南方医科大学学报 ›› 2026, Vol. 46 ›› Issue (4): 742-752.doi: 10.12122/j.issn.1673-4254.2026.04.03
• • 上一篇
吴博1( ), 宋茹1, 高宁1, 邢可尧1, 张鹏辉1, 屈沫怡1, 张慧敏2
), 宋茹1, 高宁1, 邢可尧1, 张鹏辉1, 屈沫怡1, 张慧敏2
收稿日期:2025-11-18
出版日期:2026-04-20
发布日期:2026-04-24
通讯作者:
吴博
E-mail:bwu@yau.edu.cn
基金资助:
Bo WU1( ), Ru SONG1, Ning GAO1, Keyao XING1, Penghui ZHANG1, Moyi QU1, Huimin ZHANG2
), Ru SONG1, Ning GAO1, Keyao XING1, Penghui ZHANG1, Moyi QU1, Huimin ZHANG2
Received:2025-11-18
Online:2026-04-20
Published:2026-04-24
Contact:
Bo WU
E-mail:bwu@yau.edu.cn
Supported by:摘要:
目的 探讨长链非编码RNA LASTR在头颈鳞癌细胞中的调控作用、分子机制及临床意义。 方法 利用TCGA、GEO数据库转录组测序数据分析LASTR在头颈鳞癌中的表达水平及其与患者生存时间的相关性,qPCR检测LASTR在头颈鳞癌细胞和正常对照细胞中的表达水平。利用siRNA建立LASTR敲降的头颈鳞癌细胞模型,通过高内涵细胞计数、CCK-8实验、ATP检测分析增殖能力,Transwell实验检测细胞迁移和侵袭能力。生物信息学分析LASTR候选靶基因并进行功能注释,qPCR和免疫印迹验证LASTR对候选靶基因BCAM的调控作用。RNA pulldown和双荧光素酶报告分析研究LASTR对BCAM的调控机制。利用TCGA和GEO转录组数据分析BCAM在头颈鳞癌中的表达水平,细胞功能实验研究BCAM对头颈鳞癌细胞增殖、迁移和侵袭能力的影响。挽救实验探究LASTR是否通过调控BCAM表达水平影响头颈鳞癌细胞增殖、迁移和侵袭能力。 结果 与正常对照相比,头颈鳞癌组织和细胞中LASTR表达水平升高(P<0.01)。LASTR高表达与头颈鳞癌预后不良显著相关(P<0.05)。敲降LASTR抑制头颈鳞癌细胞增殖、迁移和侵袭能力(P<0.01);生物信息学分析发现78个LASTR候选靶基因,这些基因主要富集于血管生成、低氧应答、血管内皮生长因子受体信号通路、细胞群增殖的正向调节、MAPK信号通路、ErbB信号通路、Ras信号通路等生物过程和信号通路;敲降LASTR导致头颈鳞癌细胞BCAM表达水平降低(P<0.01);机制研究揭示LASTR通过特异性结合miR-4476上调靶基因BCAM表达水平。敲降BCAM抑制头颈鳞癌细胞增殖、迁移和侵袭能力,而过表达BCAM能够逆转敲降LASTR对头颈鳞癌细胞增殖、迁移和侵袭能力的抑制作用。 结论 LASTR在头颈鳞癌组织和细胞中表达水平上调,且LASTR高表达与头颈鳞癌预后不良相关。LASTR通过特异性结合miR-4476上调BCAM表达水平,从而促进头颈鳞癌细胞增殖、迁移和侵袭能力。本研究结果将为头颈鳞癌临床诊疗提供新的潜在标志物。
吴博, 宋茹, 高宁, 邢可尧, 张鹏辉, 屈沫怡, 张慧敏. 长链非编码RNA LASTR通过结合miR-4476上调BCAM表达促进头颈鳞癌恶性进展[J]. 南方医科大学学报, 2026, 46(4): 742-752.
Bo WU, Ru SONG, Ning GAO, Keyao XING, Penghui ZHANG, Moyi QU, Huimin ZHANG. Long non-coding RNA LASTR promotes progression of head and neck squamous cell carcinoma by binding to miR-4476 and upregulating BCAM expression[J]. Journal of Southern Medical University, 2026, 46(4): 742-752.
| Name | Sense (5'-3') | Antisense (5'-3') |
|---|---|---|
| si-LASTR-1 | CCUAGAGUCCUAGAGAGCUAGAGAU | AUCUCUAGCUCUCUAGGACUCUAGG |
| si-LASTR-2 | AACCAUUCUGCAGUAAGAUUGUCAU | AUGACAAUCUUACUGCAGAAUGGUU |
