南方医科大学学报 ›› 2025, Vol. 45 ›› Issue (6): 1260-1269.doi: 10.12122/j.issn.1673-4254.2025.06.15
胡琼琼1( ), 李文沛1,2, 胥丽霞1, 官瑞磊1, 张东亚1, 蒋胶胶1, 王宁1, 杨改清1,2(
), 李文沛1,2, 胥丽霞1, 官瑞磊1, 张东亚1, 蒋胶胶1, 王宁1, 杨改清1,2( )
)
收稿日期:2024-12-05
出版日期:2025-06-20
发布日期:2025-06-27
通讯作者:
杨改清
E-mail:zxyyhqq0915@163.com;zxyyygq@163.com
作者简介:胡琼琼,主治医师,E-mail: zxyyhqq0915@163.com
基金资助:
Qiongqiong HU1( ), Wenpei LI1,2, Lixia XU1, Ruilei GUAN1, Dongya ZHANG1, Jiaojiao JIANG1, Ning WANG1, Gaiqing YANG1,2(
), Wenpei LI1,2, Lixia XU1, Ruilei GUAN1, Dongya ZHANG1, Jiaojiao JIANG1, Ning WANG1, Gaiqing YANG1,2( )
)
Received:2024-12-05
Online:2025-06-20
Published:2025-06-27
Contact:
Gaiqing YANG
E-mail:zxyyhqq0915@163.com;zxyyygq@163.com
摘要:
目的 探究跨膜蛋白240(TMEM240)的亚细胞定位及其生物学功能。 方法 用NCBI的BLAST工具对蛋白质序列分析,TMHMM生物信息学软件对TMEM240跨膜结构进行分析;体内实验:取18~20 d的雄性C57BL/6小鼠脑组织,设置TMEM240探针组和对照组(3只/组),采用原位杂交技术验证TMEM240在脑组织中的表达分布;采用qPCR、Western blotting检测小鼠不同组织、不同时间皮层神经元TMEM240的表达情况(3只/组);体外实验:采用质粒转染完成Tmem240过表达体系的构建及稳定克隆的筛选,采用细胞成像技术对TMEM240的亚细胞定位进行研究,确定TMEM240在细胞内的形态和分布,采用qPCR,Western blotting,免疫荧光检测TMEM240在细胞内表达情况并验证其生物学功能; 结果 人源和鼠源TMEM240蛋白质序列相似性达97.69%,TMEM240为含有两个跨膜结构的跨膜蛋白;体内实验:TMEM240的mRNA和蛋白在脑组织及神经元中特异性高表达(P<0.001),原代神经元培养9 d时TMEM240表达水平最高(P<0.001);体外实验:TMEM240在细胞内呈点状分布,属于细胞内膜性结构,并与过氧化物酶体具有80%共定位;过表达Tmem240的Neuro-2a细胞Rho GDP解离抑制因子β(ARHGDIB)的mRNA和蛋白表达均升高(P<0.05)。 结论 TMEM240是一种定位于细胞内的新型亚细胞结构,在神经元中特异性表达,在靶向细胞功能调控方面具有巨大的潜力。
胡琼琼, 李文沛, 胥丽霞, 官瑞磊, 张东亚, 蒋胶胶, 王宁, 杨改清. 神经特异性跨膜蛋白240与过氧化物酶体共定位并激活Rho GDP解离抑制因子β[J]. 南方医科大学学报, 2025, 45(6): 1260-1269.
Qiongqiong HU, Wenpei LI, Lixia XU, Ruilei GUAN, Dongya ZHANG, Jiaojiao JIANG, Ning WANG, Gaiqing YANG. Neurospecific transmembrane protein 240 colocalizes with peroxisomes and activates Rho GDP dissociation inhibitor β[J]. Journal of Southern Medical University, 2025, 45(6): 1260-1269.
| Gene | Sequences (5'-3') |
|---|---|
18S-F 18S-R | AGTCCCTGCCCTTTGTACACA CGATCCGAGGGCCTCACTA |
| Tmem240-F | TGCTTGATGGACATGAACGC |
| Tmem240-R | TAGTTCTCGGAGGCGTCCAC |
| Bex4_F | GGGGGAAATGTCCGAAGGAAA |
| Bex4_R | CCTGCACTACAAATCTCCCAAC |
| Fdft1_F | AGAGTGGCGGTTCACTGAGA |
| Fdft1_R | GAGAAAGGCCAATTCCCACCA |
| Slc2a3-F | TCATCAATGCACCTGAGACAATC |
| Slc2a3-R | GTCCCTCACTTGGTAGGTCTT |
| Arhgdib_F | CAACACACATACCGGACTGG |
