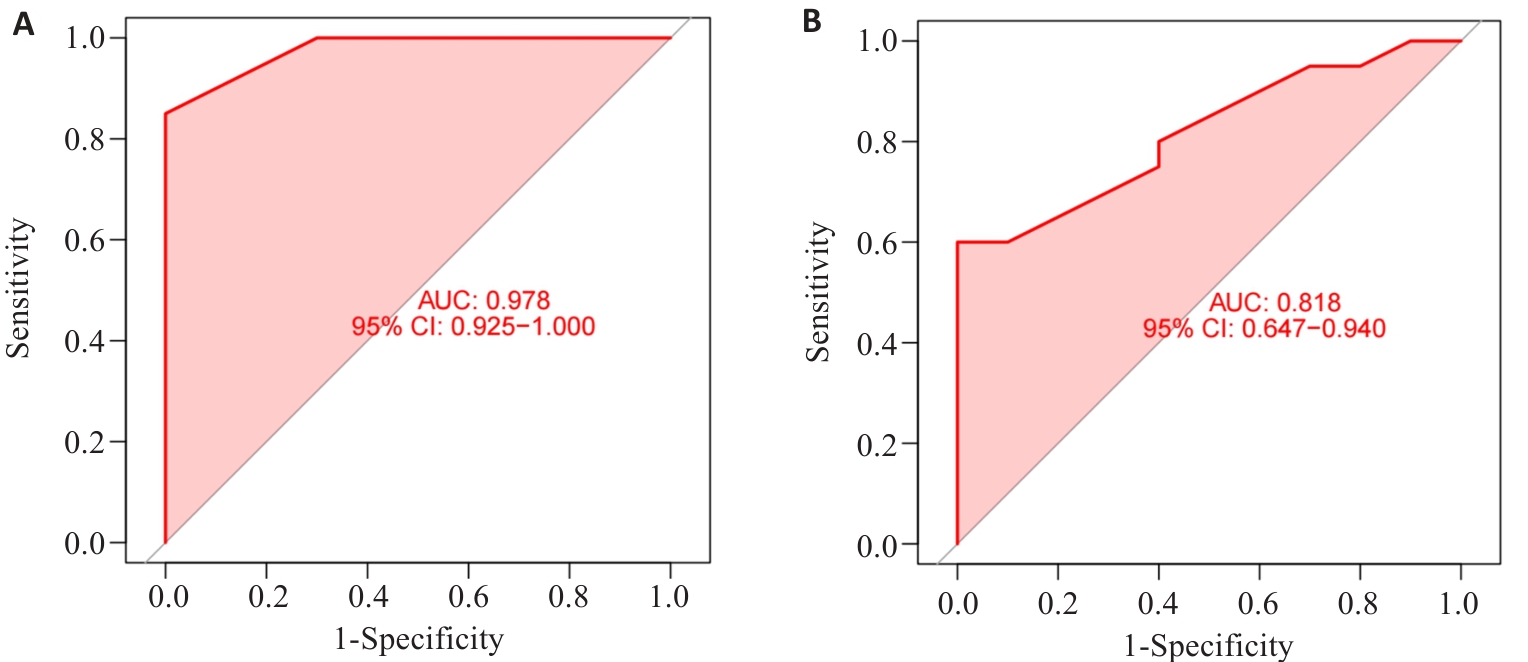
图6 临床亚组验证诊断模型的稳定性
Fig.6 Stability of the diagnostic model verified by clinical subgroups. A: ROC curve of female sample data in the diagnostic model. B: ROC curve of male sample data in the diagnostic model.