| si-LASTR-3 | GGGCUGUAAUUGAUGUGGUUCAGAG | CUCUGAACCACAUCAAUUACAGCCC |
| si-BCAM-1 | CCAACAAAGGGACACUGUCUGUGAU | AUCACAGACAGUGUCCCUUUGUUGG |
| si-BCAM-2 | GAGCCGCGAUGGCAUCUCCUGUGAA | UUCACAGGAGAUGCCAUCGCGGCUC |
| si-NC | CAGUUUCGACUCAUACCGAGUGUUU | AAACACUCGGUAUGAGUCGAAACUG |
表1 siRNA序列信息
Tab.1 Sequences for siRNA targeting LASTR and BCAM
| Name | Sense (5'-3') | Antisense (5'-3') |
|---|---|---|
| si-LASTR-1 | CCUAGAGUCCUAGAGAGCUAGAGAU | AUCUCUAGCUCUCUAGGACUCUAGG |
| si-LASTR-2 | AACCAUUCUGCAGUAAGAUUGUCAU | AUGACAAUCUUACUGCAGAAUGGUU |
| si-LASTR-3 | GGGCUGUAAUUGAUGUGGUUCAGAG | CUCUGAACCACAUCAAUUACAGCCC |
| si-BCAM-1 | CCAACAAAGGGACACUGUCUGUGAU | AUCACAGACAGUGUCCCUUUGUUGG |
| si-BCAM-2 | GAGCCGCGAUGGCAUCUCCUGUGAA | UUCACAGGAGAUGCCAUCGCGGCUC |
| si-NC | CAGUUUCGACUCAUACCGAGUGUUU | AAACACUCGGUAUGAGUCGAAACUG |
| Name | Forward (5'-3') | Reverse (5'-3') |
|---|---|---|
| LASTR | CTCCTCTGCTCTCAGTGAACG | CTGCAGACCACCATAAGGGA |
| BCAM | AAGGGGGCTCCGCCGCCAGG | AGGTTCTTGGCTCAGCACTC |
| GAPDH | CTGGGCTACACTGAGCACC | AAGTGGTCGTTGAGGGCAATG |
表2 qPCR引物序列信息
Tab.2 qPCR primer sequence information
| Name | Forward (5'-3') | Reverse (5'-3') |
|---|---|---|
| LASTR | CTCCTCTGCTCTCAGTGAACG | CTGCAGACCACCATAAGGGA |
| BCAM | AAGGGGGCTCCGCCGCCAGG | AGGTTCTTGGCTCAGCACTC |
| GAPDH | CTGGGCTACACTGAGCACC | AAGTGGTCGTTGAGGGCAATG |
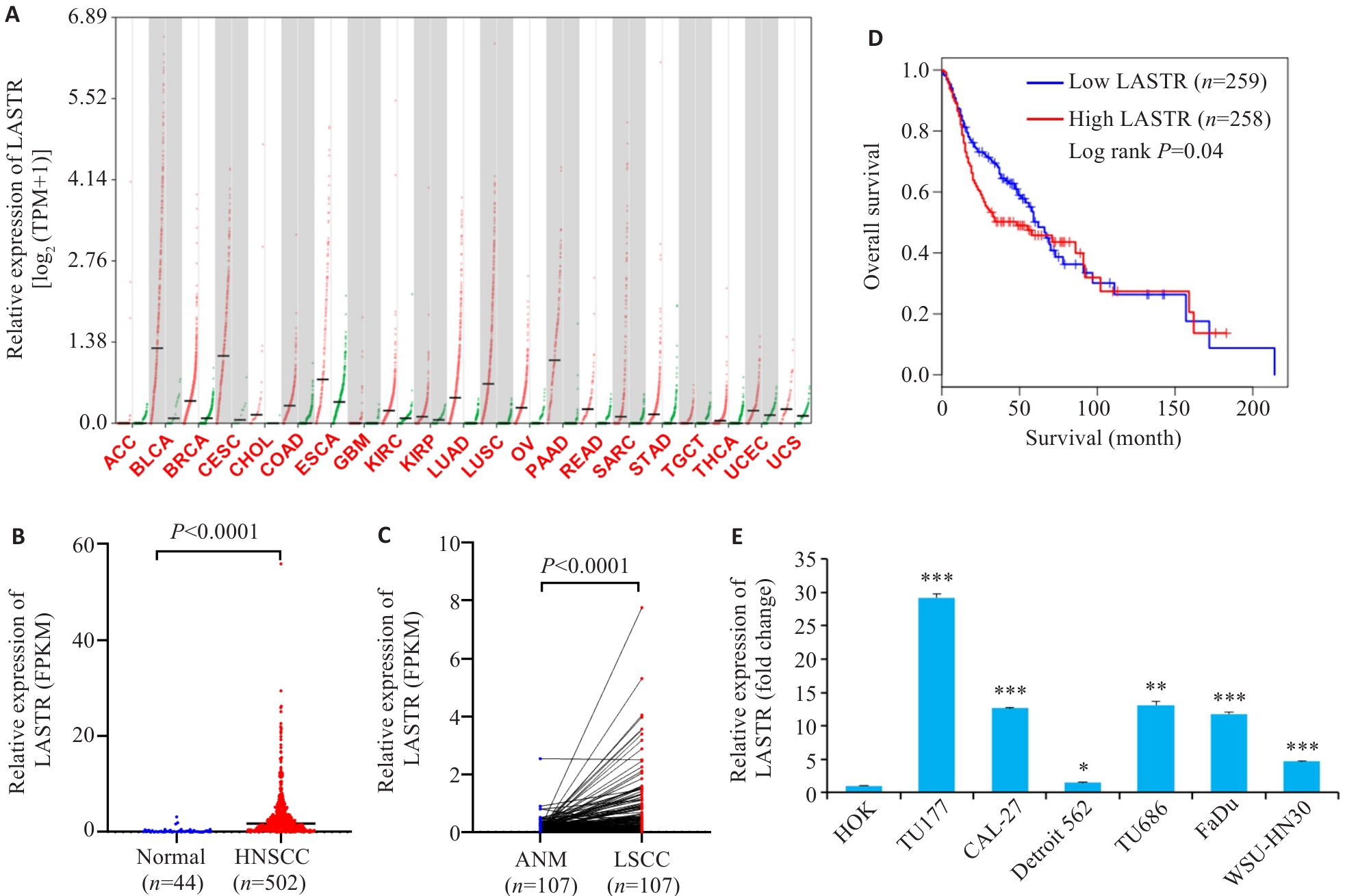
图1 头颈鳞癌组织和细胞中LASTR表达水平及其与患者生存相关性分析
Fig.1 Expression of LASTR in HNSCC tissues and cells and its correlation with survival time of HNSCC patients. A: Expression of LASTR in pan-cancer analyzed by GEPIA. B: Expression of LASTR in HNSCC tissues and normal control tissues analyzed using GEPIA. C: Expression of LASTR in laryngeal squamous cell carcinoma (LSCC) tissues and adjacent normal mucosal (ANM) tissues analyzed using transcriptome data from GEO database. D: Correlation between LASTR expression level and overall survival time of HNSCC patients analyzed using Kaplan-Meir method. E: Expression of LASTR in HNSCC cells and normal control cells determined by qPCR (Mean±SD, n=3). *P<0.05, **P<0.01, ***P<0.001 vs HOK cells.