| Arhgdib_R | TGTTTGTCGTCATCCGTGAAG |
| Insig1_F | CTAGTGCTCTTCTCATTTGGCG |
| Insig1_R | AGGGATACAGTAAACCGACAACA |
| Hmgcr_F | AGAGCGAGTGCATTAGCAAAG |
| Hmgcr_R | GATTGCCATTCCACGAGCTAT |
| Hmgcs1_F | AACTGGTGCAGAAATCTCTAGC |
| Hmgcs1_R | GGTTGAATAGCTCAGAACTAGCC |
| SPP1_F | AGCAAGAAACTCTTCCAAGCAA |
| SPP1_R | GTGAGATTCGTCAGATTCATCCG |
| Fam78b_F | TGCGCTACAAGACCCCTTACT |
| Fam78b_R | CTGATGGCTTTTACTCTCCCTTC |
| Arhgdia-F | AAGGACGATGAAAGCCTCCG |
| Arhgdia-R | GGTCAGTCGAGTCACAATGACA |
| Arhgdig-F | GTCAACTCCATCAGATGAGGTG |
| Arhgdig-R | GGGGTCCATAATGGGTGGC |
表1 引物列表
Tab.1 Primer sequences for qPCR
| Gene | Sequences (5'-3') |
|---|---|
18S-F 18S-R | AGTCCCTGCCCTTTGTACACA CGATCCGAGGGCCTCACTA |
| Tmem240-F | TGCTTGATGGACATGAACGC |
| Tmem240-R | TAGTTCTCGGAGGCGTCCAC |
| Bex4_F | GGGGGAAATGTCCGAAGGAAA |
| Bex4_R | CCTGCACTACAAATCTCCCAAC |
| Fdft1_F | AGAGTGGCGGTTCACTGAGA |
| Fdft1_R | GAGAAAGGCCAATTCCCACCA |
| Slc2a3-F | TCATCAATGCACCTGAGACAATC |
| Slc2a3-R | GTCCCTCACTTGGTAGGTCTT |
| Arhgdib_F | CAACACACATACCGGACTGG |
| Arhgdib_R | TGTTTGTCGTCATCCGTGAAG |
| Insig1_F | CTAGTGCTCTTCTCATTTGGCG |
| Insig1_R | AGGGATACAGTAAACCGACAACA |
| Hmgcr_F | AGAGCGAGTGCATTAGCAAAG |
| Hmgcr_R | GATTGCCATTCCACGAGCTAT |
| Hmgcs1_F | AACTGGTGCAGAAATCTCTAGC |
| Hmgcs1_R | GGTTGAATAGCTCAGAACTAGCC |
| SPP1_F | AGCAAGAAACTCTTCCAAGCAA |
| SPP1_R | GTGAGATTCGTCAGATTCATCCG |
| Fam78b_F | TGCGCTACAAGACCCCTTACT |
| Fam78b_R | CTGATGGCTTTTACTCTCCCTTC |
| Arhgdia-F | AAGGACGATGAAAGCCTCCG |
| Arhgdia-R | GGTCAGTCGAGTCACAATGACA |
| Arhgdig-F | GTCAACTCCATCAGATGAGGTG |
| Arhgdig-R | GGGGTCCATAATGGGTGGC |
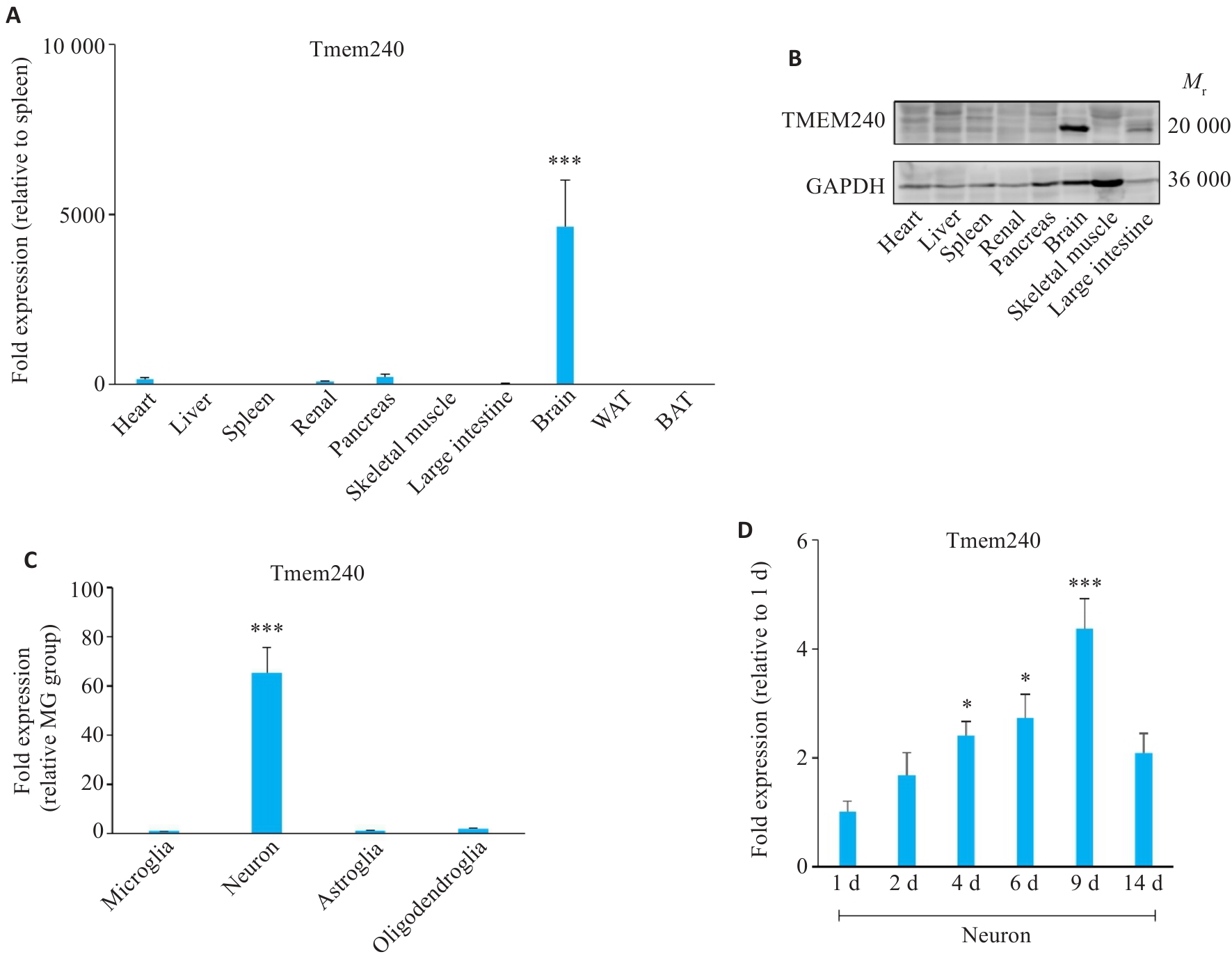
图1 TMEM240在神经元中特异性表达
Fig.1 TMEM240 shows neuron-specific expression. A: qPCR detection of Tmem240 mRNA levels in various tissues of C57BL/6 mice (n=3; ***P<0.001 vs spleen). B: Western blotting for analyzing TMEM240 protein levels in different tissues of C57BL/6 mice (n=3). C: qPCR profiling of Tmem240 mRNA in neurons, astrocytes, oligodendrocytes, and microglia of C57BL/6 mice (n=3; ***P<0.001 vs microglia group). D: qPCR quantification of Tmem240 mRNA in cultured neurons at different time points (n=3; *P<0.05, ***P<0.001 vs 1 day group).
