Journal of Southern Medical University ›› 2025, Vol. 45 ›› Issue (5): 1063-1073.doi: 10.12122/j.issn.1673-4254.2025.05.20
Previous Articles Next Articles
Xiaojuan GUO1( ), Ruijuan DU1,2, Liping CHEN1,2, Kelei GUO1,2, Biao ZHOU1,2, Hua BIAN1,2, Li HAN1,2(
), Ruijuan DU1,2, Liping CHEN1,2, Kelei GUO1,2, Biao ZHOU1,2, Hua BIAN1,2, Li HAN1,2( )
)
Received:2024-08-13
Online:2025-05-20
Published:2025-05-23
Contact:
Li HAN
E-mail:3152044@nyist.edu.cn;hanli@nyist.edu.cn
Supported by:Xiaojuan GUO, Ruijuan DU, Liping CHEN, Kelei GUO, Biao ZHOU, Hua BIAN, Li HAN. WW domain-containing ubiquitin E3 ligase 1 regulates immune infiltration in tumor microenvironment of ovarian cancer[J]. Journal of Southern Medical University, 2025, 45(5): 1063-1073.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.j-smu.com/EN/10.12122/j.issn.1673-4254.2025.05.20
| Characteristics | Low expression of WWP1 | High expression of WWP1 | P |
|---|---|---|---|
| Case (n) | 185 | 187 | |
| Figo Clinical stage [n (%)] | 0.0012 | ||
| Stage III | 152 (40.9%) | 139 (37.4%) | |
| Stage IV | 20 (5.4%) | 37 (9.9%) | |
| Stage II | 11 (2.9%) | 12 (3.2%) | |
| Stage I | 1 (0.2%) | - | |
| Primary therapy outcome [n (%)] | <0.0001 | ||
| PD | 13 (3.5%) | 11 (3.0%) | |
| SD | 11 (3.0%) | 11 (3.0%) | |
| PR | 18 (4.8%) | 25 (6.7%) | |
| CR | 109 (29.3%) | 101 (27.2%) | |
| Race [n (%)] | 0.846 | ||
| Asian | 7 (1.6%) | 4 (1.6%) | |
| Black or african american | 14 (3.8%) | 11 (3%) | |
| White | 155 (41.7%) | 168 (45.2%) | |
| Age [median (IQR)] | 61 (51, 71) | 58 (51, 65) | 0.039 |
Tab.1 Clinical characteristics of ovarian cancer patients with low and high WWP1 expression levels
| Characteristics | Low expression of WWP1 | High expression of WWP1 | P |
|---|---|---|---|
| Case (n) | 185 | 187 | |
| Figo Clinical stage [n (%)] | 0.0012 | ||
| Stage III | 152 (40.9%) | 139 (37.4%) | |