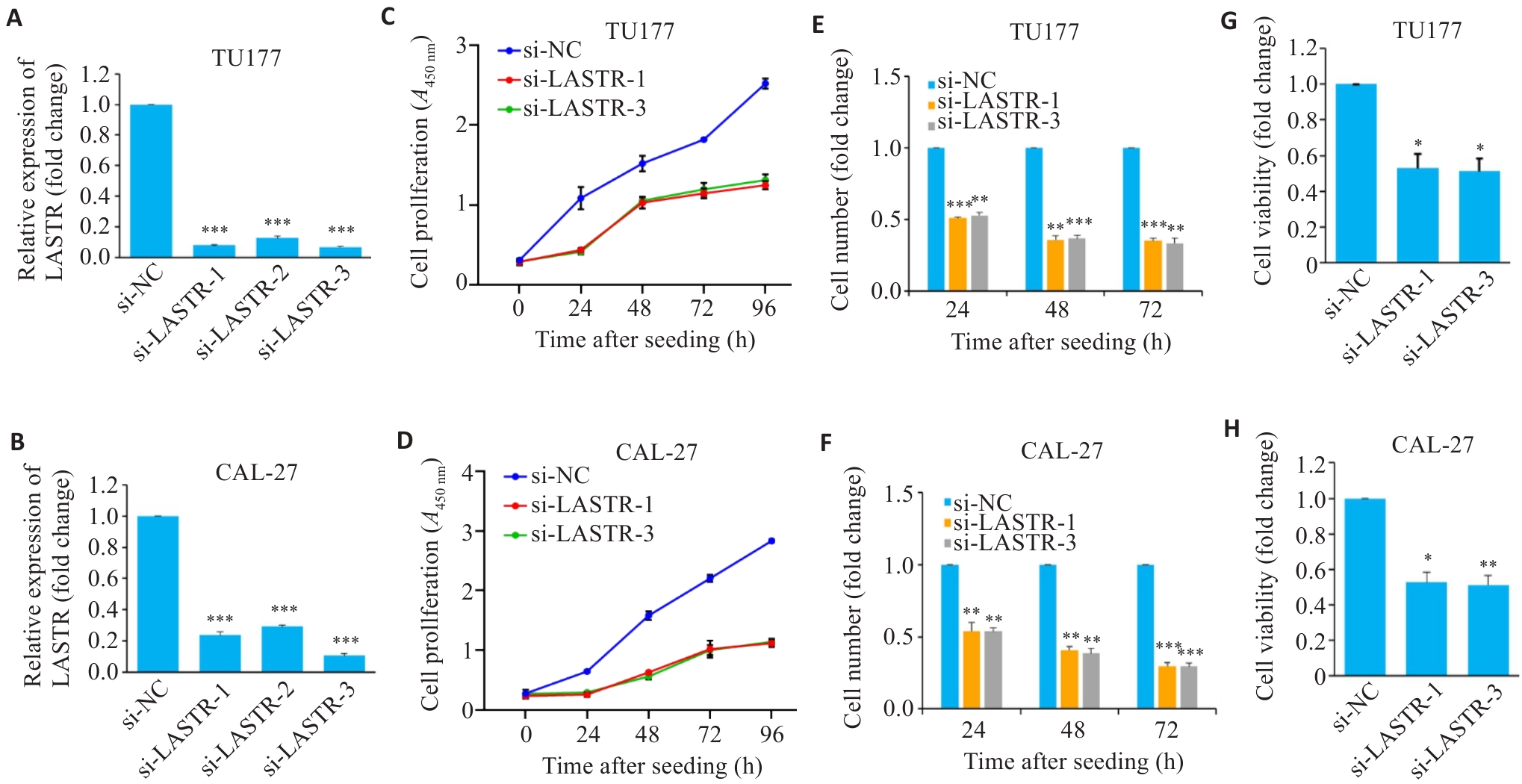
图2 敲降LASTR抑制头颈鳞癌细胞增殖和活力
Fig.2 Knockdown of LASTR inhibits proliferation and viability of HNSCC cells. A, B: TU177 and CAL-27 cells transfected with siRNAs targeting LASTR (si-LASTR) or negative control (si-NC). After 48 h of transfection, LASTR knockdown efficiency was determined by qPCR (n=3). C, D: Proliferation of TU177 and CAL-27 cells with LASTR knockdown determined by CCK8 assay (n=3). E, F: TU177 and CAL-27 cell counts under a high-content system at 24, 48 and 72 h after LASTR knockdown and seeding in 96-well plate (n=3). G, H: TU177 and CAL-27 cell viability after LASTR knockdown determined by CellTiter-Glo® Luminescent Cell Viability Assay. Data are presented as Mean±SD. *P<0.05, **P<0.01, ***P<0.001 vs si-NC group.
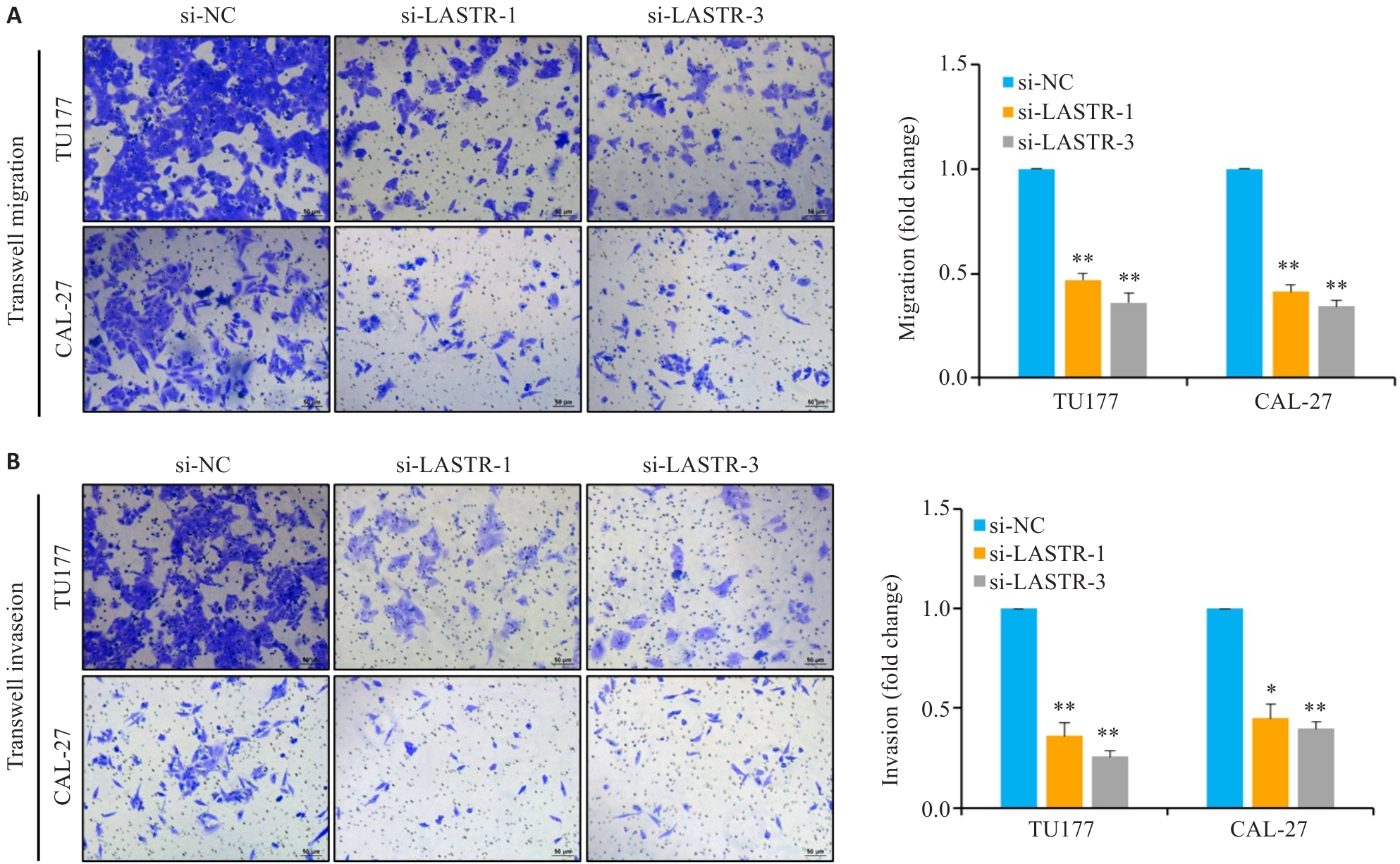
图3 敲降LASTR抑制头颈鳞癌细胞迁移和侵袭能力
Fig.3 Knockdown of LASTR inhibits migration and invasion of HNSCC cells. A, B: TU177 and CAL-27 cells transfected with siRNAs targeting LASTR (si-LASTR) or negative control (si-NC). After 24 h of transfection, the cells were seeded into Transwell insert for 48 h and examined for migration (A) and invasion (B) capacity using Transwell assay (Scale bar=50 μm). Data are presented as Mean±SD (n=3). *P<0.05, **P<0.01 vs si-NC group.