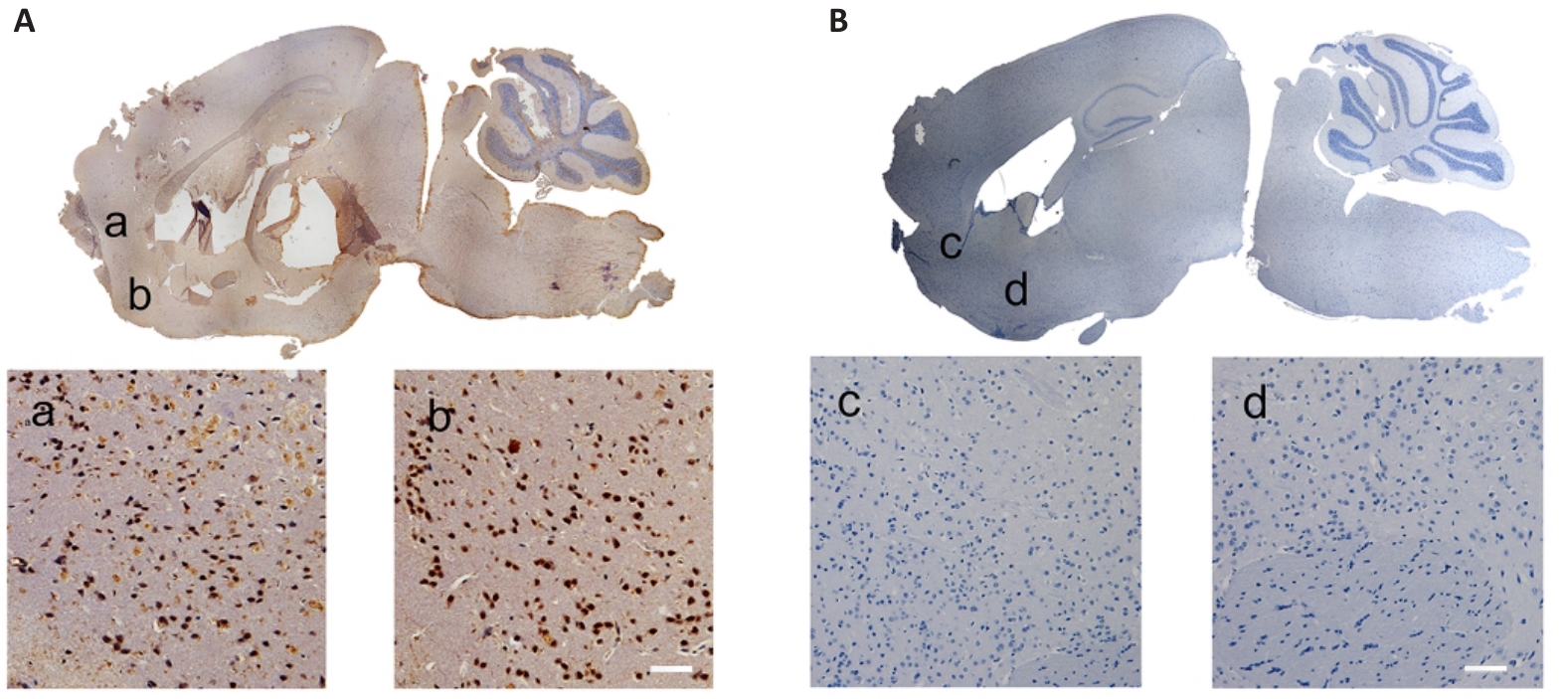
图2 TMEM240 在脑组织的表达分布
Fig.2 Distribution of TMEM240 expression in mouse brain tissues (Scale bar=40 μm). A: In situ hybridization of C57BL/6 mouse brain tissue reveals abundant TMEM240 expression. B: No probe control group. a-d shows magnified views of the corresponding selected areas.
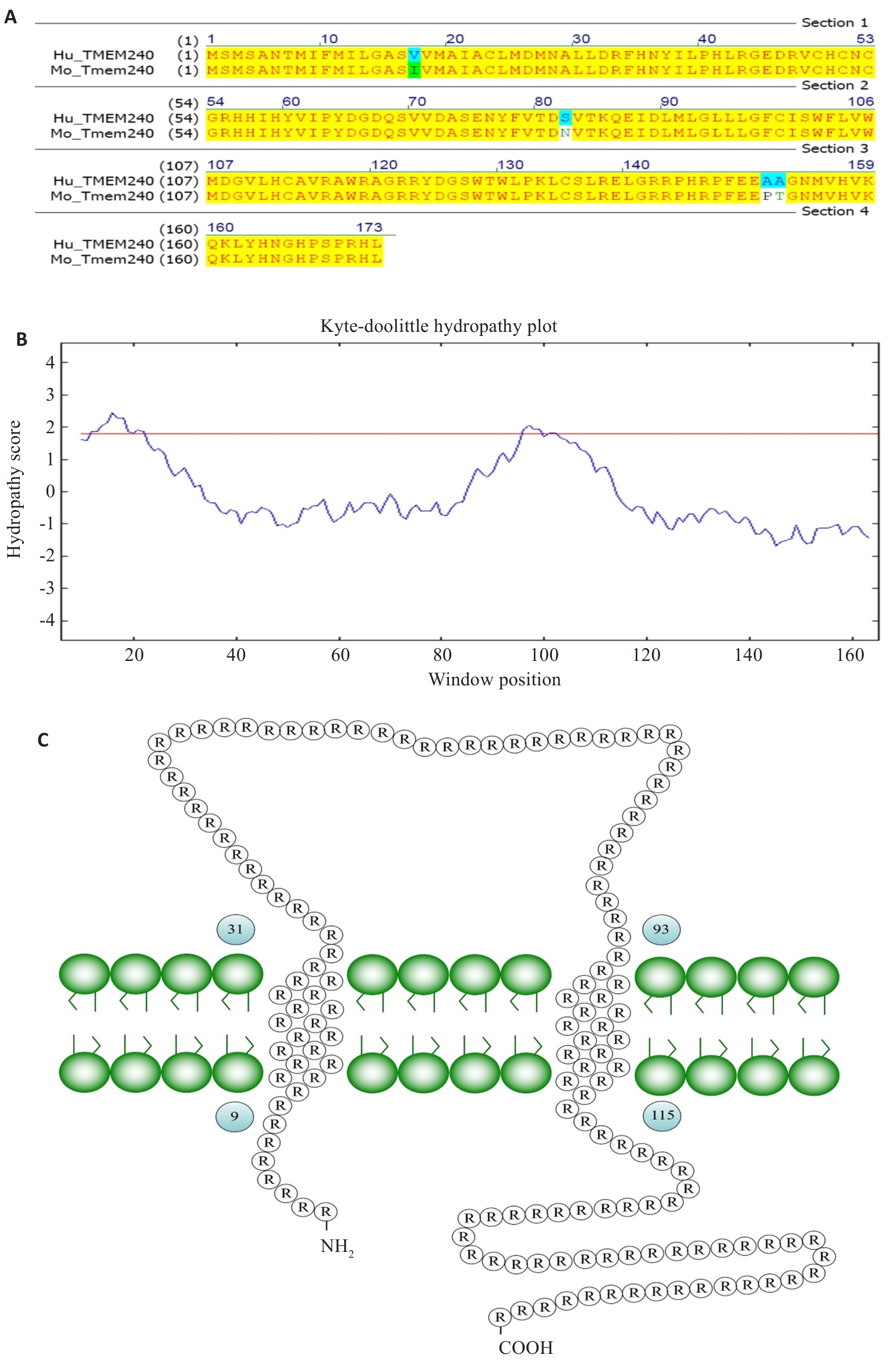
图3 TMEM240的跨膜结构分析
Fig.3 Analysis of transmembrane structure of TMEM240. A: The sequence homology between human and murine TMEM240 proteins is 97.69%. B: Hydrophobicity plot of TMEM240. C: Schematic representation of transmembrane topology of TMEM240.
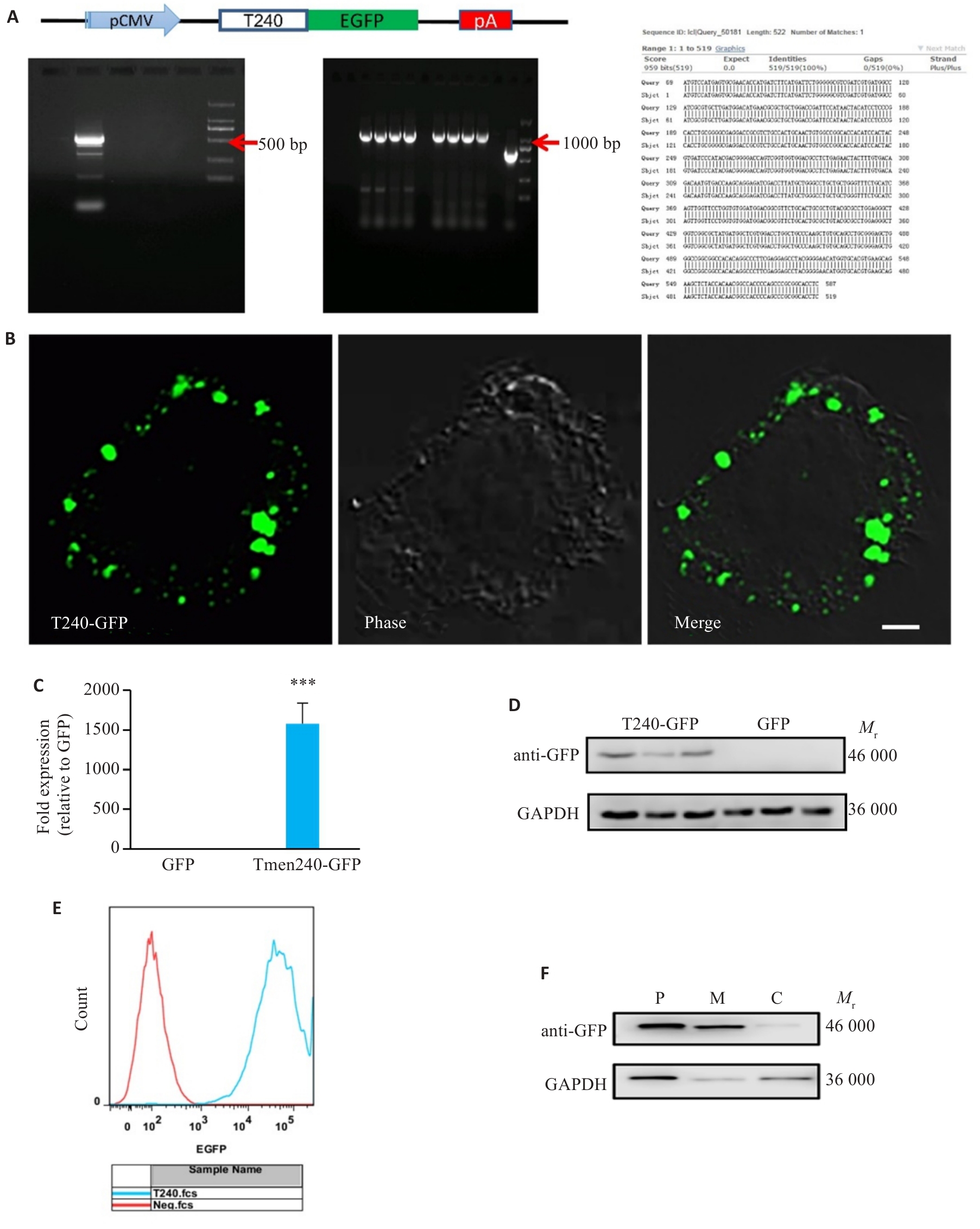
图4 Tmem240在Neuro-2a细胞中过表达体系的构建
Fig.4 Construction of Tmem240-overexpressing Neuro-2a cells. A: Schematic diagram of the Tmem240-pEGFP-N1 plasmid construct. B: Confocal microscopy observation of Neuro 2a cells transfected with pEGFP-Tmem240 eukaryotic expression plasmid showing distribution intracellular Tmem240 expression (Scale bar=2 μm). C: qPCR validation of Tmem240 overexpression in Neuro 2a cells at the mRNA level (n=3). D: Western blotting confirming TMEM240 overexpression in Neuro 2a cells (n=3). E: Flow cytometry detection of Tmem240-pEGFP-N1 positive rate. F: Western blotting of TMEM240 expression in membrane (M), cytoplasmic (C), and total proteins (P) of Neuro 2a cells overexpressing Tmem240. ***P<0.001 vs GFP control.
