Journal of Southern Medical University ›› 2026, Vol. 46 ›› Issue (2): 456-465.doi: 10.12122/j.issn.1673-4254.2026.02.23
Haokun ZHU1( ), Yanbu GUO1,2(
), Yanbu GUO1,2( ), Xiangjun XIN1(
), Xiangjun XIN1( ), Chaoyang LI1, Dongming ZHOU3,4
), Chaoyang LI1, Dongming ZHOU3,4
Received:2025-07-09
Online:2026-02-20
Published:2026-03-10
Contact:
Yanbu GUO, Xiangjun XIN
E-mail:332316040996@zzuli.edu.cn;guoyanbu@ zzuli.edu.cn;2004006@zzuli.edu.cn
Supported by:Haokun ZHU, Yanbu GUO, Xiangjun XIN, Chaoyang LI, Dongming ZHOU. Drug repositioning prediction based on dynamic feature learning on heterogeneous graphs[J]. Journal of Southern Medical University, 2026, 46(2): 456-465.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.j-smu.com/EN/10.12122/j.issn.1673-4254.2026.02.23
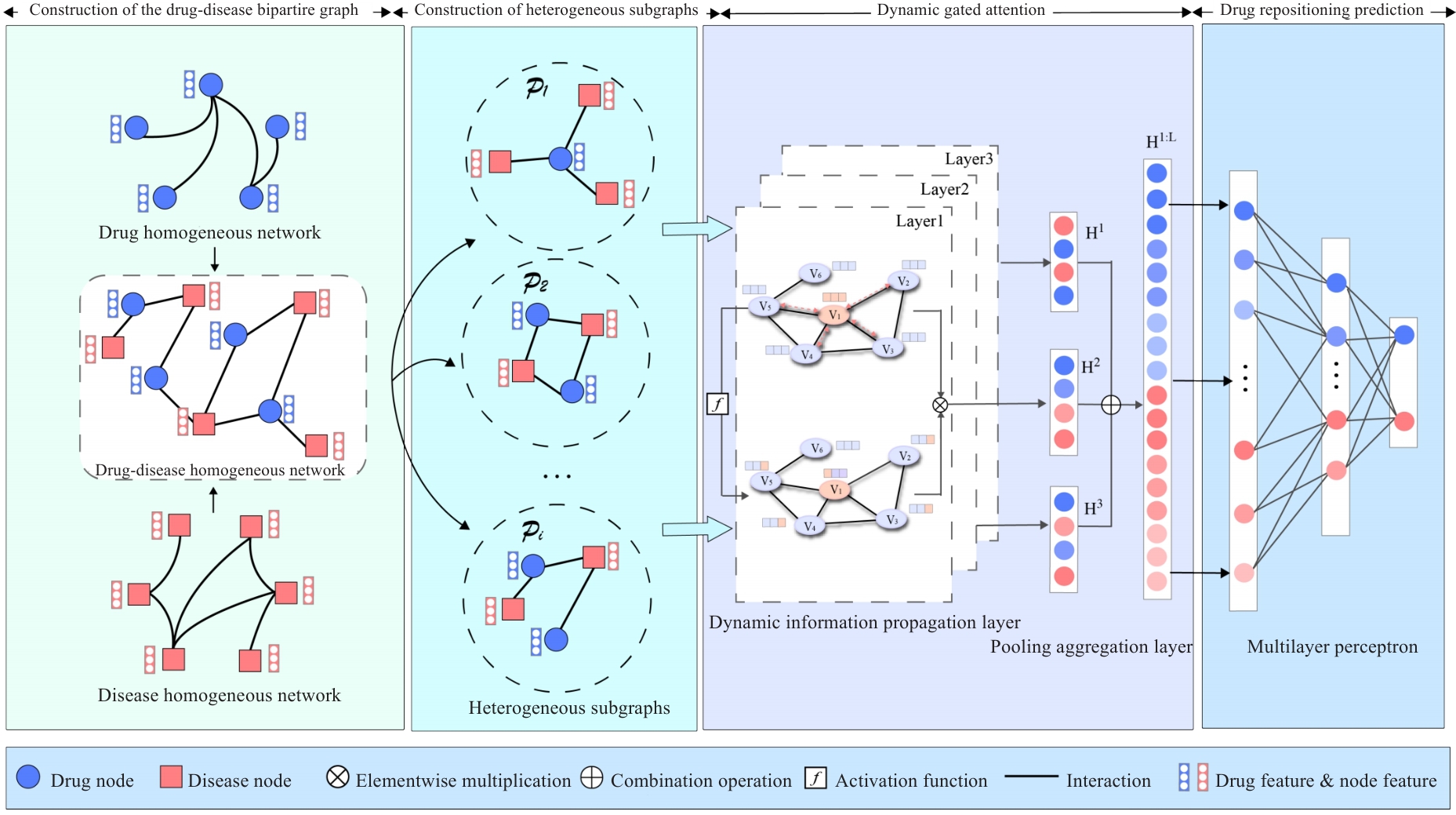
Fig.1 Overall architecture of the DRLHG, consisting of construction of a drug-disease bipartite graph, heterogeneous subgraphs, dynamic gated attention, and drug repositioning prediction.
