Journal of Southern Medical University ›› 2025, Vol. 45 ›› Issue (7): 1460-1470.doi: 10.12122/j.issn.1673-4254.2025.07.12
Previous Articles Next Articles
Ting XIE( ), Yunyun WANG, Ting GUO, Chunhua YUAN(
), Yunyun WANG, Ting GUO, Chunhua YUAN( )
)
Received:2025-04-01
Online:2025-07-20
Published:2025-07-17
Contact:
Chunhua YUAN
E-mail:424464387@qq.com;yuanch@smu.edu.cn
Supported by:Ting XIE, Yunyun WANG, Ting GUO, Chunhua YUAN. The peptide toxin components and nucleotide metabolites in Macrothele raveni venom synergistically inhibit cancer cell proliferation by activating the pro-apoptotic pathways[J]. Journal of Southern Medical University, 2025, 45(7): 1460-1470.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.j-smu.com/EN/10.12122/j.issn.1673-4254.2025.07.12
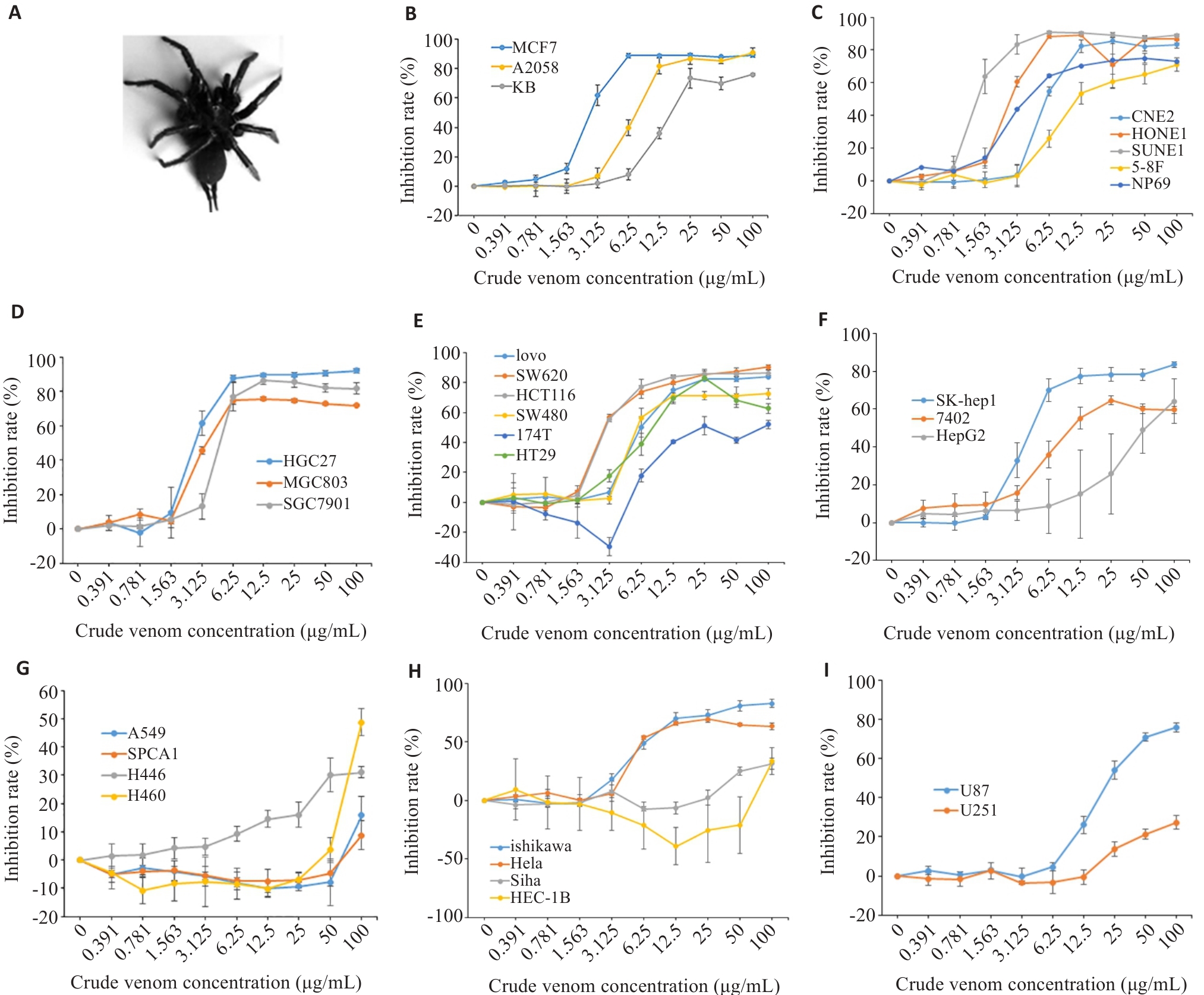
Fig.1 Inhibitory effects of Macrothele raven venom in different cancer cells assessed using CCK-8 assay at 48 h. A: The spider Macrothele raven. B: Inhibitory effects of the venom on proliferation of human breast cancer MCF7 cells, A2058 human Melanoma cells, and human oral epidermoid carcinoma KB cells. C-G: Inhibitory effects of the venom on proliferation of nasopharyngeal carcinoma cells (C), gastric cancer cells (D), colon cancer cells (E), hepatoma cells (F), and lung cancer cells (G). H: Inhibitory effects of the venom on human endometrial cancer Ishikawa cells, human cervical cancer HeLa cells, Siha cells, and human endometrial adenocarcinoma HEC-1B cells. I: Inhibitory effects of the venom on human glioma cells.