| Stage IV | 20 (5.4%) | 37 (9.9%) | |
| Stage II | 11 (2.9%) | 12 (3.2%) | |
| Stage I | 1 (0.2%) | - | |
| Primary therapy outcome [n (%)] | <0.0001 | ||
| PD | 13 (3.5%) | 11 (3.0%) | |
| SD | 11 (3.0%) | 11 (3.0%) | |
| PR | 18 (4.8%) | 25 (6.7%) | |
| CR | 109 (29.3%) | 101 (27.2%) | |
| Race [n (%)] | 0.846 | ||
| Asian | 7 (1.6%) | 4 (1.6%) | |
| Black or african american | 14 (3.8%) | 11 (3%) | |
| White | 155 (41.7%) | 168 (45.2%) | |
| Age [median (IQR)] | 61 (51, 71) | 58 (51, 65) | 0.039 |
| Characteristics | Case (n) | Univariate analysis | Multivariate analysis | |||
|---|---|---|---|---|---|---|
| Hazard ratio (95% CI) | P | Hazard ratio (95% CI) | P | |||
| WWP1 | 372 | |||||
| Low | 185 | Reference | Reference | |||
| High | 187 | 1.499 (1.156-1.942) | 0.002 | 0.024 | ||
| Clinical stage | 372 | |||||
| I & II | 24 | Reference | ||||
| III | 291 | 2.058 (0.911-4.649) | 0.083 | |||
| IV | 57 | 2.556 (1.085-6.025) | 0.032 | |||
| Tumor status | 337 | |||||
| Tumor free | 72 | Reference | Reference | |||
| With tumor | 265 | 9.598 (4.487-20.532) | < 0.001 | 15.691 (3.811-64.606) | <0.001 | |
| Primary therapy outcome | 299 | < 0.001 | ||||
| PD | 24 | Reference | Reference | |||
| SD | 22 | 0.441 (0.217-0.896) | 0.024 | 0.397 (0.187-0.845) | 0.016 | |
| PR | 43 | 0.652 (0.384-1.108) | 0.114 | 0.659 (0.374-1.160) | 0.148 | |
| CR | 210 | 0.154 (0.095-0.250) | <0.001 | 0.179 (0.106-0.302) | <0.001 | |
| Tumor residual | 336 | |||||
| No | 68 | Reference | ||||
| Yes | 268 | 2.223 (1.441-3.430) | <0.001 | |||
| Age (year) | 372 | |||||
| ≤60 | 207 | Reference | ||||
| >60 | 165 | 1.352 (1.045-1.749) | 0.022 | |||
Tab.2 Univariate and multivariate analyses of the risk factors for poor prognosis of ovarian cancer patients
| Characteristics | Case (n) | Univariate analysis | Multivariate analysis | |||
|---|---|---|---|---|---|---|