| Name of drug | P value | Combined Score | Genes |
|---|---|---|---|
| Valproic acid | 0.04785374 | 365.04628 | SOAT1; GMCL1 |
| Vorinostat | 0.04785374 | 124.87380 | SOAT1; GMPR; GMCL1 |
| Sulpiride | 0.04785374 | 123.67904 | SOAT1; PTPN12; GMCL1 |
| Rifabutin | 0.04785374 | 287.50862 | GMPR; GMCL1 |
| Epivincamine | 0.04785374 | 266.22331 | PTPN12; GMCL1 |
| Scriptaid | 0.04785374 | 105.14834 | SOAT1; GMPR; GMCL1 |
| Trichostatin A | 0.08122109 | 72.89887 | SOAT1; GMPR; GMCL1 |
| Anisomycin | 0.09425913 | 109.41726 | UBL3; GMCL1 |
| Isopentenyladenine | 0.09425913 | 828.65378 | PTPN12 |
| Hc-toxin | 0.09425913 | 97.98479 | SOAT1; GMPR |
表3 酒精暴露和ONFH交集基因的靶向药物
Tab.3 Drugs targeting the intersection genes of alcohol exposure and ONFH
| Name of drug | P value | Combined Score | Genes |
|---|---|---|---|
| Valproic acid | 0.04785374 | 365.04628 | SOAT1; GMCL1 |
| Vorinostat | 0.04785374 | 124.87380 | SOAT1; GMPR; GMCL1 |
| Sulpiride | 0.04785374 | 123.67904 | SOAT1; PTPN12; GMCL1 |
| Rifabutin | 0.04785374 | 287.50862 | GMPR; GMCL1 |
| Epivincamine | 0.04785374 | 266.22331 | PTPN12; GMCL1 |
| Scriptaid | 0.04785374 | 105.14834 | SOAT1; GMPR; GMCL1 |
| Trichostatin A | 0.08122109 | 72.89887 | SOAT1; GMPR; GMCL1 |
| Anisomycin | 0.09425913 | 109.41726 | UBL3; GMCL1 |
| Isopentenyladenine | 0.09425913 | 828.65378 | PTPN12 |
| Hc-toxin | 0.09425913 | 97.98479 | SOAT1; GMPR |
| Gene | Control group | Alcohol group | t | P |
|---|---|---|---|---|
| PTPN12 | 1.027±0.244 | 1.468±0.345 | -3.135 | 0.006 |
| UBL3 | 1.075±0.434 | 2.407±1.014 | -3.622 | 0.002 |
| SOAT1 | 1.008±0.134 | 2.649±0.592 | -8.101 | <0.001 |
| GMC1 | 1.011±0.160 | 2.899±0.688 | -8.015 | <0.001 |
| GMPR | 1.016±0.187 | 0.661±0.152 | 4.417 | <0.001 |
| CISD2 | 1.023±0.226 | 0.704±0.190 | 3.240 | 0.005 |
| ST3GAL6 | 1.051±0.342 | 3.341±0.761 | -8.232 | <0.001 |
| AHSP | 1.053±0.341 | 0.693±0.179 | 2.807 | 0.013 |
表4 构建预测模型8个基因在两组细胞中的相对表达
Tab.4 Relative expression of the 8 genes used to construct the diagnostic model in the two groups of cells (n=9)
| Gene | Control group | Alcohol group | t | P |
|---|---|---|---|---|
| PTPN12 | 1.027±0.244 | 1.468±0.345 | -3.135 | 0.006 |
| UBL3 | 1.075±0.434 | 2.407±1.014 | -3.622 | 0.002 |
| SOAT1 | 1.008±0.134 | 2.649±0.592 | -8.101 | <0.001 |
| GMC1 | 1.011±0.160 | 2.899±0.688 | -8.015 | <0.001 |
| GMPR | 1.016±0.187 | 0.661±0.152 | 4.417 | <0.001 |
| CISD2 | 1.023±0.226 | 0.704±0.190 | 3.240 | 0.005 |
| ST3GAL6 | 1.051±0.342 | 3.341±0.761 | -8.232 | <0.001 |
| AHSP | 1.053±0.341 | 0.693±0.179 | 2.807 | 0.013 |
| [1] | Li LY, Zhao SK, Leng ZK, et al. Pathological mechanisms and related markers of steroid-induced osteonecrosis of the femoral head[J]. Ann Med, 2024, 56(1): 2416070. doi:10.1080/07853890.2024.2416070 |