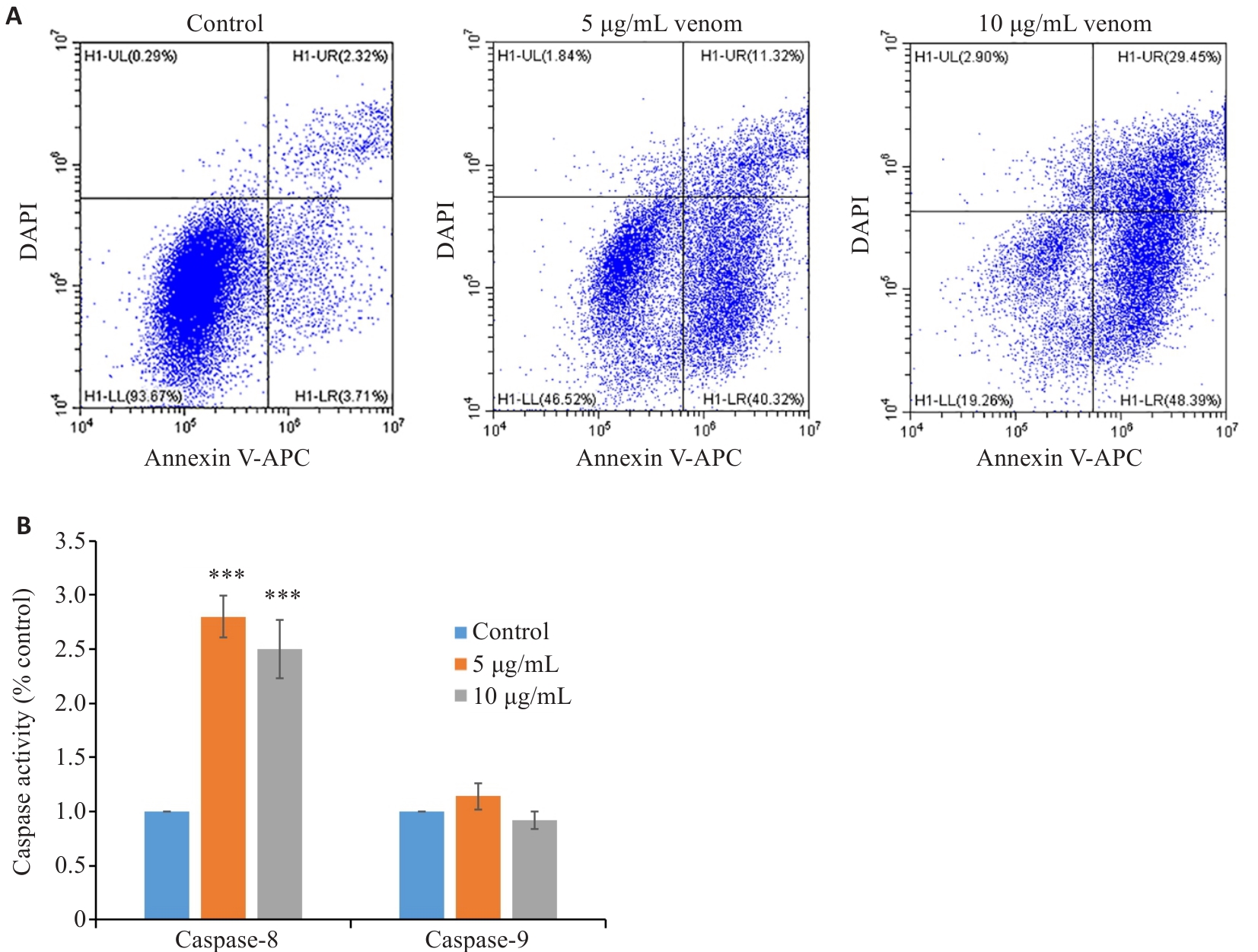
Fig.3 Apoptosis of MCF7 cells following treatment with Macrothele raven venom. A: Flow cytometry of MCF7 cells treated with 5 and 10 µg/mL venom for 24 h. B: Analysis of caspase-8 and 9 activities in MCF7 cells treated with the venom at 5 and 10 µg/mL for 24 h (n=3). ***P<0.001 vs Control.