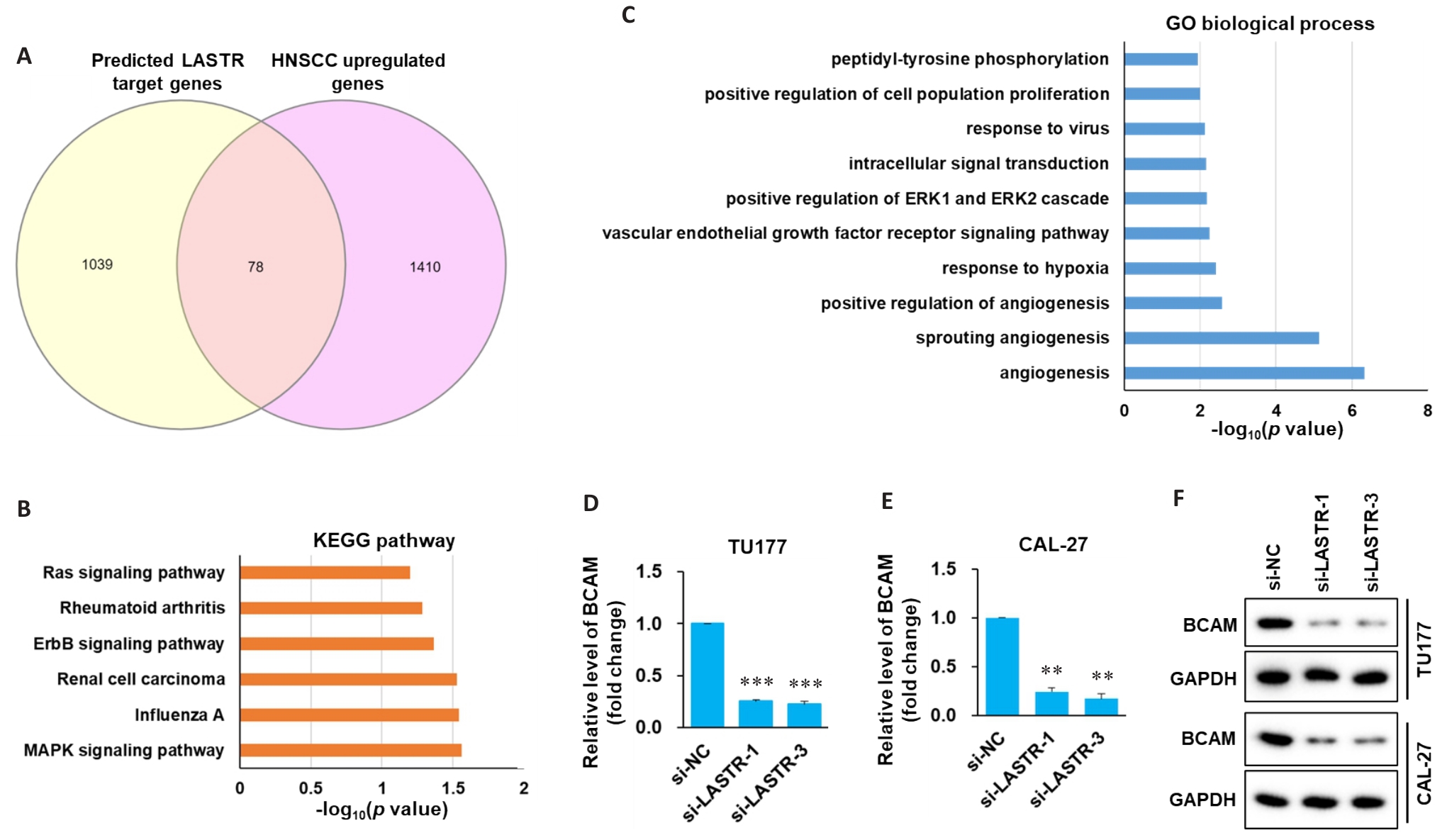
图4 头颈鳞癌细胞LASTR靶基因预测及验证
Fig.4 Prediction and validation of the target genes of LASTR in HNSCC cells. A: Venn diagram of predicted LASTR target genes and genes that upregulated in HNSCC tissues. B: KEGG pathway analysis of LASTR target genes. C: GO analysis of biological process involving LASTR target genes. D, E: TU177 and CAL-27 cells transfected with siRNAs targeting LASTR (si-LASTR) and negative control (si-NC) respectively. After 48 hours of transfection, the expression of BCAM mRNA was determined by qPCR (n=3). F: Western blotting for detecting expression of BCAM in HNSCC cells with LASTR knockdown. Data are presented as Mean±SD. **P<0.01, ***P<0.001 vs si-NC.