| Hazard ratio (95% CI) | P | Hazard ratio (95% CI) | P | |||
| WWP1 | 372 | |||||
| Low | 185 | Reference | Reference | |||
| High | 187 | 1.499 (1.156-1.942) | 0.002 | 0.024 | ||
| Clinical stage | 372 | |||||
| I & II | 24 | Reference | ||||
| III | 291 | 2.058 (0.911-4.649) | 0.083 | |||
| IV | 57 | 2.556 (1.085-6.025) | 0.032 | |||
| Tumor status | 337 | |||||
| Tumor free | 72 | Reference | Reference | |||
| With tumor | 265 | 9.598 (4.487-20.532) | < 0.001 | 15.691 (3.811-64.606) | <0.001 | |
| Primary therapy outcome | 299 | < 0.001 | ||||
| PD | 24 | Reference | Reference | |||
| SD | 22 | 0.441 (0.217-0.896) | 0.024 | 0.397 (0.187-0.845) | 0.016 | |
| PR | 43 | 0.652 (0.384-1.108) | 0.114 | 0.659 (0.374-1.160) | 0.148 | |
| CR | 210 | 0.154 (0.095-0.250) | <0.001 | 0.179 (0.106-0.302) | <0.001 | |
| Tumor residual | 336 | |||||
| No | 68 | Reference | ||||
| Yes | 268 | 2.223 (1.441-3.430) | <0.001 | |||
| Age (year) | 372 | |||||
| ≤60 | 207 | Reference | ||||
| >60 | 165 | 1.352 (1.045-1.749) | 0.022 | |||
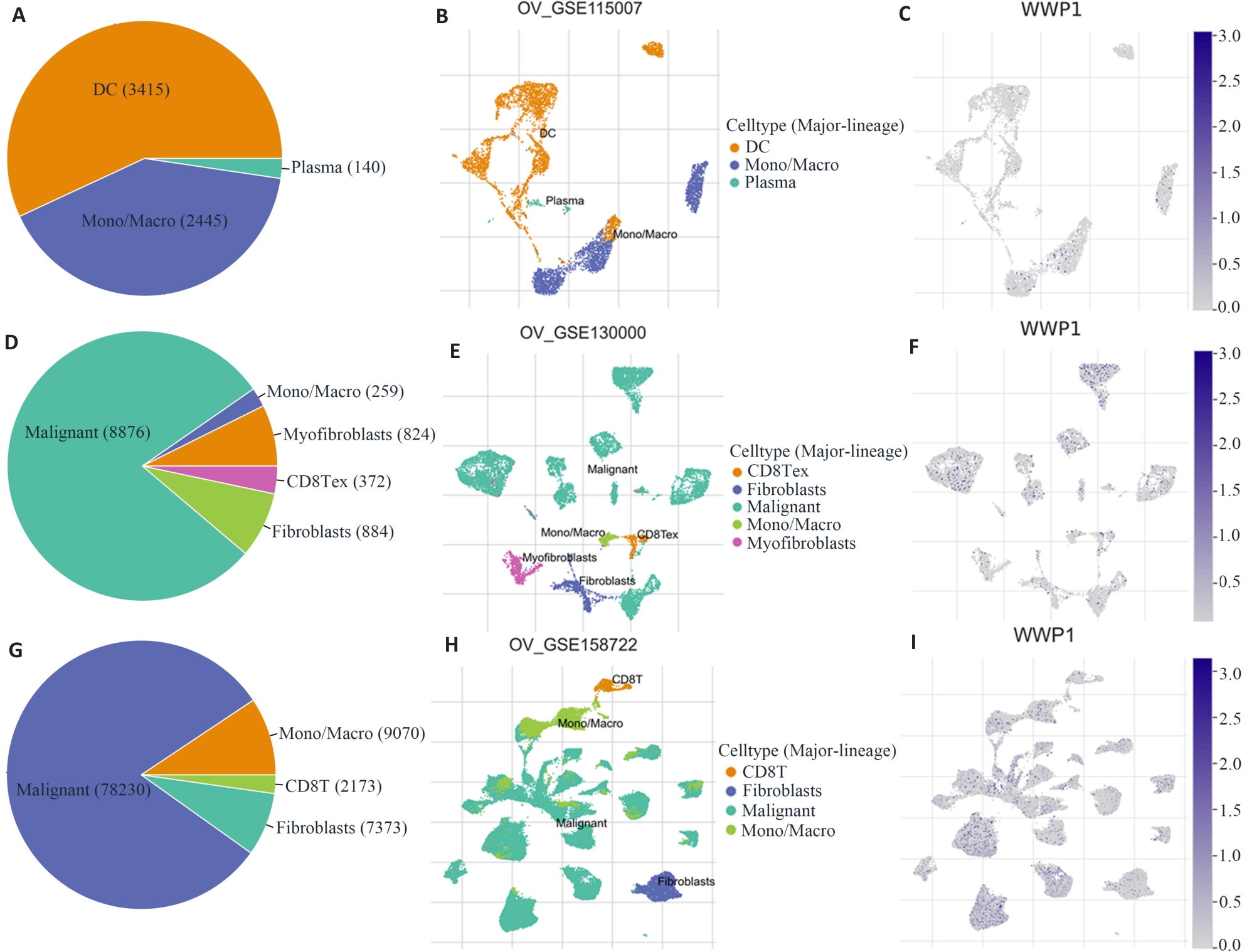
Fig.2 Analysis of WWP1-related cell type distribution in primary tumor, primary plus metastatic tumor, and primary tumor plus chemotherapy using scRNA seq database. A, B, D, E, G, H: Cell types and their distribution. C, F, I: Distribution of WWP1 in different cells in OV_GSE115007, OV_GSE130000 and OV_GSE158722 datasets.

Fig.3 Correlation between immune infiltration and WWP1 expression in ovarian cancer. XCELL, CIBERSORT, CIBERSORT-ABS, QUANTISEQ, EPIC, MCPCOUNTER and TIMER are different immune infiltration algorithms.
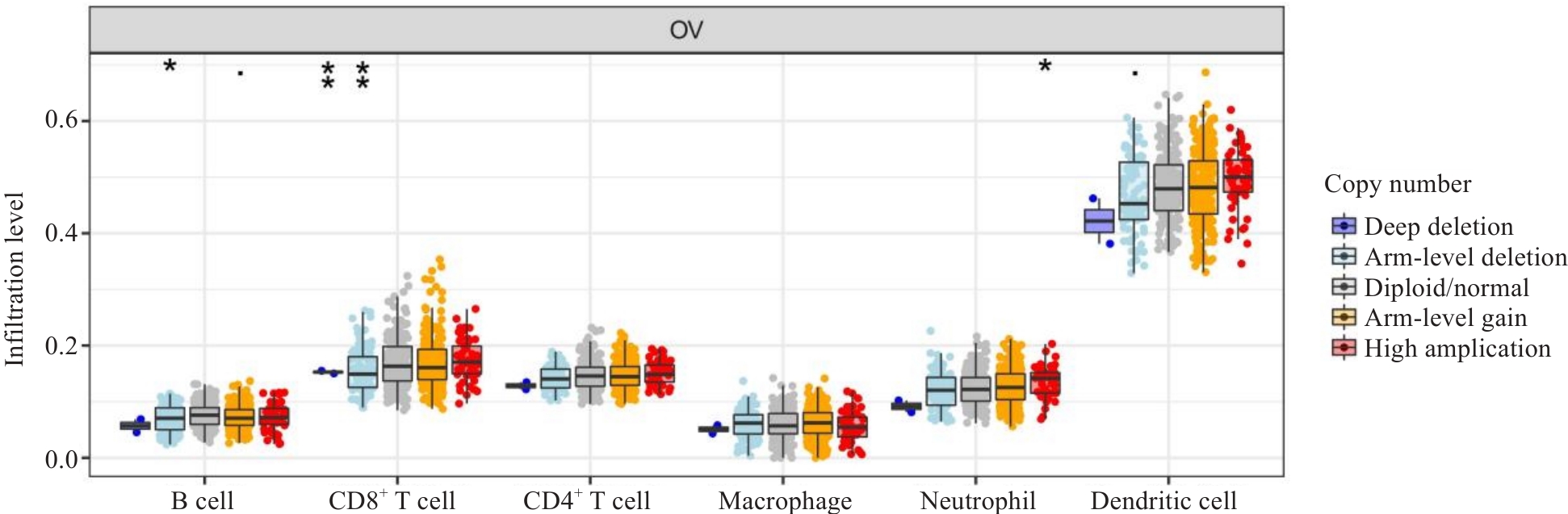
Fig.4 Correlation between immune infiltration and WWP1 expression in ovarian cancer. XCELL,CIBERSORT, CIBERSORT-ABS, QUANTISEQ, EPIC, MCPCOUNTER and TIMER are different immune infiltration algorithms. *P <0.05, **P<0.01.