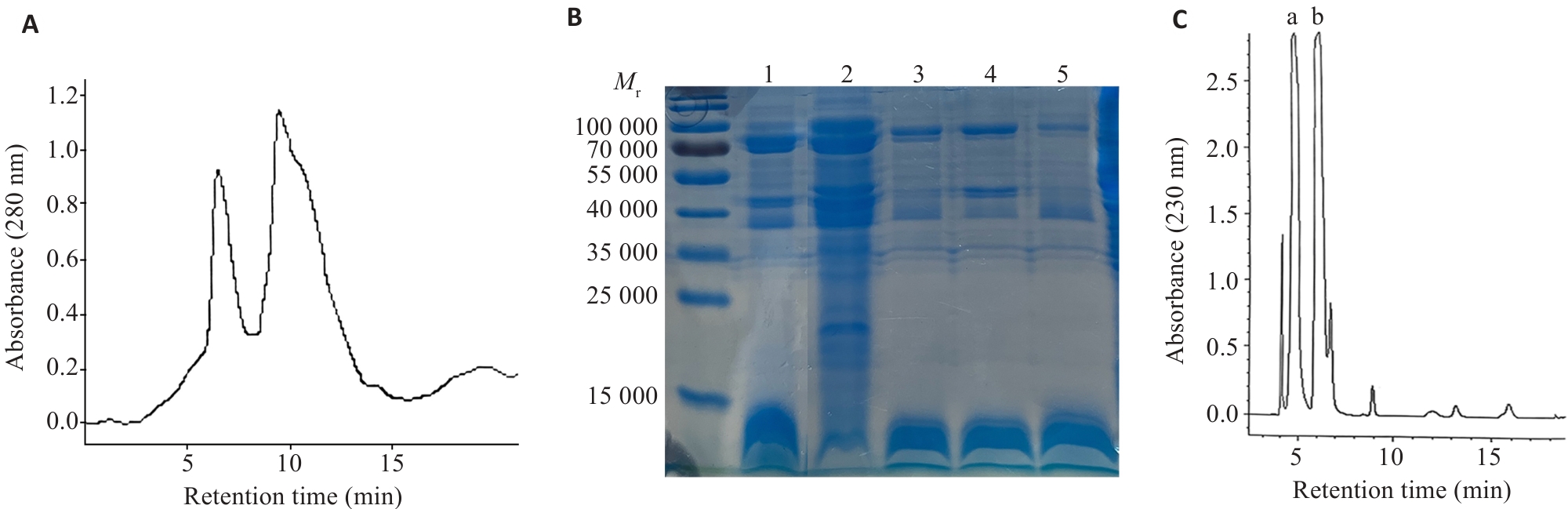
Fig.4 Isolation, purification, and SDS-PAGE molecular weight determination of Macrothele raven venom. A: Separation of the crude venom by gel filtration (Sephadex G-75 column, 10 mm×100 mm). B: Analysis of the eluted fractions by SDS-PAGE with Coomassie Brilliant Blue staining. 1: Crude venom; 2: Fraction eluted at 5 min; 3, 4, 5: Fractions eluted at 10 min of different batches. C: Separation of the small molecule compounds from the crude venom by reversed-phase HPLC.
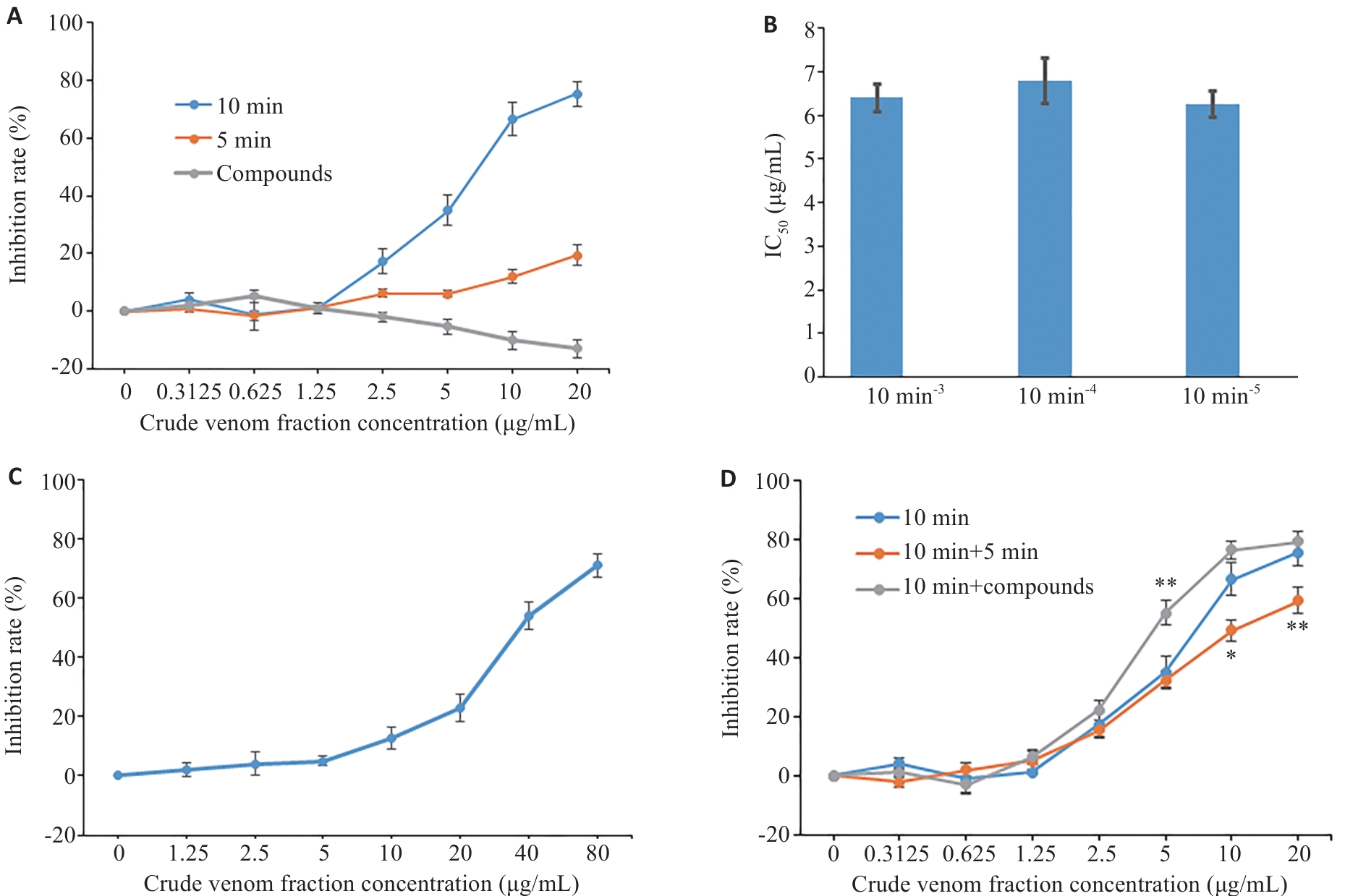
Fig.5 Inhibitory effects of the fractions from Macrothele raveni crude venom on proliferation of MCF-7 cells. A: Inhibitory effects of the 5-min and 10-min fractions separated by gel filtration and small compounds separated by HPLC on human breast cancer MCF-7 cells. B: Inhibitory effects of the fractions eluted at 10 min of different batches on MCF7 cells. C: Inhibitory effects of the 10 min-3 fraction on HEK-293 cells. D: Inhibitory effects of the combinations on human breast cancer MCF-7 cells. Data are presented as Mean±SD (n=3). *P<0.05, **P<0.01 vs 10-min peptide fraction.