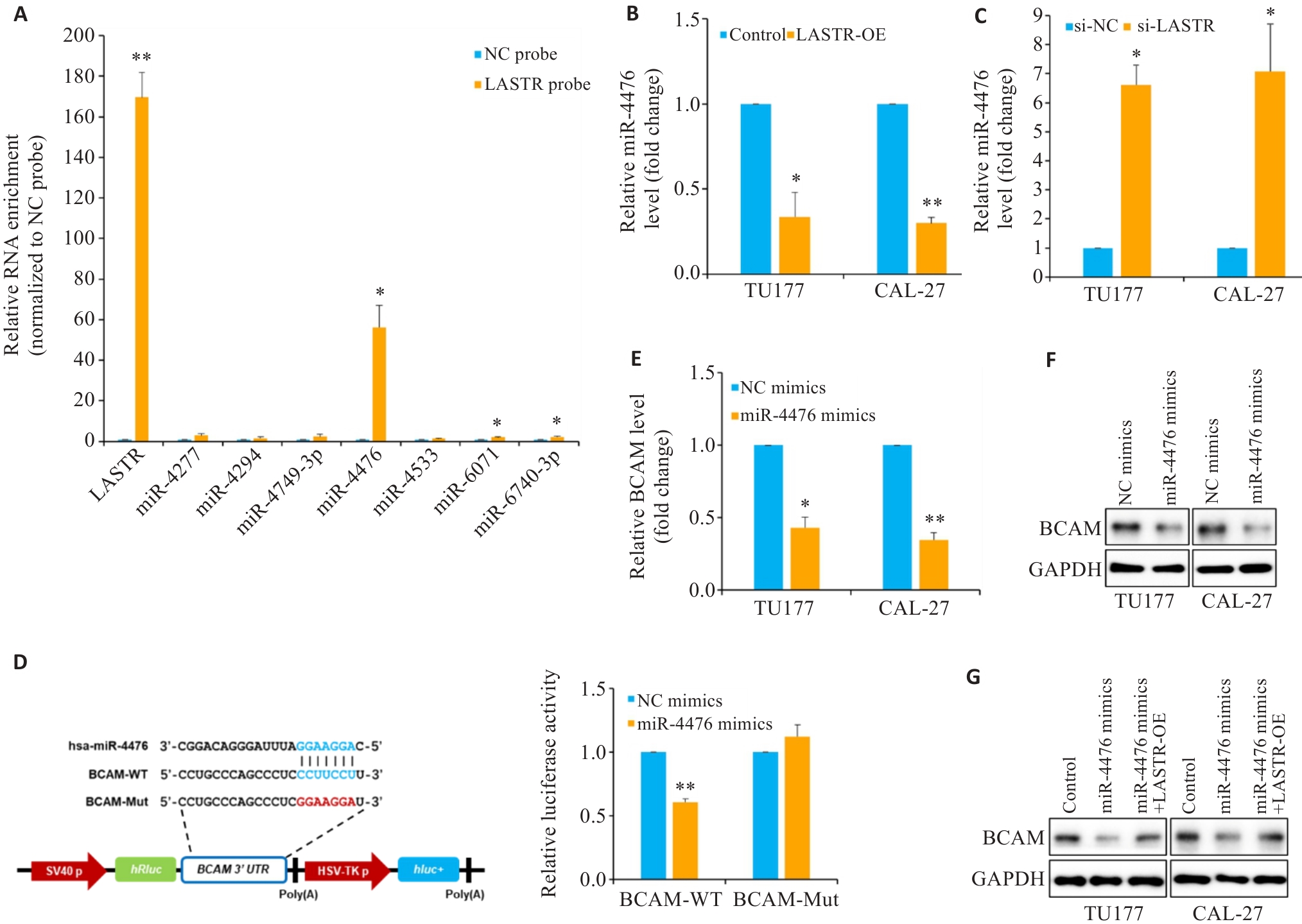
图5 LASTR调控靶基因BCAM表达的机制研究
Fig. 5 Mechanistic study of regulation of BCAM expression by LASTR. A: RNA pulldown assay using Biotin-labeled LASTR probes in TU177 cells and qPCR analysis of enrichment of the indicated miRNAs. B, C: Expression of miR-4476 in TU177 and CAL-27 cells with LASTR overexpression or knockdown determined by qPCR analysis. D: Luciferase reporter assay of TU177 cells cotransfected with miR-4476 mimics and luciferase reporter plasmid containing wild-type (WT) and miR-4476 binding site mutated (Mut) BCAM 3' UTR. E: Expression of BCAM in TU177 and CAL-27 cells transfected with miR-4476 mimics or NC mimics detected by qPCR. F: Expression of BCAM protein in TU177 and CAL-27 cells transfected with miR-4476 mimics or NC mimics detected by Western blotting. G: Expression of BCAM protein in TU177 and CAL-27 cells transfected with miR-4476 mimics or co-transfected with miR-4476 mimics and LASTR overexpressing plasmid (LASTR-OE) detected by Western blotting. Error bars represent SD of 3 independent experiments. *P<0.05, **P<0.01 vs NC/control/si-NC/NC mimics group.
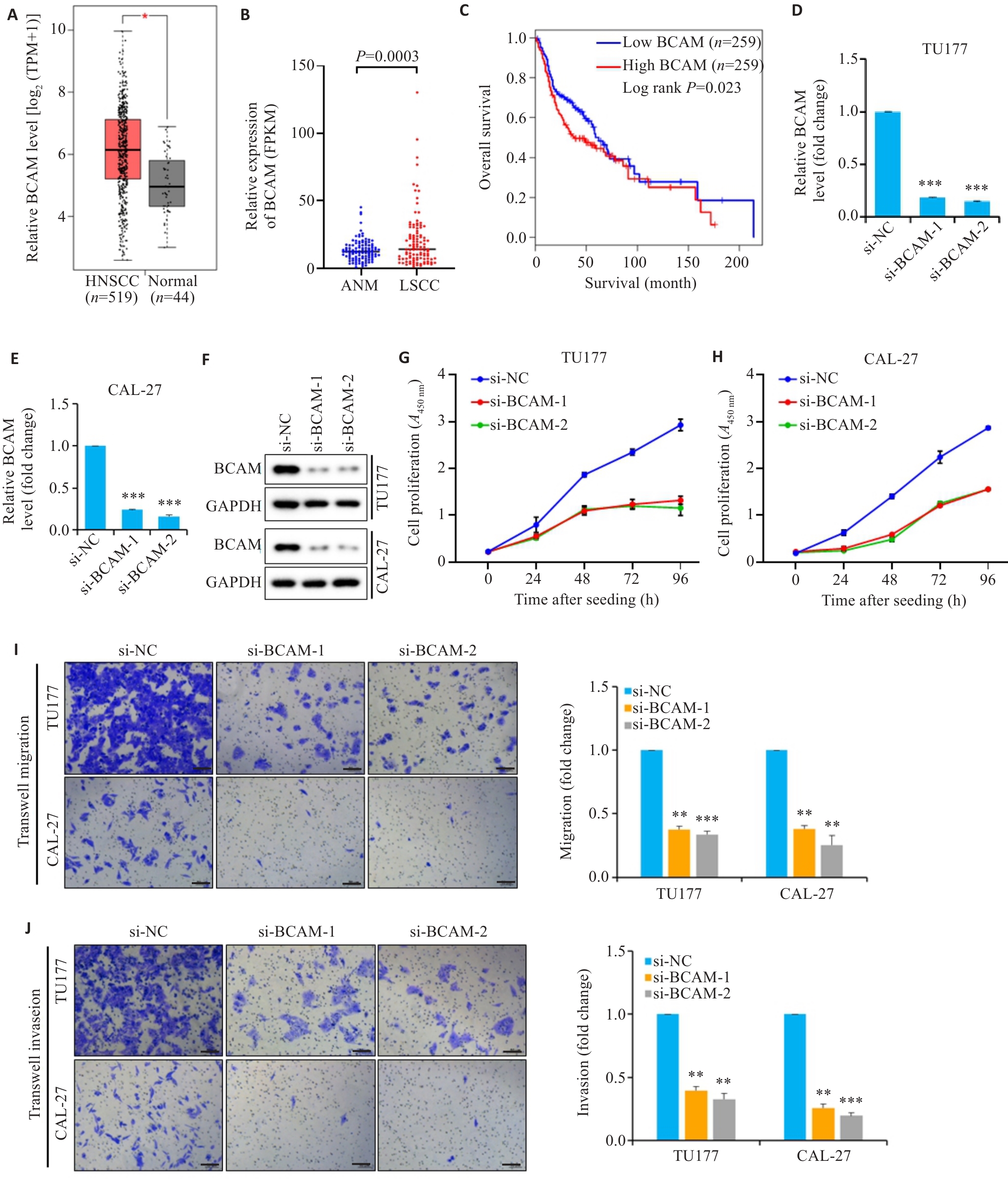
图6 BCAM在头颈鳞癌中的表达水平及生物学功能
Fig.6 Expression of BCAM and its biological functions in HNSCC. A: Expression of BCAM mRNA in HNSCC tissues and normal control tissues determined by GEPIA analysis. B: Expression of BCAM in laryngeal squamous cell carcinoma (LSCC) tissues and adjacent normal mucosal (ANM) tissues analyzed using transcriptome data from GEO database. C: Correlation between BCAM expression level and overall survival time of HNSCC patient analyzed using by Kaplan-Meir method. D, E: BCAM mRNA expression detected by qPCR in TU177 and CAL-27 cells transfected with siRNAs targeting BCAM (si-BCAM) or negative control (n=3). F: Western blotting for detecting BCAM expression in si-BCAM transfected HNSCC cells. G, H: CCK-8 assay of assessing proliferation of TU177 and CAL-27 cells 24 h after transfection with si-BCAM or si-NC (n=3). I: Transwell assay of migration capacity of HNSCC cells with BCAM knockdown (n=3). J: Invasion capacity of HNSCC cells with BCAM knockdown determined by Transwell assays. Scale bar=50 μm. Data are presented as Mean±SD. *P<0.05, **P<0.01, ***P<0.001 vs si-NC.
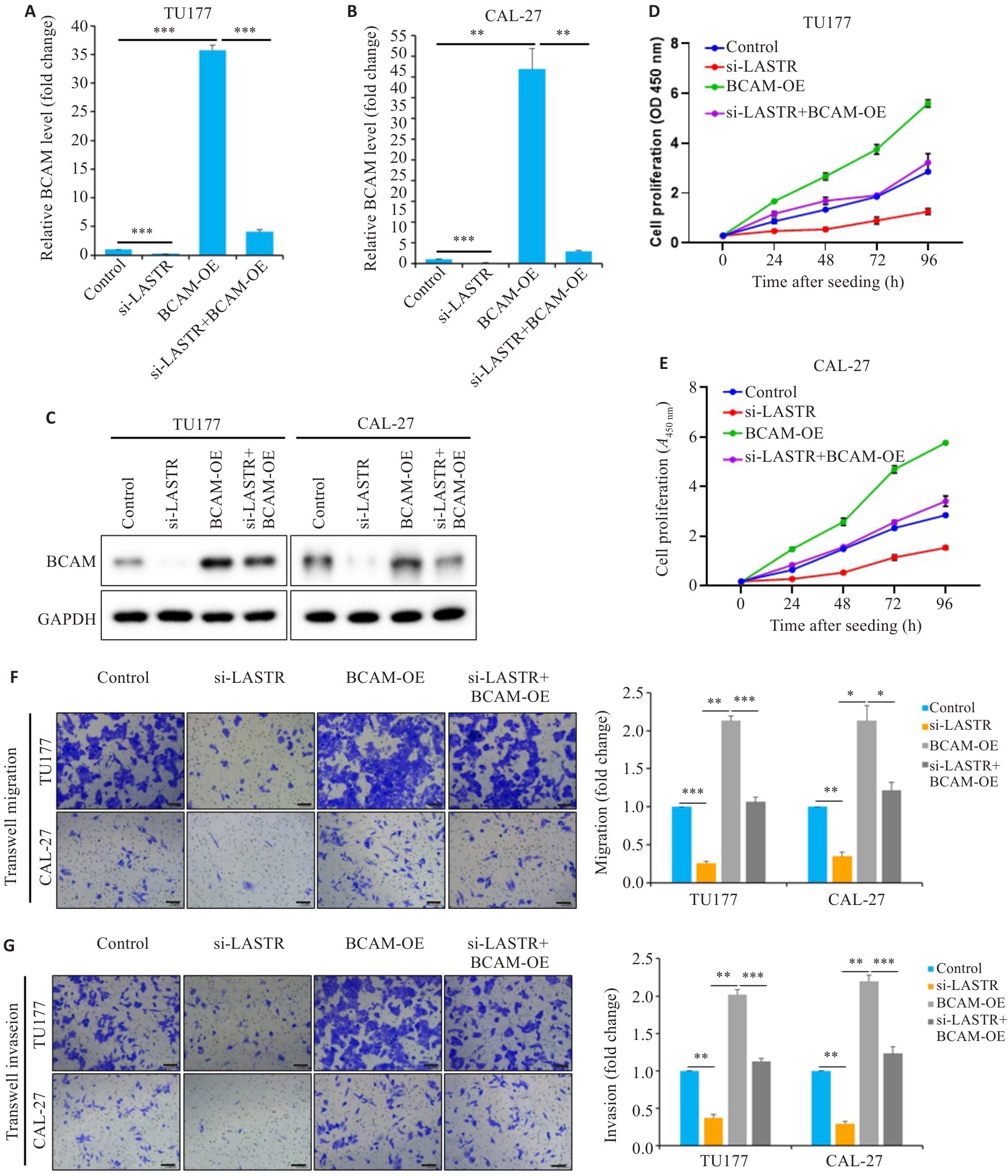
图7 LASTR通过上调BCAM表达水平促进头颈鳞癌细胞恶性表型
Fig. 7 LASTR promotes malignant phenotypes of HNSCC cells via upregulating BCAM expression. TU177 and CAL-27 cells were transfected with si-LASTR, BCAM overexpression plasmid (BCAM-OE), or co-transfected with si-LASTR and BCAM-OE, respectively. A, B: After 48 h transfection, the expression of BCAM mRNA was determined by qPCR (n=3). C: Western blotting for detecting expression of BCAM protein. D, E: CCK8 assay for assessing proliferation of the transfected cells (n=3). F: Migration capacity of the transfected cells determined by Transwell assay (n=3). G: Invasion capacity of the transfected HNSCC cells determined by Transwell assay. Scale bar=50 μm. Data are presented as Mean±SD. *P<0.05, **P<0.01, ***P<0.001.