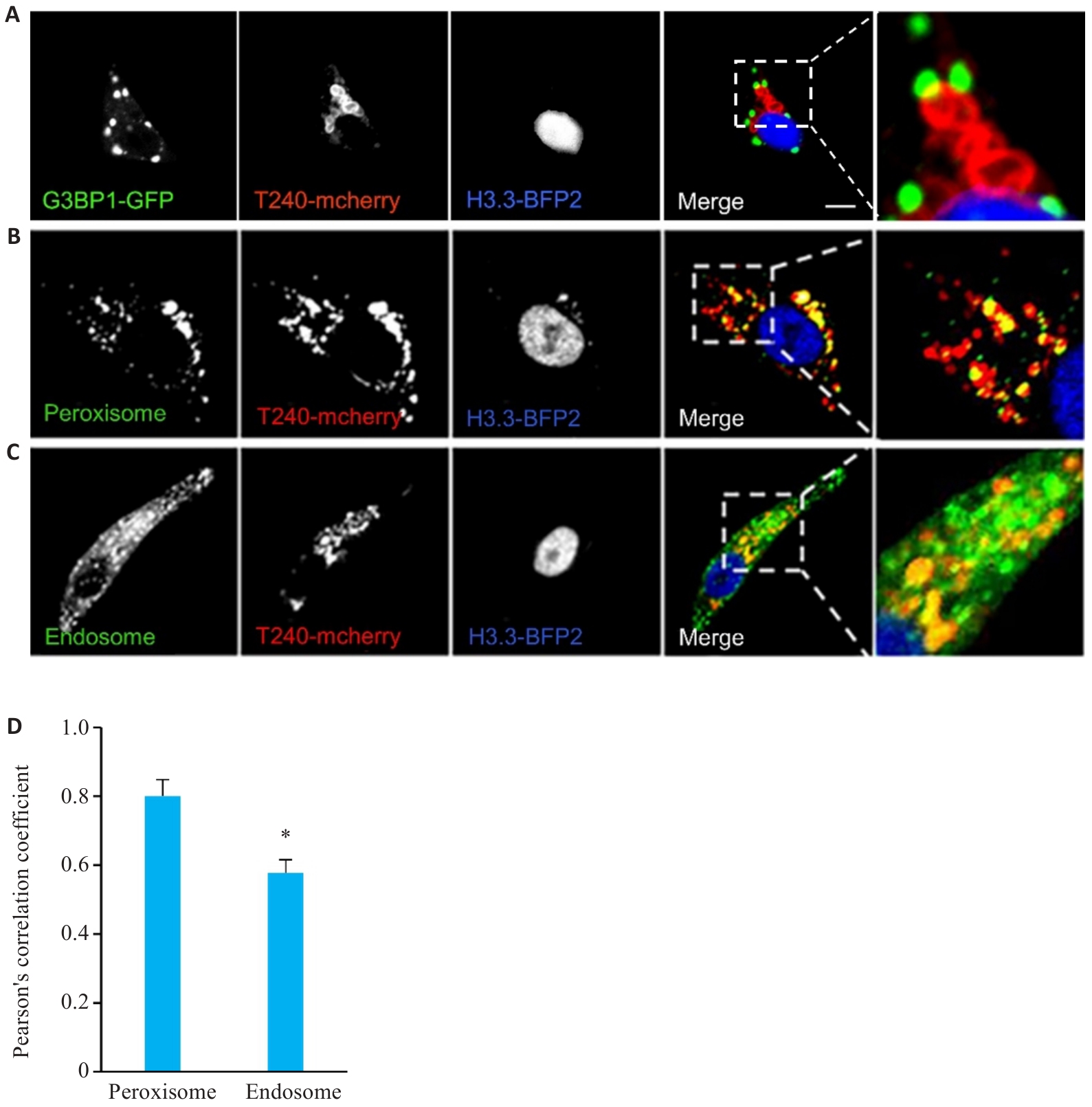
图5 TMEM240在细胞内的分布
Fig.5 Intracellular distribution of TMEM240. A: Confocal microscopy analysis of colocalization between pLVX-mcherry-TMEM240 (T240-mCherry, red) and G3BP1 (green) in HepG2 cells. B, C: Confocal microscopy observation of colocalization of pLVX-mcherry-TMEM240 (red) with peroxisome (PXMP2) and endosome (RAb5b) in HepG2 cells, with the nuclei stained blue by H3.3-BFP2 (Scale bar=10 μm). D: Pearson's correlation coefficient assessment of fluorescence overlap between pLVX-mcherry-TMEM240, peroxisome (PXMP2), and endosome (RAb5b) using Image-Pro software. Data represents mean scores (Mean±SD) from 3 independent experiments comprising a total of 60 cells for each condition. *P<0.05 vs PXMP2 group.