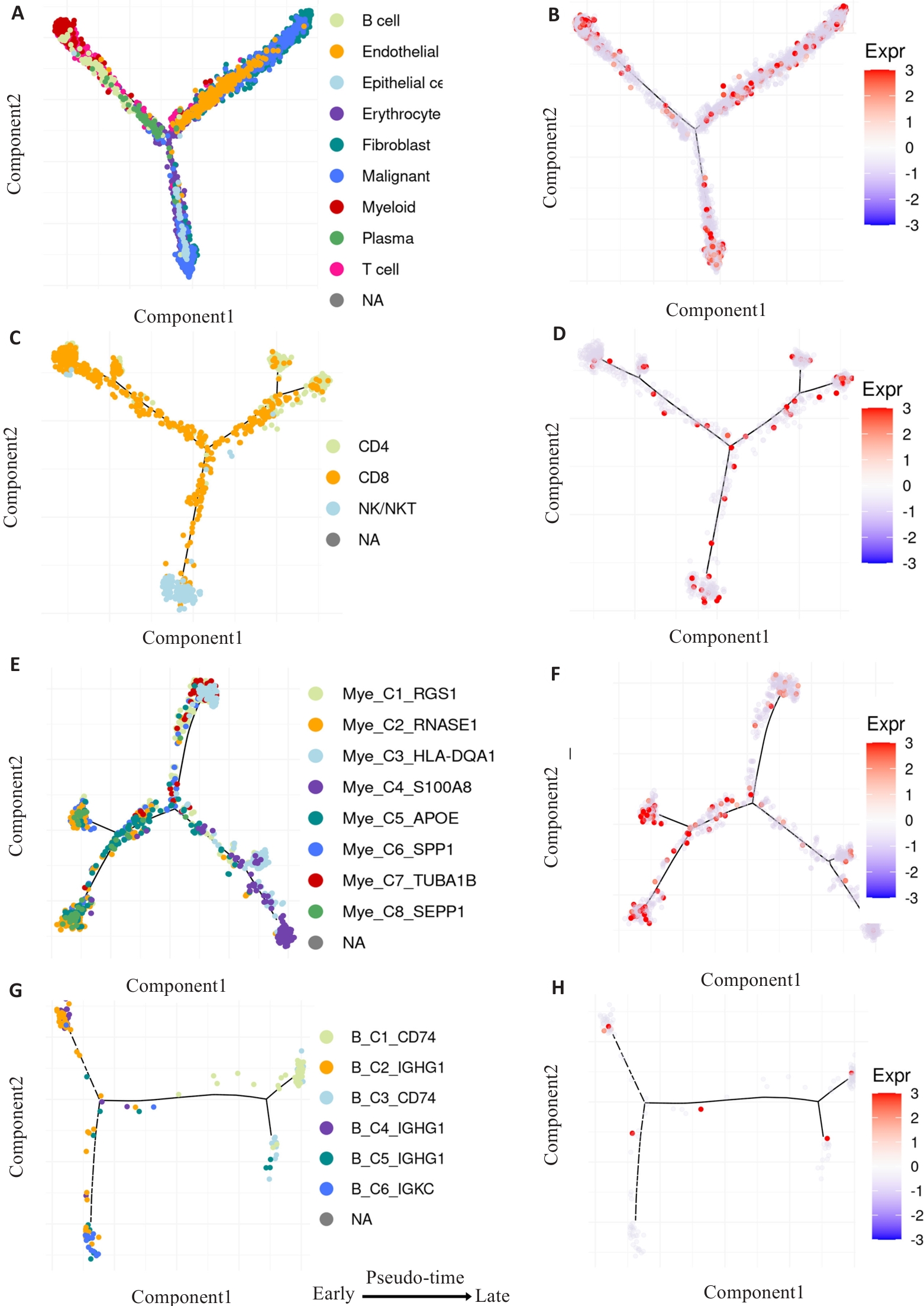
Fig.5 Pseudo-time analysis of the effect of WWP1 expression on dynamic changes of infiltrating immune cells in ovarian cancer. A, C, E, G: Developmental trajectories of the pooled infiltrating immune cells, CD4+ T cells, CD8+ T cells, NK cells, myeloid cells and B cells (The inferred direction to differentiation and maturation was from the left to the right). B, D, F, H: Dynamic expressions of WWP1 related to the differentiation and maturation along the pseudo-time axis.