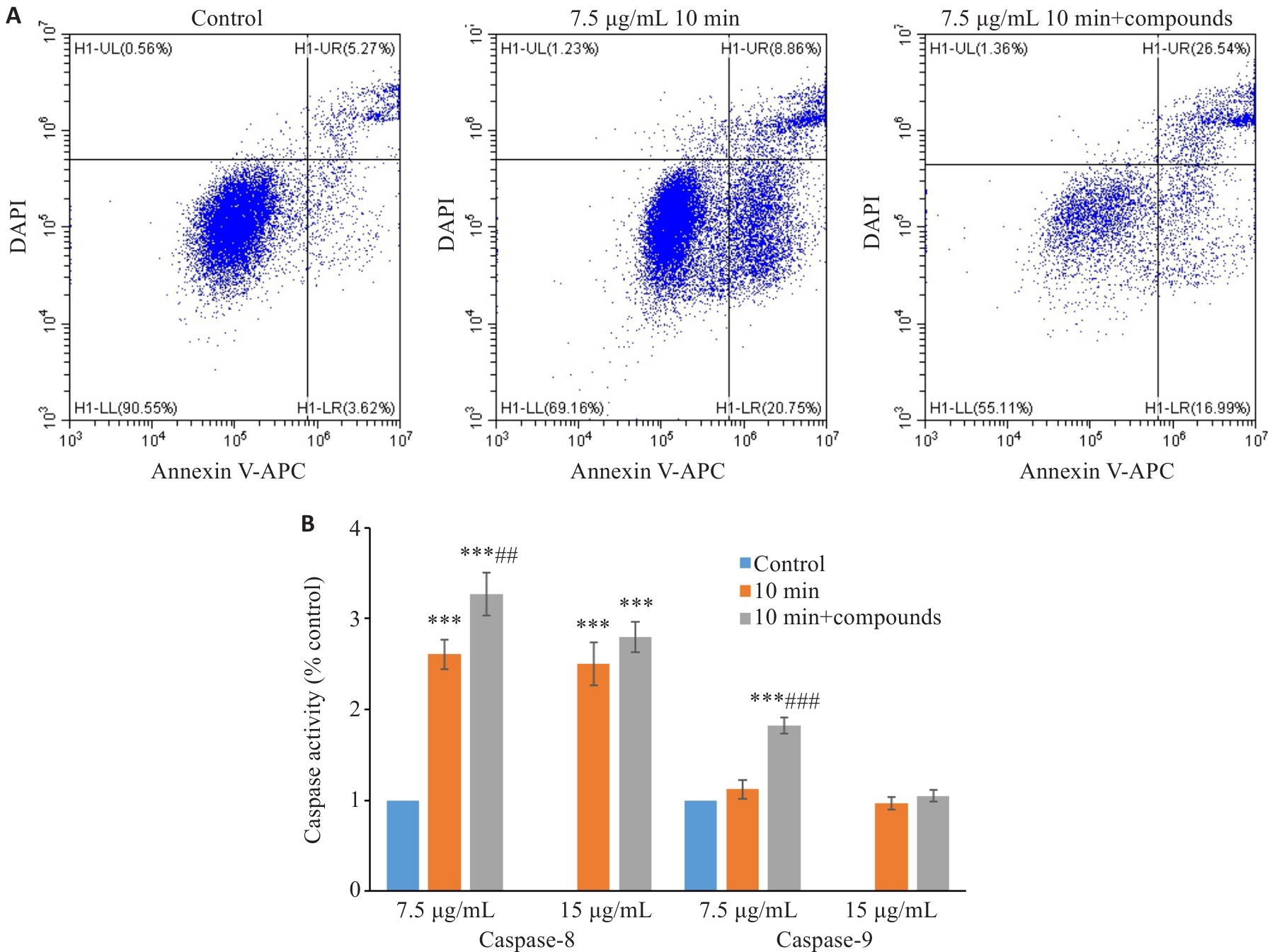
Fig.6 The small molecule compounds from Macrothele raveni venom promoted peptide-induced apoptosis of MCF7 cells. A: Apoptosis of MCF7 cells treated with 7.5 µg/mL venom fractions for 24 h analyzed by flow cytometry (10 min: 10 min peptide fraction). B: Analysis of the activities of caspase 8 and 9 in MCF7 cells treated with the venom fractions at 7.5 and 15 µg/mL for 24 h (n=3). ***P<0.001 vs Control, ##P<0.01, ###P<0.001 vs 10-min peptide fraction.