| [2] | Shao WK, Wang P, Lv X, et al. Unraveling the role of endothelial dysfunction in osteonecrosis of the femoral head: a pathway to new therapies[J]. Biomedicines, 2024, 12(3): 664. doi:10.3390/biomedicines12030664 |
| [3] | Huang K, Zeng Y, Zhang QY, et al. Less acetabular coverage predicts the failure of core decompression for osteonecrosis of the femoral head: a retrospective cohort study[J]. Orthop Surg, 2024, 16(7): 1614-21. doi:10.1111/os.14094 |
| [4] | Zhang SL, Wang HJ, Meng Q, et al. Recent advances in osteonecrosis of the femoral head: a focus on mesenchymal stem cells and adipocytes[J]. J Transl Med, 2025, 23(1): 592. doi:10.1186/s12967-025-06564-6 |
| [5] | Sun JH, Ma BW, Zhang CY, et al. To investigate the effect of neck-shaft angle in surgical hip dislocation combined with femoral neck rotational osteotomy in the treatment of osteonecrosis of the femoral head and to combine with finite element analysis[J]. Front Bioeng Biotechnol, 2025, 13: 1495292. doi:10.3389/fbioe.2025.1495292 |
| [6] | Wang YX, Luan S, Yuan Z, et al. The combined use of platelet-rich plasma clot releasate and allogeneic human umbilical cord mesenchymal stem cells rescue glucocorticoid-induced osteo-necrosis of the femoral head[J]. Stem Cells Int, 2022, 2022: 7432665. doi:10.1155/2022/7432665 |
| [7] | Zheng QY, Tao Y, Geng L, et al. Non-traumatic osteonecrosis of the femoral head induced by steroid and alcohol exposure is associated with intestinal flora alterations and metabolomic profiles[J]. J Orthop Surg Res, 2024, 19(1): 236. doi:10.1186/s13018-024-04713-z |
| [8] | Yang WB, Pan Q, Peng YZ, et al. Dual-target nanotherapy for vascular endothelium and bone mesenchymal stem cells halt steroid-induced osteonecrosis of the femoral head progression[J]. J Control Release, 2025, 380: 219-39. doi:10.1016/j.jconrel.2024.12.081 |
| [9] | Zhao DW, Yu M, Hu K, et al. Prevalence of nontraumatic osteonecrosis of the femoral head and its associated risk factors in the Chinese population: results from a nationally representative survey[J]. Chin Med J (Engl), 2015, 128(21): 2843-50. doi:10.4103/0366-6999.168017 |
| [10] | 黄 捷, 施扬华, 谭 桢, 等. 吻合血管游离腓骨移植治疗股骨头坏死[J]. 中国组织工程研究, 2024, 28(21): 3373-9. |
| [11] | Doyle A, Foley R, Houghton F. A spatial examination of alcohol availability and the level of disadvantage of schools in Ireland[J]. BMC Public Health, 2024, 24(1): 795. doi:10.1186/s12889-024-18261-y |
| [12] | Moore SC, Wood AM, Moore L, et al. A rank based social norms model of how people judge their levels of drunkenness whilst intoxicated[J]. BMC Public Health, 2016, 16: 798. doi:10.1186/s12889-016-3469-z |