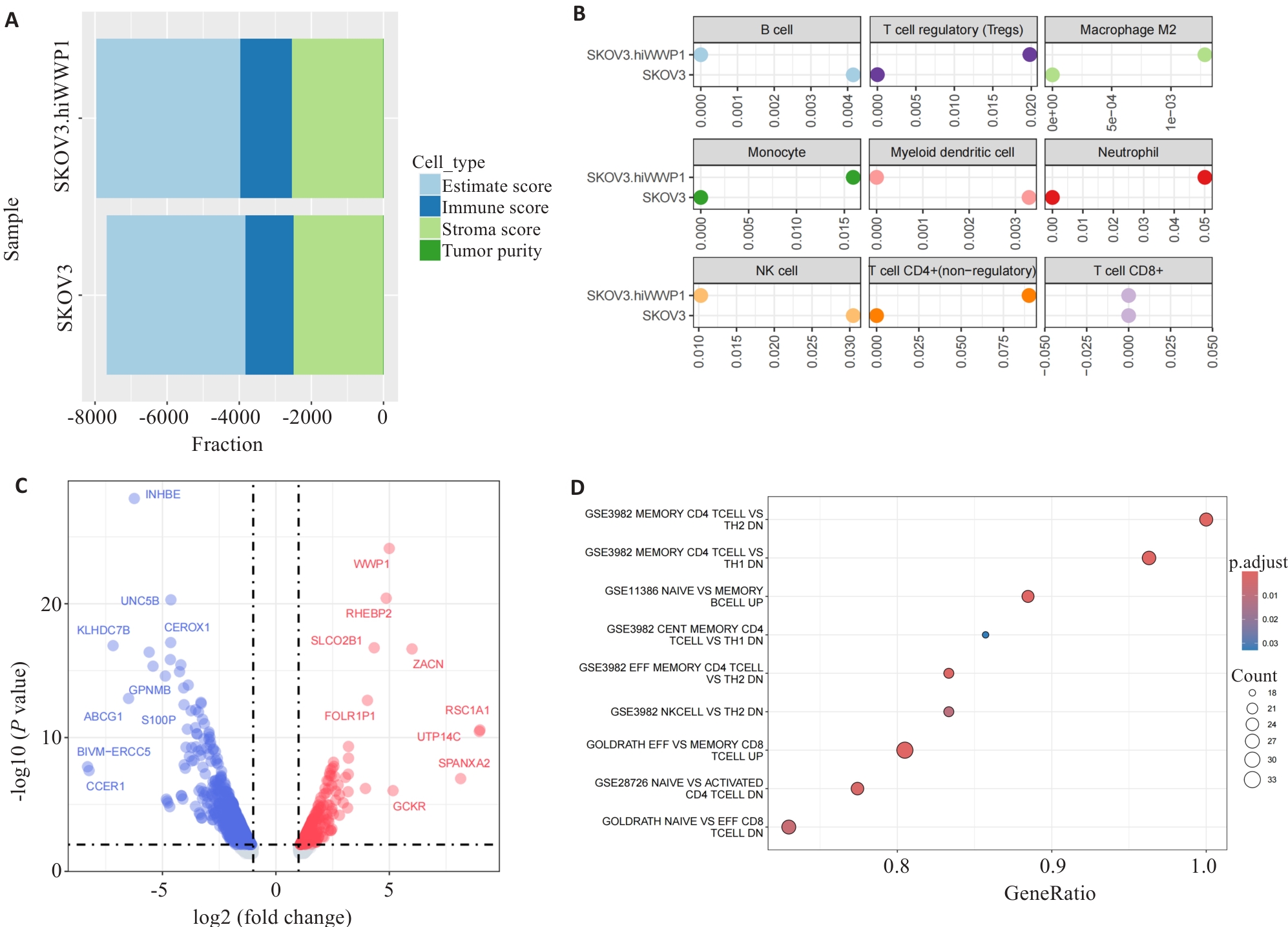
Fig.7 Effect of WWP1 overexpression on immune infiltration in SKOV3 cells. A: Comparison of immune microenvironment. B: Comparison of immune infiltration. C: Volcano map for Top 20 difference genes. D: Comparison of immune pathways of GSEA enrichment.