| [1] | Goetz JW, Rabinowits G, Kalman N, et al. A review of immunotherapy for head and neck cancer[J]. J Dent Res, 2024, 103(12): 1185-96. doi:10.1177/00220345241271992 |
| [2] | Nguyen KV, Lee CML, Choe JH. Metastatic head and neck cancer: immunotherapy and beyond[J]. Semin Radiat Oncol, 2025, 35(2): 259-70. doi:10.1016/j.semradonc.2025.02.004 |
| [3] | Mell LK, Torres-Saavedra PA, Wong SJ, et al. Radiotherapy with cetuximab or durvalumab for locoregionally advanced head and neck cancer in patients with a contraindication to cisplatin (NRG-HN004): an open-label, multicentre, parallel-group, randomised, phase 2/3 trial[J]. Lancet Oncol, 2024, 25(12): 1576-88. doi:10.1016/S1470-2045(24)00507-2 |
| [4] | Coan M, Haefliger S, Ounzain S, et al. Targeting and engineering long non-coding RNAs for cancer therapy[J]. Nat Rev Genet, 2024, 25(8): 578-95. doi:10.1038/s41576-024-00693-2 |
| [5] | Cinque S, Verheyden Y, Adnane S, et al. The assembly of cancer-specific ribosomes by the lncRNA LISRR suppresses melanoma anti-tumor immunity[J]. J Exp Med, 2026, 223(1): e20251507. doi:10.1084/jem.20251507 |
| [6] | Bian ZH, Yang F, Xu PW, et al. LINC01852 inhibits the tumorigenesis and chemoresistance in colorectal cancer by suppressing SRSF5-mediated alternative splicing of PKM[J]. Mol Cancer, 2024, 23(1): 23. doi:10.1186/s12943-024-01939-7 |
| [7] | Alkhathami AG, Pallathadka H, Shah S, et al. LncRNAs in modulating cancer cell resistance to paclitaxel (PTX) therapy[J]. Med Oncol, 2024, 42(1): 28. doi:10.1007/s12032-024-02577-1 |
| [8] | Wu ZG, Zhao Y, Peng YZ, et al. LncRNA TMEM99 complexes with IGF2BP2 to inhibit autophagy in lung adenocarcinoma[J]. Adv Sci, 2025, 12(33): e07871. doi:10.1002/advs.202507871 |
| [9] | Lu Q, Lv XH, Wang J, et al. Methylation-mediated LncRNA CRAT40 promotes colorectal cancer progression by recruiting YBX1 to initiate RelA transcription[J]. Int J Biol Sci, 2025, 21(11): 4834-50. doi:10.7150/ijbs.105629 |
| [10] | Wang ZY, Li XG, Liu ZD, et al. LncRNA HANR promotes the aerobic glycolysis in prostate cancer by stabilizing TPI1[J]. Exp Cell Res, 2025, 452(1): 114744. doi:10.1016/j.yexcr.2025.114744 |
| [11] | Yu ZL, Xu KY, Shang YN, et al. LncRNA LINC00667 inhibits breast cancer progression by regulating POTEE to suppress mitochondrial oxidative phosphorylation[J]. Cell Signal, 2025, 135: 112074. doi:10.1016/j.cellsig.2025.112074 |
| [12] | Wang GJ, Sun LB, Wang SS, et al. Ferroptosis-related long non-coding RNAs and the roles of LASTR in stomach adenocarcinoma[J]. Mol Med Rep, 2022, 25(4). DOI:10.3892/mmr.2022.12634 . |
| [13] | Kong FM, Yang XY, Lu ZC, et al. A novel long noncoding RNA (lncRNA), LINC02657(LASTR), is a prognostic biomarker associated with immune infiltrates of lung adenocarcinoma based on unsupervised cluster analysis[J]. PeerJ, 2023, 11: e16167. doi:10.7717/peerj.16167 |
| [14] | Liu JY, Yao J, Liu JJ, et al. LASTR is a novel prognostic biomarker and predicts response to cancer immunotherapy in gastric cancer[J]. Front Oncol, 2022, 12: 1020255. doi:10.3389/fonc.2022.1020255 |
| [15] | De Troyer L, Zhao PH, Pastor T, et al. Stress-induced lncRNA LASTR fosters cancer cell fitness by regulating the activity of the U4/U6 recycling factor SART3[J]. Nucleic Acids Res, 2020, 48(5): 2502-17. doi:10.1093/nar/gkz1237 |
| [16] | Xia MH, Zhu WB, Tao CM, et al. LncRNA LASTR promote lung cancer progression through the miR-137/TGFA/PI3K/AKT axis through integration analysis[J]. J Cancer, 2022, 13(4): 1086-96. doi:10.7150/jca.66067 |
| [17] | Sivakumar S, Lieber S, Dietze R, et al. Basal cell adhesion molecule (BCAM) promotes mesothelial-to-mesenchymal transition and tumor angiogenesis through paracrine signaling[J]. Cell Commun Signal, 2025, 23(1): 136. doi:10.1186/s12964-025-02128-9 |
| [18] | Sivakumar S, Lieber S, Librizzi D, et al. Basal cell adhesion molecule promotes metastasis-associated processes in ovarian cancer[J]. Clin Transl Med, 2023, 13(1): e1176. doi:10.1002/ctm2.1176 |
| [19] | Burela S, He MN, Trontzas IP, et al. BCAM (basal cell adhesion molecule) protein expression in different tumor populations[J]. Discov Oncol, 2024, 15(1): 381. doi:10.1007/s12672-024-01244-1 |