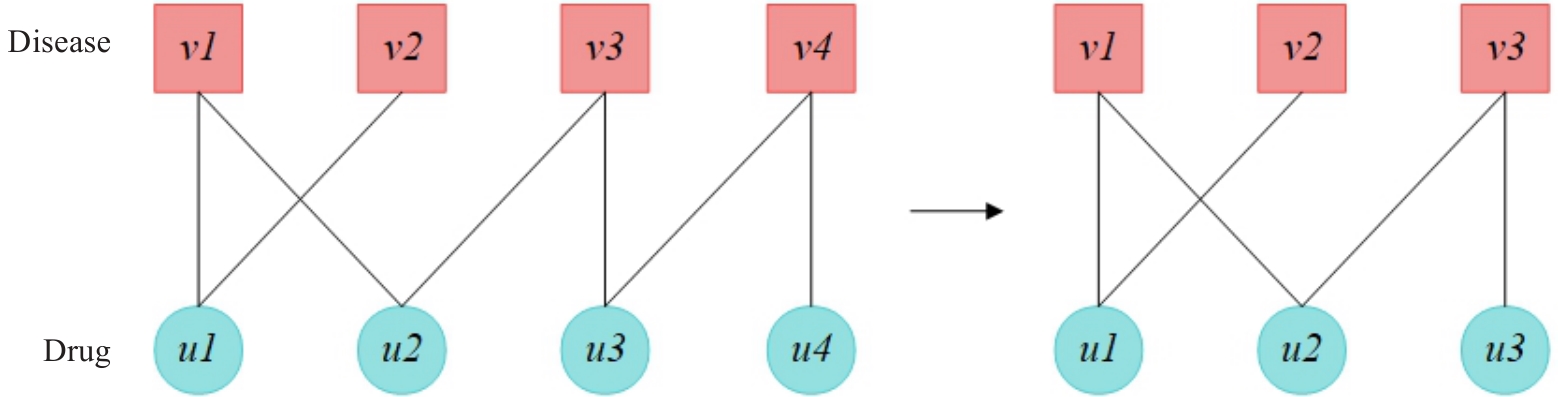
Fig. 2 Heterogeneous subgraph extraction process. For the drug central node u₂ and the disease central node v₂, their second-order neighbor nodes are extracted and combined into a heterogeneous subgraph.