| 1 | 中国抗癌协会妇科肿瘤专业委员会. 卵巢恶性肿瘤诊断与治疗指南(2021年版)[J]. 中国癌症杂志, 2021, 31(6): 490-500. DOI: 10.19538/j.fk2021060111 |
| 2 | Sosinsky A, Ambrose J, Cross W, et al. Insights for precision oncology from the integration of genomic and clinical data of 13, 880 tumors from the 100, 000 Genomes Cancer Programme[J]. Nat Med, 2024, 30(1): 279-89. |
| 3 | Liu J, Berchuck A, Backes FJ, et al. NCCN guidelines® insights: ovarian cancer/fallopian tube cancer/primary peritoneal cancer, version 3.2024[J]. J Natl Compr Canc Netw, 2024, 22(8): 512-9. |
| 4 | Zou XQ, Zhao YJ, Liang XT, et al. Double insurance for OC: miRNA-mediated platinum resistance and immune escape[J]. Front Immunol, 2021, 12: 641937. |
| 5 | Xu JF, Fang YF, Chen KL, et al. Single-cell RNA sequencing reveals the tissue architecture in human high-grade serous ovarian cancer[J]. Clin Cancer Res, 2022, 28(16): 3590-602. |
| 6 | Du YH, Shi JT, Wang JX, et al. Integration of pan-cancer single-cell and spatial transcriptomics reveals stromal cell features and therapeutic targets in tumor microenvironment[J]. Cancer Res, 2024, 84(2): 192-210. |
| 7 | El-Tanani M, Rabbani SA, Babiker R, et al. Unraveling the tumor microenvironment: Insights into cancer metastasis and therapeutic strategies[J]. Cancer Lett, 2024, 591: 216894. |
| 8 | Zhong GH, Zhao DS, Li JW, et al. WWP1 deficiency alleviates cardiac remodeling induced by simulated microgravity[J]. Front Cell Dev Biol, 2021, 9: 739944. |
| 9 | Novelli G, Liu J, Biancolella M, et al. Inhibition of HECT E3 ligases as potential therapy for COVID-19[J]. Cell Death Dis, 2021, 12(4): 310. |
| 10 | Kishikawa T, Higuchi H, Wang LM, et al. WWP1 inactivation enhances efficacy of PI3K inhibitors while suppressing their toxicities in breast cancer models[J]. J Clin Invest, 2021, 131(24): e140436. |
| 11 | Jiang H, Li ZX, Xu W, et al. WWP1 targeting PTEN for polyubiquitination to promote bone metastasis of luminal breast cancer[J]. Sci Rep, 2024, 14(1): 29950. |
| 12 | Ye P, Chi XX, Cha JH, et al. Potential of E3 ubiquitin ligases in cancer immunity: opportunities and challenges[J]. Cells, 2021, 10(12): 3309. |
| 13 | Schrock MS, Stromberg BR, Scarberry L, et al. APC/C ubiquitin ligase: Functions and mechanisms in tumorigenesis[J]. Semin Cancer Biol, 2020, 67(Pt 2): 80-91. |
| 14 | Liao CH, Yu LP, Pang Z, et al. WWP1 targeting MUC1 for ubiquitin-mediated lysosomal degradation to suppress carcinogenesis[J]. Signal Transduct Target Ther, 2021, 6(1): 297. |
| 15 | Han Y, Wang YT, Dong X, et al. TISCH2: expanded datasets and new tools for single-cell transcriptome analyses of the tumor microenvironment[J]. Nucleic Acids Res, 2023, 51(D1): D1425-31. |
| 16 | Chen ZH, Luo ZW, Zhang D, et al. TIGER A web portal of tumor immunotherapy gene expression resource[J]. Genomics Proteomics Bioinformatics, 2023, 21(2): 337-48. |
| 17 | Qian JB, Olbrecht S, Boeckx B, et al. A pan-cancer blueprint of the heterogeneous tumor microenvironment revealed by single-cell profiling[J]. Cell Res, 2020, 30(9): 745-62. |
| 18 | Rong RC, Sheng H, Jin KW, et al. A deep learning approach for histology-based nucleus segmentation and tumor microenvironment characterization[J]. Mod Pathol, 2023, 36(8): 100196. |
| 19 | Wu TZ, Hu EQ, Xu SB, et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data[J]. Innovation, 2021, 2(3): 100141. |
| 20 | 韩 立, 郭晓娟, 杜瑞娟, 等. 芍药内酯苷对人卵巢癌多药耐药的逆转作用[J]. 中国药理学通报, 2023, 39(2): 268-74. DOI: 10.12360/CPB202102039 |
| 21 | Wehrli M, Guinn S, Birocchi F, et al. Mesothelin CAR T cells secreting anti-FAP/anti-CD3 molecules efficiently target pancreatic adenocarcinoma and its stroma[J]. Clin Cancer Res, 2024, 30(9): 1859-77. |
| 22 | Luo YK, Xia Y, Liu D, et al. Neoadjuvant PARPi or chemotherapy in ovarian cancer informs targeting effector Treg cells for homologous-recombination-deficient tumors[J]. Cell, 2024, 187(18): 4905-25.e24. |
| 23 | de Visser KE, Joyce JA. The evolving tumor microenvironment: From cancer initiation to metastatic outgrowth[J]. Cancer Cell, 2023, 41(3): 374-403. |
| 24 | 关深元, 沈智勇, 林名岛, 等. 泛癌组织STIP1的表达与肿瘤免疫浸润及预后相关: 基于生物信息学方法[J]. 南方医科大学学报, 2023, 43(7): 1179-93. DOI: 10.12122/j.issn.1673-4254.2023.07.15 |