| [13] | Fu ZW, Zhou JF, Pan HY, et al. QingGan LiDan capsules improved alcoholic liver injury by regulating liver lipid transport and oxidative stress in mice[J]. Front Pharmacol, 2025, 16: 1575280. doi:10.3389/fphar.2025.1575280 |
| [14] | Yarimay R, Beyer DKE, Mundorf A, et al. Expression of GDAP1 gene correlates with alcohol deprivation effect[J]. Genes Brain Behav, 2025, 24(2): e70022. doi:10.1111/gbb.70022 |
| [15] | Wang TC, Jia ZK, An CH, et al. The protective effect of Auricularia cornea var. Li. polysaccharide on alcoholic liver disease and its effect on intestinal microbiota[J]. Molecules, 2023, 28(24): 8003. doi:10.3390/molecules28248003 |
| [16] | Song YX, Li HX, Ma SH, et al. Losartan protects human stem cell-derived cardiomyocytes from angiotensin II-induced alcoholic cardiotoxicity[J]. Cell Death Discov, 2022, 8(1): 134. doi:10.1038/s41420-022-00945-2 |
| [17] | Chen HZ, Zhang YF, Guo TS, et al. Genetic variant rs72613567 of HSD17B13 gene reduces alcohol-related liver disease risk in Chinese Han population[J]. Liver Int, 2020, 40(9): 2194-202. doi:10.1111/liv.14616 |
| [18] | Ha Y, Jeong I, Kim TH. Alcohol-related liver disease: an overview on pathophysiology, diagnosis and therapeutic perspectives[J]. Biomedicines, 2022, 10(10): 2530. doi:10.3390/biomedicines10102530 |
| [19] | Gondim Teixeira PA, Dubois L, Hossu G, et al. Quantitative dynamic contrast-enhanced MRI of bone marrow perfusion at the proximal femur: influence of femoral head osteonecrosis risk factor and overt osteonecrosis[J]. Eur Radiol, 2023, 33(4): 2340-9. doi:10.1007/s00330-022-09250-z |
| [20] | Yue C, Ma MX, Guo JY, et al. Altered gut microbe metabolites in patients with alcohol-induced osteonecrosis of the femoral head: an integrated omics analysis[J]. Exp Ther Med, 2024, 28(2): 311. doi:10.3892/etm.2024.12599 |
| [21] | 郭锦丽, 曲成毅, 白 帆, 等. 酗酒与骨质疏松和股骨头坏死的相关性研究[J]. 中华流行病学杂志, 2013, 34(7): 732-5. doi:10.3760/cma.j.issn.0254-6450.2013.07.017 |
| [22] | Yoon BH, Kim TY, Shin IS, et al. Alcohol intake and the risk of osteonecrosis of the femoral head in Japanese populations: a dose-response meta-analysis of case-control studies[J]. Clin Rheumatol, 2017, 36(11): 2517-24. doi:10.1007/s10067-017-3740-4 |
| [23] | Tan B, Li WL, Zeng P, et al. Epidemiological study based on China osteonecrosis of the femoral head database[J]. Orthop Surg, 2021, 13(1): 153-60. doi:10.1111/os.12857 |
| [24] | Li JW, Zhang Y, You YW, et al. Unraveling the mechanisms of NK cell dysfunction in aging and Alzheimer's disease: insights from GWAS and single-cell transcriptomics[J]. Front Immunol, 2024, 15: 1360687. doi:10.3389/fimmu.2024.1360687 |
| [25] | Liu DY, Chen JH, Zhou JN, et al. Macrophage-related inflammatory responses to degradation products of biodegradable molybdenum implants[J]. Mater Today Bio, 2025, 31: 101519. doi:10.1016/j.mtbio.2025.101519 |