| [20] | Gao W, Guo HN, Niu M, et al. circPARD3 drives malignant progression and chemoresistance of laryngeal squamous cell carcinoma by inhibiting autophagy through the PRKCI-Akt-mTOR pathway[J]. Mol Cancer, 2020, 19(1): 166. doi:10.1186/s12943-020-01279-2 |
| [21] | Luo LH, Li LY, Liu L, et al. A necroptosis-related lncRNA-based signature to predict prognosis and probe molecular characteristics of stomach adenocarcinoma[J]. Front Genet, 2022, 13: 833928. doi:10.3389/fgene.2022.833928 |
| [22] | Lee TL, Chen TH, Kuo YJ, et al. Tumor-associated tissue eosinophilia promotes angiogenesis and metastasis in head and neck squamous cell carcinoma[J]. Neoplasia, 2023, 35: 100855. doi:10.1016/j.neo.2022.100855 |
| [23] | Huang WH, Li N, Zhu S, et al. Imaging hypoxia for head and neck cancer: current status, challenges, and prospects[J]. Theranostics, 2025, 15(16): 8012-30. doi:10.7150/thno.112781 |
| [24] | Ngan HL, Law CH, Choi YCY, et al. Precision drugging of the MAPK pathway in head and neck cancer[J]. NPJ Genom Med, 2022, 7(1): 20. doi:10.1038/s41525-022-00293-1 |
| [25] | Jagadeeshan S, Novoplansky OZ, Cohen O, et al. New insights into RAS in head and neck cancer[J]. Biochim Biophys Acta Rev Cancer, 2023, 1878(6): 188963. doi:10.1016/j.bbcan.2023.188963 |
| [26] | Akiyama H, Iwahana Y, Suda M, et al. The FBI1/Akirin2 target gene, BCAM acts as a suppressive oncogene[J]. PLoS One, 2013, 8(11): e78716. doi:10.1371/journal.pone.0078716 |
| [27] | Bartolini A, Cardaci S, Lamba S, et al. BCAM and LAMA5 mediate the recognition between tumor cells and the endothelium in the metastatic spreading of KRAS-mutant colorectal cancer[J]. Clin Cancer Res, 2016, 22(19): 4923-33. doi:10.1158/1078-0432.ccr-15-2664 |
| [28] | Li Y, Liu XF, Dong YD, et al. Angiogenesis causes and vasculogenic mimicry formation in the context of cancer stem cells[J]. Biochim Biophys Acta Rev Cancer, 2025, 1880(3): 189323. doi:10.1016/j.bbcan.2025.189323 |
| [29] | 苏 维, 赖厚桦, 唐 昕, 等. Apelin通过激活FGF2/FGFR1通路促进膀胱癌增殖、迁移及血管生成[J]. 南方医科大学学报, 2025, 45(6): 1289-96. |
| [30] | Vassilakopoulou M, Psyrri A, Argiris A. Targeting angiogenesis in head and neck cancer[J]. Oral Oncol, 2015, 51(5): 409-15. doi:10.1016/j.oraloncology.2015.01.006 |
| [31] | Yan TT, Shi JH. Angiogenesis and EMT regulators in the tumor microenvironment in lung cancer and immunotherapy[J]. Front Immunol, 2024, 15: 1509195. doi:10.3389/fimmu.2024.1509195 |
| [32] | Ribatti D, Vacca A. The role of microenvironment in tumor angiogenesis[J]. Genes Nutr, 2008, 3(1): 29-34. doi:10.1007/s12263-008-0076-3 |
| [1] | 苏维, 赖厚桦, 唐昕, 周群, 唐亚纯, 符浩, 陈选才. Apelin通过激活 FGF2/FGFR1通路促进膀胱癌增殖、迁移及血管生成[J]. 南方医科大学学报, 2025, 45(6): 1289-1296. |
| [2] | 邱晓慧, 王檬, 唐江解, 周建大, 金晨. 负载人脐带间充质干细胞外泌体的壳聚糖水凝胶促进慢性糖尿病大鼠的伤口愈合[J]. 南方医科大学学报, 2025, 45(10): 2082-2091. |
| [3] | 李菲凡, 向俊馨, 刘佳慧, 王效静, 江浩. LncRNA FEZF1-AS1通过miR-130a-5p/CCND1轴促进非小细胞肺癌发展的分子机制研究[J]. 南方医科大学学报, 2024, 44(5): 841-850. |
| [4] | 任自敬, 周佩洋, 田静. 基于ceRNA芯片研究帕金森病患者血浆长链非编码RNA表达谱并验证lnc-CTSD-5∶1在PD细胞模型中的作用[J]. 南方医科大学学报, 2024, 44(11): 2146-2155. |
| [5] | 罗 瑞, 田龙海, 杨永曜. 高良姜素通过下调lncRNA H19的表达抑制ox-LDL诱导的人源正常主动脉内皮细胞血管生成活性[J]. 南方医科大学学报, 2024, 44(1): 52-59. |
| [6] | 陈韦任, 杜 辉, 沙 媛, 周玉杰, 梁 静, 陈韵岱, 马 茜, 吴雪萍, 钱 赓. 长链非编码RNA H19促进血管钙化:基于抑制Bax抑制因子1/视神经萎缩蛋白1通路[J]. 南方医科大学学报, 2023, 43(9): 1469-1475. |
| [7] | 冯 雯, 赖跃兴, 王 静, 徐 萍. 长链非编码RNA ABHD11-AS1促进胃癌细胞糖酵解并加速肿瘤恶性进展[J]. 南方医科大学学报, 2023, 43(9): 1485-1492. |
| [8] | 赵齐林, 王 楠, 李亚霁, 吴庆琛, 吴兰香. Lnc-TMEM132D-AS1高表达明显降低非小细胞肺癌对奥希替尼的敏感性[J]. 南方医科大学学报, 2023, 43(2): 242-250. |
| [9] | 吴小凤, 詹日明, 程大钊, 陈 黎, 王天雨, 唐旭东. 非小细胞肺癌细胞外泌体源性FZD10促进体外血管生成[J]. 南方医科大学学报, 2022, 42(9): 1351-1358. |
| [10] | 王 静, 陈雪祎, 孙 丽, 陈雪梅, 李 慧, 熊彬睿, 王海华. 长链非编码RNA ZEB1-AS1通过HMGB1/TLR-4信号轴加剧大鼠脑缺血/再灌注损伤[J]. 南方医科大学学报, 2022, 42(8): 1134-1142. |
| [11] | 刘俊平, 石宇彤, 吴敏敏, 徐梦岐, 张凤梅, 何志强, 唐 敏. JAG1影响血管生成并促进三阴性乳腺癌细胞的迁移、侵袭和粘附[J]. 南方医科大学学报, 2022, 42(7): 1100-1108. |
| [12] | 王 惠, 黎雯雯, 张冬艳. 泛癌分析长链非编码RNA MIR22HG的表达特征[J]. 南方医科大学学报, 2022, 42(4): 473-485. |
| [13] | 蔡 虹, 刘 勉, 林妙玲, 李 红, 沈 朗, 全 松. CCN5在子宫内膜异位症患者组织的表达及其作用机制[J]. 南方医科大学学报, 2022, 42(1): 86-92. |
| [14] | 高莉莉, 张 雄, 窦思雨, 岳小丁, 杨捷玲. 干扰长链编码RNA FOXCUT能抑制鼻咽癌细胞上皮间质转化及诱导线粒体损伤[J]. 南方医科大学学报, 2021, 41(9): 1334-1341. |
| [15] | 孙艳秋, 刘 健, 忻 凌, 周 琴, 陈晓露, 丁 香, 张先恒. 长链非编码RNA Linc00638对类风湿关节炎患者炎症和氧化应激的影响[J]. 南方医科大学学报, 2021, 41(7): 965-971. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||