| 25 | Malmgren JA, Mayer M, Atwood MK, et al. Differential presentation and survival of de novo and recurrent metastatic breast cancer over time: 1990-2010[J]. Breast Cancer Res Treat, 2018, 167(2): 579-90. |
| 26 | Karagiannis GS, Pastoriza JM, Wang YR, et al. Neoadjuvant chemotherapy induces breast cancer metastasis through a TMEM-mediated mechanism[J]. Sci Transl Med, 2017, 9(397): eaan0026. |
| 27 | Long XH, Zhang SL, Wang YL, et al. Targeting JMJD1C to selectively disrupt tumor Treg cell fitness enhances antitumor immunity[J]. Nat Immunol, 2024, 25(3): 525-36. |
| 28 | Gu Y, Liu YF, Fu L, et al. Tumor-educated B cells selectively promote breast cancer lymph node metastasis by HSPA4-targeting IgG[J]. Nat Med, 2019, 25(2): 312-22. |
| 29 | Wang B, Xiong YQ, Li R, et al. Shorter telomere length increases the risk of lymphocyte immunodeficiency: a Mendelian randomization study[J]. Immun Inflamm Dis, 2024, 12(4): e1251. |
| 30 | Hollern D. Memory B cell fitness and anergy has significant links to cancer lethality[J]. Cell, 2024, 187(17): 4551-3. |
| 31 | Zeng Q, Mousa M, Nadukkandy AS, et al. Understanding tumour endothelial cell heterogeneity and function from single-cell omics[J]. Nat Rev Cancer, 2023, 23(8): 544-64. |
| 32 | Ng MSF, Kwok I, Tan L, et al. Deterministic reprogramming of neutrophils within tumors[J]. Science, 2024, 383(6679): eadf6493. |
| 33 | Wen ZW, Sun HY, Zhang ZH, et al. High baseline tumor burden-associated macrophages promote an immunosuppressive microenvironment and reduce the efficacy of immune checkpoint inhibitors through the IGFBP2-STAT3-PD-L1 pathway[J]. Cancer Commun, 2023, 43(5): 562-81. |
| 34 | Tang PC, Chung JY, Xue VW, et al. Smad3 promotes cancer-associated fibroblasts generation via macrophage-myofibroblast transition[J]. Adv Sci, 2022, 9(1): e2101235. |
| 35 | Alspach E, Lussier DM, Miceli AP, et al. MHC-II neoantigens shape tumour immunity and response to immunotherapy[J]. Nature, 2019, 574(7780): 696-701. |
| 36 | Dean I, Lee CYC, Tuong ZK, et al. Rapid functional impairment of natural killer cells following tumor entry limits anti-tumor immunity[J]. Nat Commun, 2024, 15(1): 683. |
| 37 | Spranger S, Dai D, Horton B, et al. Tumor-residing Batf3 dendritic cells are required for effector T cell trafficking and adoptive T cell therapy[J]. Cancer Cell, 2017, 31(5): 711-23.e4. |
| 38 | Zheng ST, Ma JJ, Li JN, et al. Lower PTEN may be associated with CD8+ T cell exhaustion in diffuse large B-cell lymphoma[J]. Hum Immunol, 2023, 84(10): 551-60. |
| 39 | Exposito F, Redrado M, Houry M, et al. PTEN loss confers resistance to anti-PD-1 therapy in non-small cell lung cancer by increasing tumor infiltration of regulatory T cells[J]. Cancer Res, 2023, 83(15): 2513-26. |
| 40 | Nian ZG, Dou YC, Shen YQ, et al. Interleukin-34-orchestrated tumor-associated macrophage reprogramming is required for tumor immune escape driven by p53 inactivation[J]. Immunity, 2024, 57(10): 2344-61. e7. |
| 41 | Mathieu NA, Levin RH, Spratt DE. Exploring the roles of HERC2 and the NEDD4L HECT E3 ubiquitin ligase subfamily in p53 signaling and the DNA damage response[J]. Front Oncol, 2021, 11: 659049. |
| 42 | Borna S, Drobek A, Kralova J, et al. Transmembrane adaptor protein WBP1L regulates CXCR4 signalling and murine haematopoiesis[J]. J Cell Mol Med, 2020, 24(2): 1980-92. |
| 43 | Zhu GQ, Tang Z, Huang R, et al. CD36+ cancer-associated fibroblasts provide immunosuppressive microenvironment for hepatocellular carcinoma via secretion of macrophage migration inhibitory factor[J]. Cell Discov, 2023, 9(1): 25. |
| 44 | Sakata T, Yoshio S, Yamazoe T, et al. Immunoglobulin-like transcript 2 as an impaired anti-tumor cytotoxicity marker of natural killer cells in patients with hepatocellular carcinoma[J]. Front Immunol, 2024, 15: 1389411. |
| 45 | Chen Q, Shen MY, Yan M, et al. Targeting tumor-infiltrating CCR8+ regulatory T cells induces antitumor immunity through functional restoration of CD4+ Tconvs and CD8+ T cells in colorectal cancer[J]. J Transl Med, 2024, 22(1): 709. |
| 46 | da Silva SF, Murta EF, Michelin MA. ICAM2 is related to good prognosis in dendritic cell immunotherapy for cancer[J]. Immunotherapy, 2024, 16(3): 173-85. |