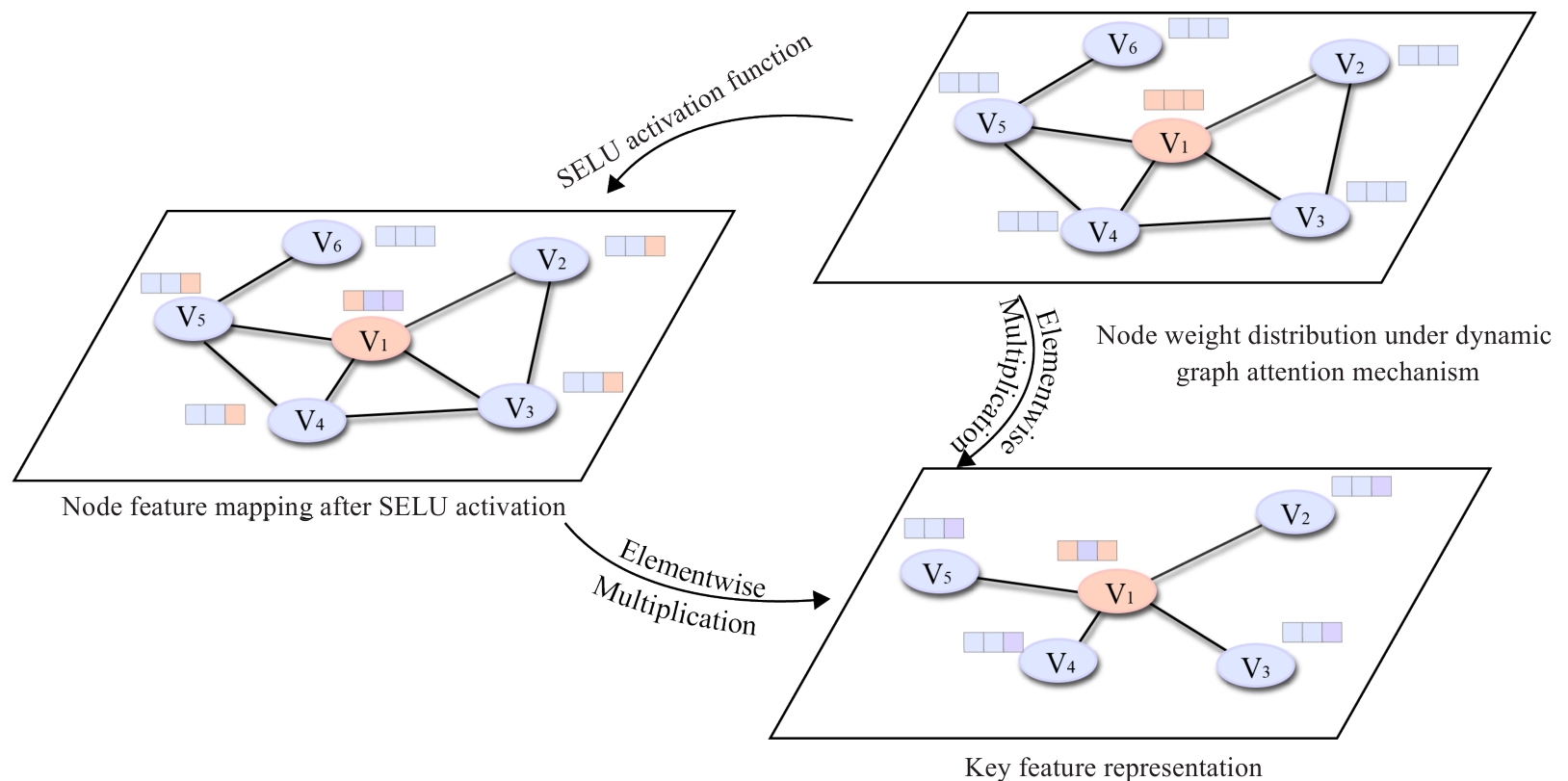
Fig.3 Schematic diagram of the gating mechanism. The key feature representations are obtained by introducing SELU to model complex nonlinear relationships and applying a Sigmoid gate to selectively filter noise.