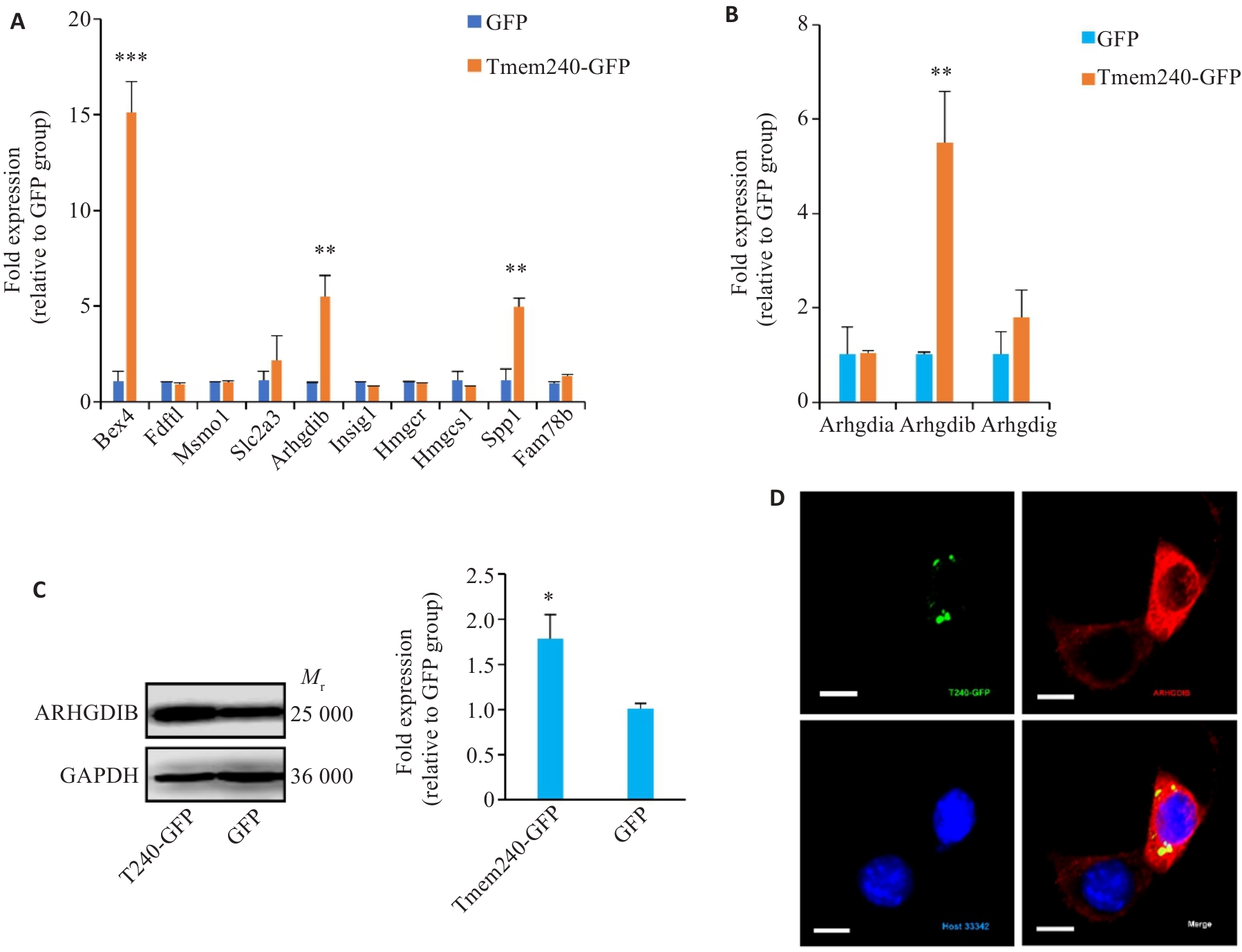
图6 TMEM240的生物学功能研究
Fig.6 Investigation of biological functions of TMEM240. A: qPCR analysis of gene expression levels in Neuro2A cells overexpressing Tmem240 (n=3). B: qPCR validation of Arhgdia, Arhgdib, and Arhgdig mRNA expression levels in Neuro2a cells overexpressing Tmem240 (n=3). C: Western blotting analysis of the impact of Tmem240 overexpression on ARHGDIB protein levels in Neuro2a cells (n=3). D: Immunofluorescence assessment of the effect of Tmem240 overexpression on ARHGDIB expression levels in Neuro2a cells (×800). *P<0.05, **P<0.01, ***P<0.001 vs GFP controll.