| Accession No. | Protein | #PSMs | MW | Score sequest HT |
|---|---|---|---|---|
| Q9NFL4 | Hemocyanin G chain | 8 | 71.766 | 17.8 |
| P02241 | Hemocyanin D chain | 9 | 72.077 | 17.6 |
| P02242 | Hemocyanin E chain | 8 | 71.63 | 15.3 |
| P41340 | Actin-3 | 6 | 41.781 | 11.7 |
| Q9NFH9 | Hemocyanin B chain | 7 | 72.052 | 10.7 |
| P14750 | Hemocyanin A chain | 4 | 72.273 | 6.0 |
| P0C2V1 | U15-hexatoxin-Mg1a | 2 | 6.946 | 4.7 |
| P80476 | Hemocyanin AA6 chain | 3 | 71.74 | 3.8 |
| W4VS99 | Neprilysin-1 | 1 | 82.004 | 2.4 |
| B4GEL3 | E3 UFM1-protein ligase 1 homolog | 1 | 87.486 | 2.1 |
| Q9NFL6 | Hemocyanin C chain | 2 | 72.521 | 2.1 |
| Q9VL52 | Putative ATP-dependent RNA helicase | 1 | 168.374 | 2.0 |
Tab.1 Identified proteins in the 5-min component separated by gel filtration chromatography
| Accession No. | Protein | #PSMs | MW | Score sequest HT |
|---|---|---|---|---|
| Q9NFL4 | Hemocyanin G chain | 8 | 71.766 | 17.8 |
| P02241 | Hemocyanin D chain | 9 | 72.077 | 17.6 |
| P02242 | Hemocyanin E chain | 8 | 71.63 | 15.3 |
| P41340 | Actin-3 | 6 | 41.781 | 11.7 |
| Q9NFH9 | Hemocyanin B chain | 7 | 72.052 | 10.7 |
| P14750 | Hemocyanin A chain | 4 | 72.273 | 6.0 |
| P0C2V1 | U15-hexatoxin-Mg1a | 2 | 6.946 | 4.7 |
| P80476 | Hemocyanin AA6 chain | 3 | 71.74 | 3.8 |
| W4VS99 | Neprilysin-1 | 1 | 82.004 | 2.4 |
| B4GEL3 | E3 UFM1-protein ligase 1 homolog | 1 | 87.486 | 2.1 |
| Q9NFL6 | Hemocyanin C chain | 2 | 72.521 | 2.1 |
| Q9VL52 | Putative ATP-dependent RNA helicase | 1 | 168.374 | 2.0 |
| Accession No. | Protein | #PSMs | MW | Score sequest HT |
|---|---|---|---|---|
| P68423 | U4-theraphotoxin-Hs1a | 59 | 3.134 | 254.7 |
| P0C2V2 | U15-hexatoxin-Mg1b | 32 | 6.904 | 101.0 |
| P0C2V1 | U15-hexatoxin-Mg1a | 30 | 6.946 | 88.0 |
| Q75WH4 | U10-hexatoxin-Mg1a | 9 | 8.876 | 35.2 |
| Q75WH1 | Mu-hexatoxin-Mg1c | 14 | 11.64 | 34.2 |
| P02241 | Hemocyanin D chain | 8 | 72.077 | 26.9 |
| Q9NFL4 | Hemocyanin G chain | 8 | 71.766 | 25.2 |
| P83558 | Mu-hexatoxin-Mg1a | 8 | 14.179 | 24.0 |
| Q75WG7 | U13-hexatoxin-Mg1a | 8 | 13.18 | 21.2 |
| P0DL75 | RTX-VII | 5 | 8.706 | 18.8 |
| P83561 | Beta-hexatoxin-Mg1a | 5 | 8.785 | 17.5 |
| P14750 | Hemocyanin A chain | 5 | 72.273 | 15.2 |