| [26] | Gu XF, Yu ZX, Qian TW, et al. Transcriptomic analysis identifies the shared diagnostic biomarkers and immune relationship between Atherosclerosis and abdominal aortic aneurysm based on fatty acid metabolism gene set[J]. Front Mol Biosci, 2024, 11: 1365447. doi:10.3389/fmolb.2024.1365447 |
| [27] | 陶振宇, 张月雷, 陈 华, 等. 细胞自噬在酒精性股骨头坏死中的作用及相关机制研究[J]. 中医正骨, 2018, 30(10): 4-11. doi:10.3969/j.issn.1001-6015.2018.10.002 |
| [28] | Jeong HJ, Park JW, Lee YK, et al. Comparison between osteonecrosis of the humeral and femoral heads-epidemiological analysis of the surgical trend using the nationwide claims database of the republic of Korea[J]. BMC Musculoskelet Disord, 2023, 24(1): 878. doi:10.1186/s12891-023-07022-4 |
| [29] | Zhao G, Liu YJ, Zheng YJ, et al. Hip arthroscopy debridement combined with multiple small-diameter fan-shaped low-speed drilling decompression in the treatment of early and middle stage osteonecrosis of the femoral head: 14 years follow-up[J]. Orthop Surg, 2024, 16(3): 604-12. doi:10.1111/os.13985 |
| [30] | Cevolani L, Focaccia M, Spazzoli B, et al. Is core decompression and bone marrow concentrate with demineralized bone matrix and platelet-rich fibrin suitable for treating femoral head osteonecrosis?[J]. J Hip Preserv Surg, 2024, 11(4): 263-70. doi:10.1093/jhps/hnae031 |
| [31] | Go MJ, Kim JM, Lee HL, et al. Hepatoprotective effect of Allium ochotense extracts on chronic alcohol-induced fatty liver and hepatic inflammation in C57BL/6 mice[J]. Int J Mol Sci, 2024, 25(6): 3496. doi:10.3390/ijms25063496 |
| [32] | Meenashi Sundaram D, Madesh VP, Rambrahma Reddy D, et al. Multiple dyselectrolytemia in a chronic alcohol abuser: a case report[J]. Cureus, 2023, 15(3): e36389. |
| [33] | Mrabet S, Attia S, Chaieb R. Pain assessment, management and impact among Hemodialysis patients: a study from Tunisia[J]. BMC Nurs, 2025, 24(1): 581. doi:10.1186/s12912-025-03130-9 |
| [34] | Sadek KM, El Moshy S, Radwan IA, et al. Molecular basis beyond interrelated bone resorption/regeneration in periodontal diseases: a concise review[J]. Int J Mol Sci, 2023, 24(5): 4599. doi:10.3390/ijms24054599 |
| [35] | Sturm R, Haag F, Bergmann CB, et al. Alcohol drinking leads to sex-dependent differentiation of T cells[J]. Eur J Trauma Emerg Surg, 2025, 51(1): 87. doi:10.1007/s00068-024-02732-3 |
| [36] | McTernan PM, Levitt DE, Welsh DA, et al. Alcohol impairs immunometabolism and promotes Naïve T cell differentiation to pro-inflammatory Th1 CD4+ T cells[J]. Front Immunol, 2022, 13: 839390. doi:10.3389/fimmu.2022.839390 |
| [37] | Brahadeeswaran S, Tamizhselvi R. Consequence of alcohol intoxication-mediated efferocytosis impairment[J]. Front Immunol, 2024, 15: 1386658. doi:10.3389/fimmu.2024.1386658 |