| Dataset | Drugs | Diseases | Drug-disease associations |
|---|---|---|---|
| Gdataset | 593 | 313 | 1933 |
| Cdataset | 663 | 409 | 2352 |
| LRSSL | 763 | 681 | 3051 |
Tab.1 Drug-disease association dataset information
| Dataset | Drugs | Diseases | Drug-disease associations |
|---|---|---|---|
| Gdataset | 593 | 313 | 1933 |
| Cdataset | 663 | 409 | 2352 |
| LRSSL | 763 | 681 | 3051 |
| Model | Gdataset | Cdataset | LRSSL | |||
|---|---|---|---|---|---|---|
| AUROC | AUPR | AUROC | AUPR | AUROC | AUPR | |
| NMFDR | 0.9452±0.0089 | 0.9478±0.0104 | 0.9511±0.0053 | 0.9567±0.0124 | 0.9336±0.0049 | 0.9442±0.0115 |
| DPNetinfer | 0.9120±0.0055 | 0.9218±0.0071 | 0.9197±0.0036 | 0.9307±0.0055 | 0.9230±0.0044 | 0.9342±0.0093 |
| FuHLDR | 0.9063±0.0067 | 0.8970±0.0126 | 0.9423±0.0104 | 0.9235±0.0103 | 0.9311±0.0113 | 0.9196±0.0093 |
| DRWBNCF | 0.9121±0.0132 | 0.9314±0.0239 | 0.9276±0.0141 | 0.9395±0.0149 | 0.9253±0.0218 | 0.9389±0.0150 |
| DRAGNN | 0.9442±0.0086 | 0.9504±0.0093 | 0.9479±0.0077 | 0.9522±0.0102 | 0.9504±0.0103 | 0.9595±0.0083 |
| PSGCN | 0.9484±0.0101 | 0.9551±0.0094 | 0.9561±0.0120 | 0.9606±0.0077 | 0.9405±0.0137 | 0.9412±0.0103 |
| HDGAT | 0.9478±0.0102 | 0.9575±0.0117 | 0.9524±0.0097 | 0.9613±0.0103 | 0.9493±0.0133 | 0.9581±0.0106 |
| DRLHG | 0.9551±0.0079 | 0.9606±0.0058 | 0.9610±0.0039 | 0.9675±0.0027 | 0.9557±0.0075 | 0.9615±0.0054 |
Tab.2 Performance comparison of DRLHG and the baseline models in 10-fold cross-validation
| Model | Gdataset | Cdataset | LRSSL | |||
|---|---|---|---|---|---|---|
| AUROC | AUPR | AUROC | AUPR | AUROC | AUPR | |
| NMFDR | 0.9452±0.0089 | 0.9478±0.0104 | 0.9511±0.0053 | 0.9567±0.0124 | 0.9336±0.0049 | 0.9442±0.0115 |
| DPNetinfer | 0.9120±0.0055 | 0.9218±0.0071 | 0.9197±0.0036 | 0.9307±0.0055 | 0.9230±0.0044 | 0.9342±0.0093 |
| FuHLDR | 0.9063±0.0067 | 0.8970±0.0126 | 0.9423±0.0104 | 0.9235±0.0103 | 0.9311±0.0113 | 0.9196±0.0093 |
| DRWBNCF | 0.9121±0.0132 | 0.9314±0.0239 | 0.9276±0.0141 | 0.9395±0.0149 | 0.9253±0.0218 | 0.9389±0.0150 |
| DRAGNN | 0.9442±0.0086 | 0.9504±0.0093 | 0.9479±0.0077 | 0.9522±0.0102 | 0.9504±0.0103 | 0.9595±0.0083 |
| PSGCN | 0.9484±0.0101 | 0.9551±0.0094 | 0.9561±0.0120 | 0.9606±0.0077 | 0.9405±0.0137 | 0.9412±0.0103 |
| HDGAT | 0.9478±0.0102 | 0.9575±0.0117 | 0.9524±0.0097 | 0.9613±0.0103 | 0.9493±0.0133 | 0.9581±0.0106 |
| DRLHG | 0.9551±0.0079 | 0.9606±0.0058 | 0.9610±0.0039 | 0.9675±0.0027 | 0.9557±0.0075 | 0.9615±0.0054 |
| Disease name | Drug | Drug ID | Evidence |
|---|---|---|---|
| Alzheimer's disease | Benazepril | DB00542 | [ |
| Lisinopril | DB00722 | [ | |
| Rosuvastatin | DB01098 | [ | |
| Oxaprozin | DB00991 | [ | |
| Nizatidine | DB00585 | N | |
| Mesoridazine | DB00933 | N | |
| Simvastatin | DB00641 | [ | |
| Dextromethorphan | DB00514 | [ | |
| Diazepam | DB00829 | [ | |
| Sertraline | DB01104 | [ |
Tab.3 Top 10 candidate drugs for Alzheimer's disease predicted by DRLHG
| Disease name | Drug | Drug ID | Evidence |
|---|---|---|---|
| Alzheimer's disease | Benazepril | DB00542 | [ |
| Lisinopril | DB00722 | [ | |
| Rosuvastatin | DB01098 | [ | |
| Oxaprozin | DB00991 | [ | |
| Nizatidine | DB00585 | N | |
| Mesoridazine | DB00933 | N | |
| Simvastatin | DB00641 | [ | |
| Dextromethorphan | DB00514 | [ | |
| Diazepam | DB00829 | [ | |
| Sertraline | DB01104 | [ |