| 1 | Kang H, Lee CJ. Transmembrane proteins with unknown function (TMEMs) as ion channels: electrophysiological properties, structure, and pathophysiological roles[J]. Exp Mol Med, 2024, 56(4): 850-60. doi:10.1038/s12276-024-01206-1 |
| 2 | Herrera-Quiterio GA, Encarnación-Guevara S. The transmembrane proteins (TMEM) and their role in cell proliferation, migration, invasion, and epithelial-mesenchymal transition in cancer[J]. Front Oncol, 2023, 13: 1244740. doi:10.3389/fonc.2023.1244740 |
| 3 | Forrest AS, Joyce TC, Huebner ML, et al. Increased TMEM16A-encoded calcium-activated chloride channel activity is associated with pulmonary hypertension[J]. Am J Physiol Cell Physiol, 2012, 303(12): C1229-43. doi:10.1152/ajpcell.00044.2012 |
| 4 | Christopher KJ, Wang BL, Kong Y, et al. Forward genetics uncovers Transmembrane protein 107 as a novel factor required for ciliogenesis and Sonic hedgehog signaling[J]. Dev Biol, 2012, 368(2): 382-92. doi:10.1016/j.ydbio.2012.06.008 |
| 5 | Zhao Y, Hu J, Miao G, et al. Transmembrane protein 208: a novel ER-localized protein that regulates autophagy and ER stress[J]. PLoS One, 2013, 8(5): e64228. doi:10.1371/journal.pone.0064228 |
| 6 | Millay DP, O’Rourke JR, Sutherland LB, et al. Myomaker is a membrane activator of myoblast fusion and muscle formation[J]. Nature, 2013, 499(7458): 301-5. doi:10.1038/nature12343 |
| 7 | Medina MW, Bauzon F, Naidoo D, et al. Transmembrane protein 55B is a novel regulator of cellular cholesterol metabolism[J]. Arterioscler Thromb Vasc Biol, 2014, 34(9): 1917-23. doi:10.1161/ATVBAHA.113.302806 |
| 8 | Zhang Z, Bai M, Barbosa GO, et al. Broadly conserved roles of TMEM131 family proteins in intracellular collagen assembly and secretory cargo trafficking[J]. Sci Adv, 2020, 6(7): eaay7667. doi:10.1126/sciadv.aay7667 |
| 9 | Marx S, Dal Maso T, Chen JW, et al. Transmembrane (TMEM) protein family members: Poorly characterized even if essential for the metastatic process[J]. Semin Cancer Biol, 2020, 60: 96-106. doi:10.1016/j.semcancer.2019.08.018 |
| 10 | Li CY, Ou RW, Chen YP, et al. Mutation analysis of TMEM family members for early-onset Parkinson's disease in Chinese population[J]. Neurobiol Aging, 2021, 101: 299.e1-299.e6. doi:10.1016/j.neurobiolaging.2020.11.005 |
| 11 | Jiao HS, Yuan P, Yu JT. TMEM106B aggregation in neuro-degenerative diseases: linking genetics to function[J]. Mol Neurodegener, 2023, 18(1): 54. doi:10.1186/s13024-023-00644-1 |
| 12 | Li L, Khatibi NH, Hu Q, et al. Transmembrane protein 166 regulates autophagic and apoptotic activities following focal cerebral ischemic injury in rats[J]. Exp Neurol, 2012, 234(1): 181-90. doi:10.1016/j.expneurol.2011.12.038 |
| 13 | Seki T, Sato M, Kibe Y, et al. Lysosomal dysfunction and early glial activation are involved in the pathogenesis of spinocerebellar Ataxia type 21 caused by mutant transmembrane protein 240[J]. Neurobiol Dis, 2018, 120: 34-50. doi:10.1016/j.nbd.2018.08.022 |
| 14 | Delplanque J, Devos D, Huin V, et al. TMEM240 mutations cause spinocerebellar Ataxia 21 with mental retardation and severe cognitive impairment[J]. Brain, 2014, 137(10): 2657-63. doi:10.1093/brain/awu202 |
| 15 | Chang SC, Liew PL, Ansar M, et al. Hypermethylation and decreased expression of TMEM240 are potential early-onset biomarkers for colorectal cancer detection, poor prognosis, and early recurrence prediction[J]. Clin Epigenetics, 2020, 12(1): 67. doi:10.1186/s13148-020-00855-z |
| 16 | Lin RK, Su CM, Lin SY, et al. Hypermethylation of TMEM240 predicts poor hormone therapy response and disease progression in breast cancer[J]. Mol Med, 2022, 28(1): 67. doi:10.1186/s10020-022-00474-9 |
| 17 | Bian S, Wang Y, Zhou Y, et al. Integrative single-cell multiomics analyses dissect molecular signatures of intratumoral hetero-geneities and differentiation states of human gastric cancer[J]. Natl Sci Rev, 2023, 10(6): nwad094. doi:10.1093/nsr/nwad094 |
| 18 | Ward JM, La Spada AR. Identification of the SCA21 disease gene: remaining challenges and promising opportunities[J]. Brain, 2014, 137(10): 2626-8. doi:10.1093/brain/awu217 |
| 19 | Yahikozawa H, Miyatake S, Sakai T, et al. A Japanese family of spinocerebellar Ataxia type 21: clinical and neuropathological studies[J]. Cerebellum, 2018, 17(5): 525-30. doi:10.1007/s12311-018-0941-6 |
| 20 | Zeng S, Zeng J, He M, et al. Spinocerebellar Ataxia type 21 exists in the Chinese Han population[J]. Sci Rep, 2016, 6: 19897. doi:10.1038/srep19897 |