| [38] | Yu J, Fu L, Wu R, et al. Immunocytes in the tumor microenvironment: recent updates and interconnections[J]. Front Immunol, 2025, 16: 1517959. doi:10.3389/fimmu.2025.1517959 |
| [39] | Ma S, Lu Y, Sui S, et al. Unraveling the triad of immunotherapy, tumor microenvironment, and skeletal muscle biomechanics in oncology[J]. Front Immunol, 2025, 16: 1572821. doi:10.3389/fimmu.2025.1572821 |
| [40] | Lu KH, Hsieh YH, Lin RC, et al. Melatonin: a potential therapy for osteoporosis with insights into molecular mechanisms[J]. J Pineal Res, 2025, 77(4): e70062. doi:10.1111/jpi.70062 |
| [41] | Li ZH, Li XH, Jiang H, et al. Alcohol promotes CPT1A-induced lipid metabolism disorder to sentinel-regulate acute pancreatitis[J]. Eur J Med Res, 2025, 30(1): 35. doi:10.1186/s40001-024-02213-8 |
| [42] | Fu RR, Xue WQ, Liang JJ, et al. SOAT1 regulates cholesterol metabolism to induce EMT in hepatocellular carcinoma[J]. Cell Death Dis, 2024, 15(5): 325. doi:10.1038/s41419-024-06711-9 |
| [43] | Wu HH, Zhu Q, Liang N, et al. CISD2 regulates oxidative stress and mitophagy to maintain the balance of the follicular micro-environment in PCOS[J]. Redox Rep, 2024, 29(1): 2377870. doi:10.1080/13510002.2024.2377870 |
| [44] | Chen MY, Li WJ, Lei L, et al. Role of SOST in response to mechanical stimulation in bone and extraosseous organs[J]. Biomolecules, 2025, 15(6): 856. doi:10.3390/biom15060856 |
| [45] | Yen FS, Wang SI, Lin SY, et al. The impact of heavy alcohol consumption on cognitive impairment in young old and middle old persons[J]. J Transl Med, 2022, 20(1): 155. doi:10.1186/s12967-022-03353-3 |
| [46] | Meng C, Qi BC, Luo H, et al. Exploring the genetic association between immune cells and susceptibility to osteonecrosis using large-scale population data[J]. Heliyon, 2024, 10(14): e34547. doi:10.1016/j.heliyon.2024.e34547 |
| [47] | Sitaram P, Uyemura B, Malarkannan S, et al. Beyond the cell surface: targeting intracellular negative regulators to enhance T cell anti-tumor activity[J]. Int J Mol Sci, 2019, 20(23): 5821. doi:10.3390/ijms20235821 |
| [48] | Marino P, Mininni M, Deiana G, et al. Healthy lifestyle and cancer risk: modifiable risk factors to prevent cancer[J]. Nutrients, 2024, 16(6): 800. doi:10.3390/nu16060800 |
| [49] | Qi QY, Chen BR, Wu JR, et al. TsR-0072 inhibits colorectal cancer progression through modulating lipid and vitamin D3 metabolic reprogramming and inactivating the Wnt/β‑catenin signalling pathway[J]. Ann Med, 2025, 57(1): 2531253. doi:10.1080/07853890.2025.2531253 |
| [50] | 田方圆, 王台安, 李艳敏, 等. 鸡MOGAT和DGAT基因的克隆及表达特性研究[J]. 畜牧兽医学报, 2017, 48(2): 214-24. |
| [51] | Mahmoud AM. An overview of epigenetics in obesity: the role of lifestyle and therapeutic interventions[J]. Int J Mol Sci, 2022, 23(3): 1341. doi:10.3390/ijms23031341 |