| Disease name | Drug | Drug ID | Evidence |
|---|---|---|---|
| Lung cancer | Diethylstilbestrol | DB00255 | [ |
| Rosuvastatin | DB01098 | [ | |
| Ofloxacin | DB01165 | [ | |
| Heparin | DB01109 | [ | |
| Levothyroxine | DB00451 | N | |
| Celecoxib | DB00482 | [ | |
| Valproic acid | DB00313 | [ | |
| Carvedilol | DB01136 | [ | |
| Orciprenaline | DB00816 | N | |
| Sulfasalazine | DB00795 | [ |
Tab.4 Top 10 candidate drugs for lung cancer predicted by DRLHG
| Disease name | Drug | Drug ID | Evidence |
|---|---|---|---|
| Lung cancer | Diethylstilbestrol | DB00255 | [ |
| Rosuvastatin | DB01098 | [ | |
| Ofloxacin | DB01165 | [ | |
| Heparin | DB01109 | [ | |
| Levothyroxine | DB00451 | N | |
| Celecoxib | DB00482 | [ | |
| Valproic acid | DB00313 | [ | |
| Carvedilol | DB01136 | [ | |
| Orciprenaline | DB00816 | N | |
| Sulfasalazine | DB00795 | [ |
| [1] | Ullah A, Meng YH. Finding influential nodes via graph embedding and hybrid centrality in complex networks[J]. Chaos Solitons Fractals, 2025, 194: 116151-8. doi:10.1016/j.chaos.2025.116151 |
| [2] | Aung HN, Ohsaki H. Node embedding accelerates randoms walk on a graph[C]//2024 IEEE 48th Annual Computers, Software, and Applications Conference, 2024: 537-545. doi:10.1109/compsac61105.2024.00079 |
| [3] | Chen H, Cheng F, Li J. iDrug: Integration of drug repositioning and drug-target prediction via cross-network embedding[J]. PLoS Comput Biol, 2020, 16(7): e1008040-9. doi:10.1371/journal.pcbi.1008040 |
| [4] | Sadeghi S, Lu J, Ngom A. A network-based drug repurposing method via non-negative matrix factorization[J]. Bioinformatics, 2022, 38(5): 1369-77. doi:10.1093/bioinformatics/btab826 |
| [5] | Song SP, Liu B, Teng F, et al. Self-supervised contrastive learning for implicit collaborative filtering[J]. Eng Appl Artif Intell, 2025, 139: 109563. doi:10.1016/j.engappai.2024.109563 |
| [6] | Zeng XX, Zhu SY, Lu WQ, et al. Target identification among known drugs by deep learning from heterogeneous networks[J]. Chem Sci, 2020, 11(7): 1775-97. doi:10.1039/c9sc04336e |
| [7] | Meng Y, Wang Y, Xu J, et al. Drug repositioning based on weighted local information augmented graph neural network[J]. Brief Bioinform, 2023, 25(1): bbad431. doi:10.1093/bib/bbad431 |
| [8] | Sun X, Wang B, Zhang J, et al. Partner-specific drug repositioning approach based on graph convolutional network[J]. IEEE J Biomed Health Inform, 2022, 26(11): 5757-65. doi:10.1109/jbhi.2022.3194891 |
| [9] | 吴雪扬, 张 煜, 张 华, 等.基于注意力机制和多模态特征融合的猕猴脑磁共振图像全脑分割[J].南方医科大学学报,2023, 43 (12): 2118-25. doi:10.12122/j.issn.1673-4254.2023.12.17 |
| [10] | Jia X, Sun X, Wang K, et al. DRGCL: drug repositioning via semantic-enriched graph contrastive learning[J]. IEEE J Biomed Health Inform, 2025, 29(3): 1656-67. doi:10.1109/jbhi.2024.3372527 |
| [11] | Peng L, Yang C, Yang J, et al. Drug repositioning via multi-view representation learning with heterogeneous graph neural network[J]. IEEE J Biomed Health Inform, 2025, 29(3): 1668-79. doi:10.1109/jbhi.2024.3434439 |
| [12] | Zhang F, Hu W, Liu Y. GCMM: graph convolution network based on multimodal attention mechanism for drug repurposing[J]. BMC Bioinformatics, 2022, 23(1): 372-83. doi:10.1186/s12859-022-04911-8 |
| [13] | Zhang M, Chen Y.Inductive matrix completion based on graph neural networks[J]. arXiv, 2019: 1904. 12058. |