| 21 | D’Ambrosi N, Milani M, Apolloni S. S100A4 in the physiology and pathology of the central and peripheral nervous system[J]. Cells, 2021, 10(4): 798. doi:10.3390/cells10040798 |
| 22 | Brennan FH, Coulthard LG, Alawieh AM, et al. Editorial: complement in the development and regeneration of the nervous system[J]. Front Immunol, 2021, 12: 694810. doi:10.3389/fimmu.2021.694810 |
| 23 | Tikhonova MA, Shvaikovskaya AA, Zhanaeva SY, et al. Concordance between the in vivo content of neurospecific proteins (BDNF, NSE, VILIP-1, S100B) in the hippocampus and blood in patients with epilepsy[J]. Int J Mol Sci, 2023, 25(1): 502. doi:10.3390/ijms25010502 |
| 24 | Tikhonova MA, Zhanaeva SY, Shvaikovskaya AA, et al. Neurospecific molecules measured in periphery in humans: how do they correlate with the brain levels? A systematic review[J]. Int J Mol Sci, 2022, 23(16): 9193. doi:10.3390/ijms23169193 |
| 25 | Schröder B, Wrocklage C, Hasilik A, et al. Molecular charac-terisation of 'transmembrane protein 192' (TMEM192), a novel protein of the lysosomal membrane[J]. Biol Chem, 2010, 391(6): 695-704. doi:10.1515/bc.2010.062 |
| 26 | Hanein S, Perrault I, Roche O, et al. TMEM126A, encoding a mitochondrial protein, is mutated in autosomal-recessive nonsyn-dromic optic atrophy[J]. Am J Hum Genet, 2009, 84(4): 493-8. doi:10.1016/j.ajhg.2009.03.003 |
| 27 | Wesoly J, Pstrąg N, Derylo K, et al. Structural, topological, and functional characterization of transmembrane proteins TMEM213, 207, 116, 72 and 30B provides a potential link to ccRCC etiology[J]. Am J Cancer Res, 2023, 13(5): 1863-83. doi:10.62347/cred7532 |
| 28 | Homa M, Loyens A, Eddarkaoui S, et al. The TMEM240 protein, mutated in SCA21, is expressed in Purkinje cells and synaptic terminals[J]. Cerebellum, 2020, 19(3): 358-69. doi:10.1007/s12311-020-01112-y |
| 29 | Parker DM, Tauber D, Parker R. G3BP1 promotes intermolecular RNA-RNA interactions during RNA condensation[J]. Mol Cell, 2025, 85(3): 571-84. e7. doi:10.1016/j.molcel.2024.11.012 |
| 30 | Protter DSW, Parker R. Principles and properties of stress granules[J]. Trends Cell Biol, 2016, 26(9): 668-79. doi:10.1016/j.tcb.2016.05.004 |
| 31 | Schmit K, Michiels C. TMEM proteins in cancer: a review[J]. Front Pharmacol, 2018, 9: 1345. doi:10.3389/fphar.2018.01345 |
| 32 | 宋 玲, 张 洪, 范月莹, 等. 跨膜蛋白家族与癌症的发生发展[J]. 生命的化学, 2021, 41(10): 2168-76. |
| 33 | Liu L, Cui J, Zhao Y, et al. KDM6A-ARHGDIB axis blocks metastasis of bladder cancer by inhibiting Rac1[J]. Mol Cancer, 2021, 20(1): 77. doi:10.1186/s12943-021-01369-9 |
| [1] | 杨毓甲, 杨丽芳, 吴雅玲, 段兆达, 于春泽, 吴春云, 于建云, 杨力. 大麻二酚经PERK-eIF2α-ATF4-CHOP通路减轻多重脑震荡大鼠的神经元内质网应激和凋亡[J]. 南方医科大学学报, 2025, 45(6): 1240-1250. |
| [2] | 何光侣, 储婉玉, 李妍, 盛鑫, 罗浩, 徐爱萍, 卞命杰, 张环环, 汪萌芽, 郑超. Orexin-A通过调节促离子型谷氨酸受体促进脊髓损伤大鼠运动功能恢复[J]. 南方医科大学学报, 2025, 45(5): 1023-1030. |
| [3] | 陈昌锋, 方勤, 高银环, 王烈成, 陈磊. 丰富环境通过调控前扣带回皮层锥体神经元兴奋性缓解束缚应激诱导的小鼠焦虑样行为[J]. 南方医科大学学报, 2025, 45(5): 962-968. |
| [4] | 陈悦, 肖林雨, 任侣, 宋雪, 李静, 胡建国. 水晶兰苷通过抑制PI3K/AKT信号通路减少神经元凋亡改善脊髓损伤后小鼠的运动功能[J]. 南方医科大学学报, 2025, 45(4): 774-784. |
| [5] | 蔡巧燕, 许瑶瑶, 林雨星, 林浩伟, 郑俊鹏, 张伟祥, 赵春雨, 林育鹏, 张铃. 清达颗粒通过调控miR-124/STAT3信号轴减轻自发性高血压大鼠脑损伤[J]. 南方医科大学学报, 2025, 45(1): 18-26. |
| [6] | 李言响, 郭永馨, 曹福羊, 郭舒婷, 薛丁豪, 周志康, 郝新宇, 仝 黎, 傅 强. 抑制dmPAG区谷氨酸能神经元可减轻创伤后应激障碍小鼠的过度防御反应[J]. 南方医科大学学报, 2024, 44(3): 420-427. |
| [7] | 桂建军, 孙晓东, 温 舒, 刘 欣, 覃冰清, 桑 明. 白藜芦醇对帕金森病模型小鼠多巴胺能神经元的保护作用:基于抑制TLR4/MyD88/NF-κB通路[J]. 南方医科大学学报, 2024, 44(2): 270-279. |
| [8] | 王 娟, 杨雯钦, 刘 进, 石金凤, 肖 萍, 李美香. 脂联素通过上调PPARα/HOXA10通路改善多囊卵巢综合征大鼠的子宫内膜容受性[J]. 南方医科大学学报, 2024, 44(2): 298-307. |
| [9] | 傅鸣宇, 纪晓宏, 钟 磊, 吴 琼, 李海福, 王宁黔. 大鼠小脑浦肯野神经元发育过程中NaV亚基表达与电生理特性的发育成熟具有相关性[J]. 南方医科大学学报, 2023, 43(7): 1102-1109. |
| [10] | 殷 果, 李 荣, 刘岳飞, 王晓睛, 吴炳义. Notch信号通路抑制剂DAPT改善酒精诱导的斑马鱼神经元分化障碍[J]. 南方医科大学学报, 2023, 43(6): 889-899. |
| [11] | 曹福羊, 郭永馨, 郭舒婷, 周志康, 曹江北, 仝 黎, 米卫东. 激活小鼠ZI 区GABA 能神经元可促进七氟醚和丙泊酚的麻醉诱导而对麻醉维持及觉醒无影响[J]. 南方医科大学学报, 2023, 43(5): 718-726. |
| [12] | 徐佳佳, 李洋洋, 仲光尚, 方祝玲, 刘淳博, 马彩云, 王春景, 郭 俣, 刘长青. 人诱导多能干细胞体外定向分化为多巴胺能神经元祖细胞[J]. 南方医科大学学报, 2023, 43(2): 175-182. |
| [13] | 张 列, 苗树船, 杨中鑫, 李宗喜, 范英俊, 余 凯, 黄可阳, 黄庆喜, 夏 勋. HMGB1下调的作用:通过抑制神经元细胞自噬和凋亡减轻大鼠脑出血后的神经元损伤[J]. 南方医科大学学报, 2022, 42(7): 1050-1056. |
| [14] | 黄威龙, 梁妃学. 运动调控纹状体听觉响应的神经机制[J]. 南方医科大学学报, 2022, 42(5): 766-771. |
| [15] | 游佳鹏, 宋长宝, 梁妃学. 小鼠纹状体尾部和听皮层PV神经元的电生理性质比较[J]. 南方医科大学学报, 2022, 42(12): 1889-1895. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||