| [1] | Jinlong PANG, Xinli ZHAO, Zhen ZHANG, Haojie WANG, Xingqi ZHOU, Yumei YANG, Shanshan LI, Xiaoqiang CHANG, Feng LI, Xian LI. Overexpression of multimerin-2 promotes cutaneous melanoma cell invasion and migration and is associated with poor prognosis [J]. Journal of Southern Medical University, 2025, 45(7): 1479-1489. |
| [2] | Zhi GAO, Ao WU, Zhongxiang HU, Peiyang SUN. Bioinformatics analysis of oxidative stress and immune infiltration in rheumatoid arthritis [J]. Journal of Southern Medical University, 2025, 45(4): 862-870. |
| [3] | Yaobin WANG, Liuyan CHEN, Yiling LUO, Jiqing SHEN, Sufang ZHOU. Predictive value of NUF2 for prognosis and immunotherapy responses in pan-cancer [J]. Journal of Southern Medical University, 2025, 45(1): 137-149. |
| [4] | Zihan LU, Fangjun HUANG, Guangyao CAI, Jihong LIU, Xin ZHEN. A multi-constraint representation learning model for identification of ovarian cancer with missing laboratory indicators [J]. Journal of Southern Medical University, 2025, 45(1): 170-178. |
| [5] | Qinzhi WANG, Bing SONG, Shirui HAO, Zhiyuan XIAO, Lianhui JIN, Tong ZHENG, Fang CHAI. Bioinformatic analysis of CCND2 expression in papillary thyroid carcinoma and its impact on immune infiltration [J]. Journal of Southern Medical University, 2024, 44(5): 981-988. |
| [6] | SHAO Shan, BAI Weichao, ZHOU Pengcheng, LUO Minna, ZHAO Xinhan, LEI Jianjun. Metformin suppresses hypoxia-inducible factor-1α expression in cancer-associated fibroblasts to block tumor-stromal cross-talk in breast cancer [J]. Journal of Southern Medical University, 2024, 44(3): 428-436. |
| [7] | Fuxing ZHANG, Guoqing LIU, Rui DONG, Lei GAO, Weichen LU, Lianxia GAO, Zhongkuo ZHAO, Fei LU, Mulin LIU. High expression of CRTAC1 promotes proliferation, migration and immune cell infiltration of gastric cancer by regulating the PI3K/AKT signaling pathway [J]. Journal of Southern Medical University, 2024, 44(12): 2421-2433. |
| [8] | GUO Xiaojuan, CHEN Liping, LÜ Qin, DU Ruijuan, LUO Qin, ZHANG Yang, BIAN Hua, HAN Li. Guizhi Fuling Capsule inhibits migration and induces apoptosis of human ovarian cancer cells by regulating the NF-κB signaling pathway [J]. Journal of Southern Medical University, 2023, 43(8): 1315-1321. |
| [9] | DENG Ting, DU Boyu, XI Xueyan. Colorectal cancer cells induce the formation of cancer-associated fibroblasts by activating the ERK signaling pathway in fibroblasts [J]. Journal of Southern Medical University, 2023, 43(6): 943-951. |
| [10] | ZHANG Ziran, TAN Jiale, YU Zihang, LIU Chengdong, WANG Jian, WU Dehua, BAI Xue. FARSB stratifies prognosis and cold tumor microenvironment across different cancer types: an integrated single cell and bulk RNA sequencing analysis [J]. Journal of Southern Medical University, 2023, 43(5): 667-679. |
| [11] | MAO Jianying, YANG Wenjing, GUO He, DONG Ruili, REN Lifang, LI Shubin. A cervical cancer tissue-derived decellularized extracellular matrix scaffold for cervical cancer tissue reconstruction in vitro [J]. Journal of Southern Medical University, 2023, 43(2): 157-165. |
| [12] | ZHANG Weijian, ZOU Zhuoyue, ZHU Yongna, WANG Min, MA Caiyun, WU Junjie, SHI Xin, LIU Xi. Expression of interleukin-34 in tongue squamous cell carcinoma and its clinical implications [J]. Journal of Southern Medical University, 2023, 43(12): 2111-2117. |
| [13] | SU Lili, LIANG Wanqing, LÜ Zhenyu, HAN Xiao. PLXNA1 is highly expressed in hepatocellular carcinoma and affects patients' survival and immune microenvironment [J]. Journal of Southern Medical University, 2023, 43(11): 1909-1918. |
| [14] | ZHAO Haiyuan, LIU Gang, LI Yang, YANG Nianzhao, ZHAO Jun. High expression of ANKRD6 is an indicator for poor prognosis of colon adenocarcinoma [J]. Journal of Southern Medical University, 2023, 43(10): 1715-1724. |
| [15] | ZHANG Panyang, HE Mingmin, ZENG Yuanyuan, CAI Xiongwei. Identification of key molecules in miRNA-mRNA regulatory network associated with high-grade serous ovarian cancer recurrence using bioinformatic analysis [J]. Journal of Southern Medical University, 2023, 43(1): 8-16. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||