| [14] | 洪 永, 张 鑫, 林铭俊, 等.基于深度可分离卷积与注意力机制的单导联心房颤动轻量级分类网络[J].南方医科大学学报, 2025, 45 (3): 650-60. |
| [15] | Maas AL, Hannun AY, Ng AY. Rectifier nonlinearities improve neural network acoustic models[C].Proc. International Conference on Machine Learning, 2013: 3. |
| [16] | Klambauer G, Unterthiner T, Mayr A, et al.Self-normalizing neural networks[J]. arXiv, 2017: 1706. 02515. doi:10.1016/j.toxlet.2017.07.175 |
| [17] | 王永贵, 于 琦.结合图同构和混合阶残差门控图神经网络的会话推荐[J].计算机科学与探索, 2025, 19 (2): 502-12. |
| [18] | Ying R, You J, Morris C, et al.Hierarchical graph representation learning with differentiable pooling[J]. arXiv, 2018: 1806.08804. |
| [19] | Barakat A, Bianchi P. Convergence and dynamical behavior of the ADAM algorithm for nonconvex stochastic optimization[J]. SIAM J Optim, 2021, 31(1): 244-74. doi:10.1137/19m1263443 |
| [20] | Poernomo A, Kang DK. Biased dropout and crossmap dropout: learning towards effective dropout regularization in convolutional neural network[J]. Neural Netw, 2018, 104: 60-7. doi:10.1016/j.neunet.2018.03.016 |
| [21] | Knox C, Wilson M, Klinger CM, et al. DrugBank 6.0: the DrugBank knowledgebase for 2024[J]. Nucleic Acids Res, 2024, 52(d1): D1265-75. doi:10.1093/nar/gkad976 |
| [22] | Hamosh A, Amberger JS, Bocchini C, et al. Online mendelian inheritance in man (OMIM®): victor McKusick's magnum opus[J]. Am J Med Genet A, 2021, 185(11): 3259-65. doi:10.1002/ajmg.a.62407 |
| [23] | Yang X, Yang G, Chu J. The neural metric factorization for computational drug repositioning[J]. IEEE/ACM Trans Comput Biol Bioinform, 2023, 20(1): 731-41. doi:10.1109/tcbb.2022.3144429 |
| [24] | Wang J, Wu Z, Peng Y, et al. Pathway-based drug repurposing with DPNetinfer: a method to predict drug-pathway associations via network-based approaches[J]. J Chem Inf Model, 2021, 61(5): 2475-85. doi:10.1021/acs.jcim.1c00009 |
| [25] | Zhao BW, Wang L, Hu PW, et al. Fusing higher and lower-order biological information for drug repositioning via graph representation learning[J]. IEEE Trans Emerg Topics Comput, 2024, 12(1): 163-76. doi:10.1109/tetc.2023.3239949 |
| [26] | Meng Y, Lu C, Jin M, et al. A weighted bilinear neural collaborative filtering approach for drug repositioning[J]. Brief Bioinform, 2022, 23(2): bbab581. doi:10.1093/bib/bbab581 |
| [27] | Huang SH, Wang MH, Zheng X, et al. Hierarchical and dynamic graph attention network for drug-disease association prediction[J]. IEEE J Biomed Health Inform, 2024, 28(4): 2416-27. doi:10.1109/jbhi.2024.3363080 |
| [28] | Yang MH, Ho TC, Chang CC, et al. Utilizing proteomic approaches to uncover the neuroprotective effects of ACE inhibitors: implications for Alzheimer's disease treatment[J]. Molecules, 2023, 28(16): 5938. doi:10.3390/molecules28165938 |
| [29] | Thomas J, Smith H, Smith CA, et al. The angiotensin-converting enzyme inhibitor lisinopril mitigates memory and motor deficits in a Drosophila model of Alzheimer's disease[J]. Pathophysiology, 2021, 28(2): 307-19. doi:10.3390/pathophysiology28020020 |
| [30] | Modiri A, Abdolmaleki Z, Paryani MR. The effect of rosuvastatin coated by nano-chitosan on developing hippocampus: association with hippocampal neurogenesis and memory in an Alzheimer's induced model of rats[J]. Anat Cell Biol, 2025, 58(1): 61-75. doi:10.5115/acb.24.250 |