| P02242 | Hemocyanin E chain | 4 | 71.63 | 13.4 |
| P02572 | Actin-42A | 5 | 41.797 | 10.5 |
| Q9NFH9 | Hemocyanin B chain | 6 | 72.052 | 10.2 |
| P83559 | Mu-hexatoxin-Mg2a | 3 | 5.229 | 8.9 |
| P56676 | Mu/omega-theraphotoxin-Hs1a | 2 | 9.313 | 5.5 |
| P61233 | U5-hexatoxin-Mr1a(Raventoxin-1) | 1 | 4.845 | 4.3 |
| C0JB02 | Phospholipase D | 1 | 30.78 | 4.0 |
| Q9NFL6 | Hemocyanin C chain | 2 | 72.521 | 4.0 |
| P82959 | U1-theraphotoxin-Hs1a | 1 | 9.423 | 3.5 |
| Q86C49 | U1-theraphotoxin-Hs1f | 1 | 9.162 | 3.4 |
Tab.2 Identified peptides and proteins in the 10-min component separated by gel filtration chromatography
| Accession No. | Protein | #PSMs | MW | Score sequest HT |
|---|---|---|---|---|
| P68423 | U4-theraphotoxin-Hs1a | 59 | 3.134 | 254.7 |
| P0C2V2 | U15-hexatoxin-Mg1b | 32 | 6.904 | 101.0 |
| P0C2V1 | U15-hexatoxin-Mg1a | 30 | 6.946 | 88.0 |
| Q75WH4 | U10-hexatoxin-Mg1a | 9 | 8.876 | 35.2 |
| Q75WH1 | Mu-hexatoxin-Mg1c | 14 | 11.64 | 34.2 |
| P02241 | Hemocyanin D chain | 8 | 72.077 | 26.9 |
| Q9NFL4 | Hemocyanin G chain | 8 | 71.766 | 25.2 |
| P83558 | Mu-hexatoxin-Mg1a | 8 | 14.179 | 24.0 |
| Q75WG7 | U13-hexatoxin-Mg1a | 8 | 13.18 | 21.2 |
| P0DL75 | RTX-VII | 5 | 8.706 | 18.8 |
| P83561 | Beta-hexatoxin-Mg1a | 5 | 8.785 | 17.5 |
| P14750 | Hemocyanin A chain | 5 | 72.273 | 15.2 |
| P02242 | Hemocyanin E chain | 4 | 71.63 | 13.4 |
| P02572 | Actin-42A | 5 | 41.797 | 10.5 |
| Q9NFH9 | Hemocyanin B chain | 6 | 72.052 | 10.2 |
| P83559 | Mu-hexatoxin-Mg2a | 3 | 5.229 | 8.9 |
| P56676 | Mu/omega-theraphotoxin-Hs1a | 2 | 9.313 | 5.5 |
| P61233 | U5-hexatoxin-Mr1a(Raventoxin-1) | 1 | 4.845 | 4.3 |
| C0JB02 | Phospholipase D | 1 | 30.78 | 4.0 |
| Q9NFL6 | Hemocyanin C chain | 2 | 72.521 | 4.0 |
| P82959 | U1-theraphotoxin-Hs1a | 1 | 9.423 | 3.5 |
| Q86C49 | U1-theraphotoxin-Hs1f | 1 | 9.162 | 3.4 |
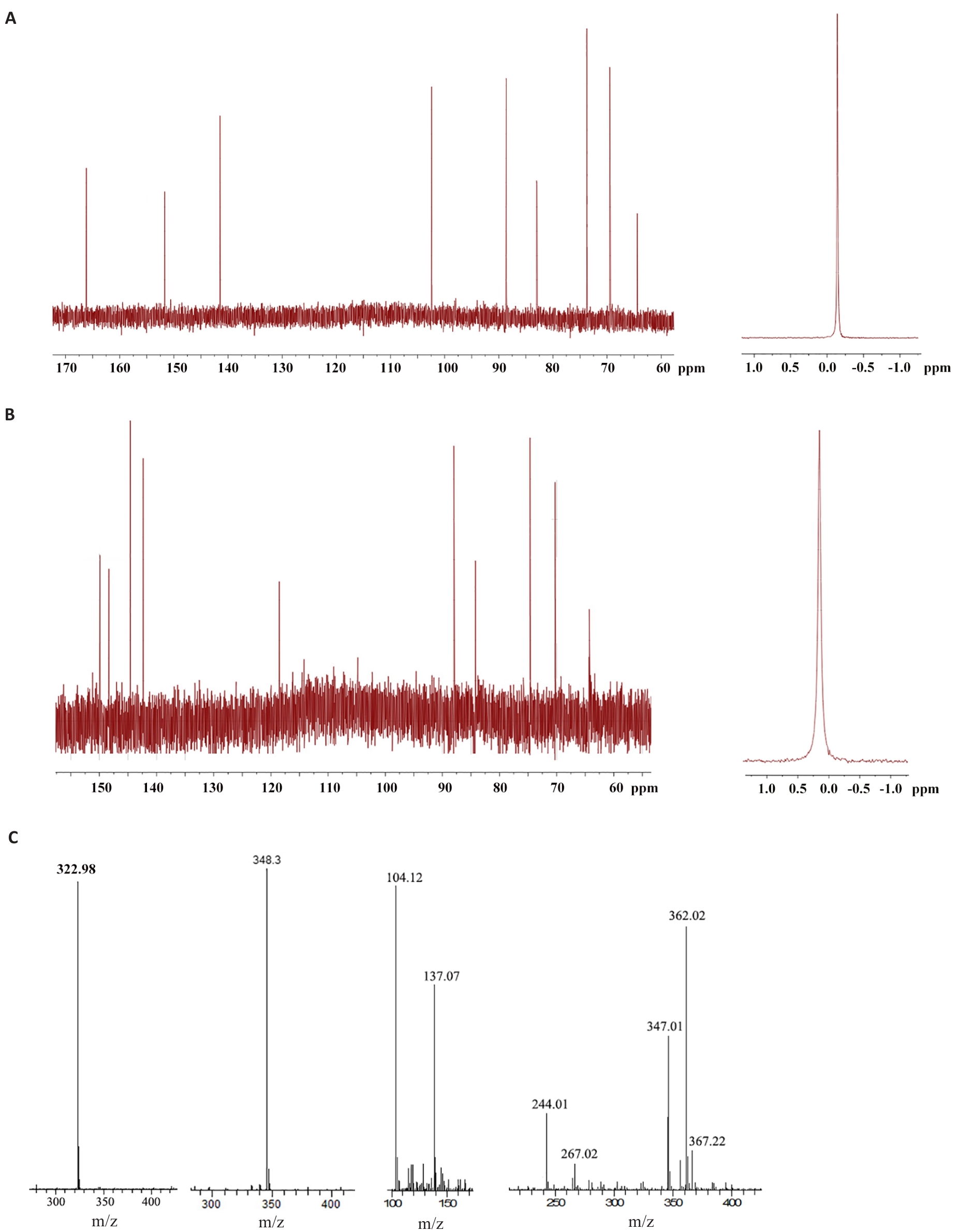
Fig.7 Structural analysis of small molecule compounds isolated from Macrothele raveni venom. A: 600 MHz 13C NMR (left) and 31P NMR (right) spectrum of sample of peak a in Fig.4C. B: 600 MHz 13C NMR (left) and 31P NMR (right) spectrum of sample of peak b in Fig.4C. C: Mass spectrometry of the small molecule compounds isolated from Macrothele raveni venom.