| [52] | Grønningsæter IS, Fredly HK, Gjertsen BT, et al. Systemic metabolomic profiling of acute myeloid leukemia patients before and during disease-stabilizing treatment based on all-trans retinoic acid, valproic acid, and low-dose chemotherapy[J]. Cells, 2019, 8(10): 1229. doi:10.3390/cells8101229 |
| [53] | Yue SR, Fan JY, Xie DL, et al. Unveiling the therapeutic potential: targeting fibroblast-like synoviocytes in rheumatoid arthritis[J]. Expert Rev Mol Med, 2025, 27: e18. doi:10.1017/erm.2025.11 |
| [54] | Chandrasekar N, Sharma K, Jain S, et al. A critical assessment on stability behaviour of Vorinostat using LC-MS-QTOF with H/D exchange and NMR[J]. J Pharm Biomed Anal, 2023, 236: 115687. doi:10.1016/j.jpba.2023.115687 |
| [55] | Gulfo MC, Lebowitz JJ, Ramos C, et al. Dopamine D2 receptors in hilar mossy cells regulate excitatory transmission and hippocampal function[J]. Proc Natl Acad Sci USA, 2023, 120(50): e2307509120. doi:10.1073/pnas.2307509120 |
| [1] | 陈君尧, 陈泽宇, 林钊杰, 方梦浩, 沈超英, 许琦, 张晓怡, 卢鲁. 饮茶对胃肠道疾病风险的双重作用:基于可解释机器学习与大语言模型的联合预测辅助模型[J]. 南方医科大学学报, 2026, 46(2): 353-361. |
| [2] | 崔运能, 冯敏清, 姚亮凤, 严杰文, 李闻瀚, 黄燕平. 基于欠采样的影像组学机器学习模型术前预测子宫肌瘤高强度聚焦超声消融效果[J]. 南方医科大学学报, 2026, 46(1): 141-149. |
| [3] | 黄启智, 谢戴鹏, 姚霖彤, 李洽轩, 吴少伟, 周海榆. 肿瘤微环境特异性CT影像组学标签预测非小细胞肺癌免疫治疗疗效[J]. 南方医科大学学报, 2025, 45(9): 1903-1918. |
| [4] | 姜君, 封硕, 孙银贵, 安燕. 经尿道前列腺钬激光剜除术后低体温风险预测模型:基于逻辑回归、决策树和支持向量机[J]. 南方医科大学学报, 2025, 45(9): 2019-2025. |
| [5] | 陈梅妹, 王洋, 雷黄伟, 张斐, 黄睿娜, 杨朝阳. 基于多种机器学习算法和语音情绪特征的阈下抑郁辨识模型构建[J]. 南方医科大学学报, 2025, 45(4): 711-717. |
| [6] | 张申尧, 卢敏, 邝高艳, 许晓彤, 付俊, 曾楚然. 过表达HDAC1通过诱导SP1去乙酰化改善类固醇诱导的小鼠骨细胞凋亡[J]. 南方医科大学学报, 2025, 45(1): 10-17. |
| [7] | 许怀文, 翁丽, 薛鸿. CXCL12可作为2型糖尿病合并慢性阻塞性肺疾病的潜在治疗靶点[J]. 南方医科大学学报, 2025, 45(1): 100-109. |
| [8] | 陈莉莉, 吴天宇, 张铭, 丁子夏, 张妍, 杨依清, 郑佳倩, 张小楠. 类风湿关节炎的潜在生物标志物及其免疫调控机制:基于GEO数据库[J]. 南方医科大学学报, 2024, 44(6): 1098-1108. |
| [9] | 申采玉, 王帅, 周锐盈, 汪雨贺, 高琴, 陈兴智, 杨枢. 慢性心力衰竭合并肺部感染患者院内死亡风险预测:基于可解释性机器学习方法[J]. 南方医科大学学报, 2024, 44(6): 1141-1148. |
| [10] | 左志威, 孟庆良, 崔家康, 郭克磊, 卞华. 基于硬皮病线粒体相关基因的人工神经网络模型的构建[J]. 南方医科大学学报, 2024, 44(5): 920-929. |
| [11] | 朱 奇, 路云翔, 彭 优, 何嘉乐, 韦泽宇, 李智勇, 陈郁鲜. α2-巨球蛋白通过调控血管内皮细胞改善小鼠激素性股骨头坏死[J]. 南方医科大学学报, 2024, 44(4): 712-719. |
| [12] | 梁一豪, 赖颖君, 袁燕文, 袁 炜, 张锡波, 张拔山, 卢志锋. 基于GEO数据库筛选胃癌差异表达基因及其功能和通路富集分析[J]. 南方医科大学学报, 2024, 44(3): 605-616. |
| [13] | 任自敬, 周佩洋, 田静. 基于ceRNA芯片研究帕金森病患者血浆长链非编码RNA表达谱并验证lnc-CTSD-5∶1在PD细胞模型中的作用[J]. 南方医科大学学报, 2024, 44(11): 2146-2155. |
| [14] | 黄晓茵, 陈凤莲, 张煜, 梁淑君. 多参数多区域MRI影像组学特征与临床信息联合模型可有效预测脑胶质瘤患者生存期[J]. 南方医科大学学报, 2024, 44(10): 2004-2014. |
| [15] | 何慧珊, 郭二嘉, 蒙文仪, 王 彧, 王 雯, 何文乐, 吴元魁, 阳 维. 基于磁共振图像机器学习放射组学模型预测脑胶质瘤的强化[J]. 南方医科大学学报, 2024, 44(1): 194-200. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||