| [31] | Willems S, Busch R, Nawa F, et al. Structural optimization of oxaprozin for selective inverse Nurr1 agonism[J]. J Med Chem, 2024, 67(15): 13324-48. doi:10.1021/acs.jmedchem.4c01218 |
| [32] | Salari A, Roghani M, Khalili M. HMG-CoA reductase inhibitor simvastatin ameliorates trimethyltin neurotoxicity and cognitive impairment through reversal of Alzheimer's-associated markers[J]. Metab Brain Dis, 2024, 40(1): 74. doi:10.1007/s11011-024-01515-4 |
| [33] | Majhi PK, Sayyad S, Gaur M, et al. Tinospora cordifolia extract enhances dextromethorphan bioavailability: implications for Alzheimer's disease[J]. ACS Omega, 2024, 9(22): 23634-48. doi:10.1021/acsomega.4c01219 |
| [34] | Chen JW, Zhang M, Shen ZY, et al. Low-dose diazepam improves cognitive function in APP/PS1 mouse models: Involvement of AMPA receptors[J]. Brain Res, 2024, 1845: 149207. doi:10.1016/j.brainres.2024.149207 |
| [35] | Wang H, Li S, Zhang J, et al. Efficacy of selective serotonin reuptake inhibitors-related antidepressants in Alzheimer's disease: a meta-analysis[J]. Eur J Med Res, 2024, 29(1): 438. doi:10.1186/s40001-024-02006-z |
| [36] | Seo Y, Jeong SB, Woo JH, et al. Diethylstilbestrol, a novel ANO1 inhibitor, exerts an anticancer effect on non-small cell lung cancer via inhibition of ANO1[J]. Int J Mol Sci, 2021, 22(13): 7100. doi:10.3390/ijms22137100 |
| [37] | Aksoy HN, Ceylan C. Comparison of the effects of statins on A549 nonsmall-cell lung cancer cell line lipids using Fourier transform infrared spectroscopy: rosuvastatin stands out[J]. Lipids, 2021, 56(3): 289-99. doi:10.1002/lipd.12296 |
| [38] | Schuette W, Nagel S, von Weikersthal LF, et al. Randomized phase III trial of docetaxel plus carboplatin with or without levofloxacin prophylaxis in elderly patients with advanced non-small cell lung cancer: the APRONTA trial[J]. J Thorac Oncol, 2011, 6(12): 2090-6. doi:10.1097/jto.0b013e3182307e3c |
| [39] | Bendahl PO, Belting M, Gezelius E. Longitudinal assessment of circulating tumor cells and outcome in small cell lung cancer: a sub-study of RASTEN-a randomized trial with low molecular weight heparin[J]. Cancers: Basel, 2023, 15(12): 3176. doi:10.3390/cancers15123176 |
| [40] | Qadir A, Khalid Z, Kashan Theba F, et al. Celecoxib and bevacizumab synergistically inhibit non-small cell lung cancer by inducing apoptosis and modulating VEGF and MMP-9 expression[J]. Pak J Pharm Sci, 2023, 36(2): 501-6. |
| [41] | Hu M, Cheng H, Yang Y, et al. Valproic acid increased the efficacy of EGFR TKIs on EGFR/TP53 co-mutated lung cancers and downregulated mutant-p53 levels[J]. Mol Carcinog, 2024, 63(2): 275-85. doi:10.1002/mc.23651 |
| [42] | Ikhmais BA, Hammad AM, Abusara OH, et al. Investigating carvedilol's repurposing for the treatment of non-small cell lung cancer via aldehyde dehydrogenase activity modulation in the presence of β-adrenergic agonists[J]. Curr Issues Mol Biol, 2023, 45(10): 7996-8012. doi:10.3390/cimb45100505 |
| [43] | Bagherpoor AJ, Shameem M, Luo X, et al. Inhibition of lung adenocarcinoma by combinations of sulfasalazine (SAS) and disulfiram-copper (DSF-Cu) in cell line models and mice[J]. Carcinogenesis, 2023, 44(4): 291-303. doi:10.1093/carcin/bgad020 |
| No related articles found! |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||