| ID | a peak | UMP | b peak | AMP |
|---|---|---|---|---|
| 1 | 64.4 | 64.4 | 64.30 | 64.25 |
| 2 | 69.5 | 71 | 64.33 | 64.28 |
| 3 | 73.74 | 74.9 | 70.27 | 70.31 |
| 4 | 83.01 | 84.9 | 74.66 | 74.67 |
| 5 | 88.63 | 89.4 | 84.21 | 84.27 |
| 6 | 102.41 | 103.5 | 84.27 | 84.32 |
| 7 | 141.47 | 143 | 87.99 | 87.91 |
| 8 | 151.69 | 152.7 | 118.54 | 118.55 |
| 9 | 166.16 | 167.1 | 142.35 | 142.21 |
| 10 | 144.61 | 145.10 | ||
| 11 | 148.35 | 148.40 | ||
| 12 | 149.91 | 150.25 |
Tab.3 13C NMR chemical shifts (ppm): comparison between sample and literature or standard data
| ID | a peak | UMP | b peak | AMP |
|---|---|---|---|---|
| 1 | 64.4 | 64.4 | 64.30 | 64.25 |
| 2 | 69.5 | 71 | 64.33 | 64.28 |
| 3 | 73.74 | 74.9 | 70.27 | 70.31 |
| 4 | 83.01 | 84.9 | 74.66 | 74.67 |
| 5 | 88.63 | 89.4 | 84.21 | 84.27 |
| 6 | 102.41 | 103.5 | 84.27 | 84.32 |
| 7 | 141.47 | 143 | 87.99 | 87.91 |
| 8 | 151.69 | 152.7 | 118.54 | 118.55 |
| 9 | 166.16 | 167.1 | 142.35 | 142.21 |
| 10 | 144.61 | 145.10 | ||
| 11 | 148.35 | 148.40 | ||
| 12 | 149.91 | 150.25 |
| Sample | Small-molecule compounds | Standards | |||
|---|---|---|---|---|---|
| MW | Retention time (min) | Standards | MW | Retention time (min) | |
| 1 | 322.98 (-) | 3.41 | UMP | 324.2 | 3.43 |
| 2 | 348.3 (-) | 3.64 | AMP | 347.2 | 3.62 |
| 3 | 104.12 (+) | 15.03 | GABA | 103.1 | 15.08 |
| 4 | 137.07 (+) | 4.14 | Hypoxanthine | 136.1 | 4.11 |
| 5 | 244.01 (-) | 4.91 | |||
| 6 | 267.02 (-) | 4.45 | Inosine | 268.2 | 4.41 |
| 7 | 347.01 (-) | 3.54 | IMP | 348.2 | 3.52 |
| 8 | 362.02 (-) | 3.71 | GMP | 363.2 | 3.77 |
| 9 | 367.22 (-) | 5.74 | |||
Tab.4 Comparison relative molecular mass and retention time between small-molecule compounds from Macrothele raveni venom and standards
| Sample | Small-molecule compounds | Standards | |||
|---|---|---|---|---|---|
| MW | Retention time (min) | Standards | MW | Retention time (min) | |
| 1 | 322.98 (-) | 3.41 | UMP | 324.2 | 3.43 |
| 2 | 348.3 (-) | 3.64 | AMP | 347.2 | 3.62 |
| 3 | 104.12 (+) | 15.03 | GABA | 103.1 | 15.08 |
| 4 | 137.07 (+) | 4.14 | Hypoxanthine | 136.1 | 4.11 |
| 5 | 244.01 (-) | 4.91 | |||
| 6 | 267.02 (-) | 4.45 | Inosine | 268.2 | 4.41 |
| 7 | 347.01 (-) | 3.54 | IMP | 348.2 | 3.52 |
| 8 | 362.02 (-) | 3.71 | GMP | 363.2 | 3.77 |
| 9 | 367.22 (-) | 5.74 | |||
| [1] | Tan KK, Bay BH, Gopalakrishnakone P. L-amino acid oxidase from snake venom and its anticancer potential[J]. Toxicon, 2018, 144: 7-13. doi:10.1016/j.toxicon.2018.01.015 |
| [2] | Ullah A. Structure-function studies and mechanism of action of snake venom L-amino acid oxidases[J]. Front Pharmacol, 2020, 11: 110. doi:10.3389/fphar.2020.00110 |
| [3] | Akhtar B, Muhammad F, Sharif A, et al. Mechanistic insights of snake venom disintegrins in cancer treatment[J]. Eur J Pharmacol, 2021, 899: 174022. doi:10.1016/j.ejphar.2021.174022 |
| [4] | Schönthal AH, Swenson SD, Chen TC, et al. Preclinical studies of a novel snake venom-derived recombinant disintegrin with antitumor activity: a review[J]. Biochem Pharmacol, 2020, 181: 114149. doi:10.1016/j.bcp.2020.114149 |
| [5] | Dardevet L, Rani D, Aziz TA, et al. Chlorotoxin: a helpful natural scorpion peptide to diagnose glioma and fight tumor invasion[J]. Toxins (Basel), 2015, 7(4): 1079-101. doi:10.3390/toxins7041079 |
| [6] | Duan XF, Zou HY, Yang JZ, et al. Melittin-incorporated nanomedicines for enhanced cancer immunotherapy[J]. J Control Release, 2024, 375: 285-99. doi:10.1016/j.jconrel.2024.08.047 |
| [7] | Zhang P, Luo W, Zhang ZX, et al. A lipid-sensitive spider peptide toxin exhibits selective anti-leukemia efficacy through multimodal mechanisms[J]. Adv Sci (Weinh), 2024, 11(32): e2404937. doi:10.1002/advs.202404937 |
| [8] | Buglione A, Alloisio G, Ciaccio C, et al. GsMTx-4 venom toxin antagonizes biophysical modulation of metastatic traits in human osteosarcoma cells[J]. Eur J Cell Biol, 2025, 104(1): 151469. doi:10.1016/j.ejcb.2024.151469 |
| [9] | 胡增祥, 杜昱蕾, 刘全喜, 等. 雷氏大疣蛛蜘蛛毒素对人肺癌细胞A549增殖的影响[J]. 中国肺癌杂志, 2010, 13(10): 933-6. doi:10.3779/j.issn.1009-3419.2010.10.02 |
| [10] | 高 莉, 沈金宝, 孙 捷, 等. 雷氏大疣蛛毒液对人肝癌HepG2细胞p21基因表达的影响[J]. 生理学报, 2007, 59(1): 58-62. doi:10.3321/j.issn:0371-0874.2007.01.010 |
| [11] | Liu ZH, Zhao Y, Li J, et al. The venom of the spider Macrothele raveni induces apoptosis in the myelogenous leukemia K562 cell line[J]. Leuk Res, 2012, 36(8): 1063-6. doi:10.1016/j.leukres.2012.02.025 |
| [12] | Gottlieb B, Trifiro M, Batist G. Why tumor genetic heterogeneity may require rethinking cancer genesis and treatment[J]. Trends Cancer, 2021, 7(5): 400-9. doi:10.1016/j.trecan.2020.10.013 |
| [13] | Fruman DA, Chiu H, Hopkins BD, et al. The PI3K pathway in human disease[J]. Cell, 2017, 170(4): 605-35. doi:10.1016/j.cell.2017.07.029 |
| [14] | Bi SY, Chen W, Fang YY, et al. Cancer cell-selective membrane receptor clustering driven by VEGF secretion for in vivo therapy[J]. J Am Chem Soc, 2023, 145(9): 5041-52. doi:10.1021/jacs.2c10428 |
| [15] | Dobrzyńska I, Szachowicz-Petelska B, Sulkowski S, et al. Changes in electric charge and phospholipids composition in human colorectal cancer cells[J]. Mol Cell Biochem, 2005, 276(1): 113-9. doi:10.1007/s11010-005-3557-3 |
| [16] | Chang Y, Chen JY, Yang J, et al. Targeting the Cell Membrane by Charge-Reversal Amphiphilic Pillar[5]arene for the Selective Killing of Cancer Cells[J]. ACS Appl Mater Interfaces, 2019, 11(42): 38497-502. doi:10.1021/acsami.9b13492 |
| [17] | Li XL, Chen ZW, Li YM, et al. Optical tweezers study of membrane fluidity in small cell lung cancer cells[J]. Opt Express, 2021, 29(8): 11976. doi:10.1364/oe.420288 |
| [18] | Sahoo G, Samal D, Khandayataray P, et al. A review on caspases: key regulators of biological activities and apoptosis[J]. Mol Neurobiol, 2023, 60(10): 5805-37. doi:10.1007/s12035-023-03433-5 |
| [19] | Yuan JY, Ofengeim D. A guide to cell death pathways[J]. Nat Rev Mol Cell Biol, 2024, 25(5): 379-95. doi:10.1038/s41580-023-00689-6 |
| [20] | Ramaswamy M, Efimova EV, Martinez O, et al. IG20 (MADD splice variant-5), a proapoptotic protein, interacts with DR4/DR5 and enhances TRAIL-induced apoptosis by increasing recruitment of FADD and caspase-8 to the DISC[J]. Oncogene, 2004, 23(36): 6083-94. doi:10.1038/sj.onc.1207804 |
| [21] | Capera J, Pérez-Verdaguer M, Peruzzo R, et al. A novel mitochondrial Kv1.3-caveolin axis controls cell survival and apoptosis[J]. eLife, 2021, 10: e69099. doi:10.7554/elife.69099 |
| [22] | Teisseyre A, Chmielarz M, Uryga A, et al. Co-application of statin and flavonoids as an effective strategy to reduce the activity of voltage-gated potassium channels Kv1.3 and induce apoptosis in human leukemic T cell line jurkat[J]. Molecules, 2022, 27(10): 3227. doi:10.3390/molecules27103227 |
| [23] | Xiao Z, You X, Chen MZ, et al. A novel spider toxin as a selective antagonist of the Kv1 subfamily of voltage-gated potassium channels through gating modulation[J]. J Biol Chem, 2025, 301(4): 108341. doi:10.1016/j.jbc.2025.108341 |
| [24] | Liang SP. An overview of peptide toxins from the venom of the Chinese bird spider Selenocosmia huwena Wang [=Ornithoctonus huwena (Wang)[J]. Toxicon, 2004, 43(5): 575-85. doi:10.1016/j.toxicon.2004.02.005 |
| [25] | Xiao X, Luo XQ, Huang CL, et al. Transcriptome analysis reveals the peptide toxins diversity of Macrothele palpator venom[J]. Int J Biol Macromol, 2023, 253(Pt 1): 126577. doi:10.1016/j.ijbiomac.2023.126577 |
| [26] | Corzo G, Gilles N, Satake H, et al. Distinct primary structures of the major peptide toxins from the venom of the spider Macrothele gigas that bind to sites 3 and 4 in the sodium channel[J]. FEBS Lett, 2003, 547(1/2/3): 43-50. doi:10.1016/s0014-5793(03)00666-5 |
| [27] | Titaux-Delgado G, Carrillo E, Mendoza A, et al. Successful refolding and NMR structure of rMagi3: a disulfide-rich insecticidal spider toxin[J]. Protein Sci, 2018, 27(3): 692-701. doi:10.1002/pro.3363 |
| [28] | Suchyna TM, Johnson JH, Hamer K, et al. Identification of a peptide toxin from Grammostola spatulata spider venom that blocks cation-selective stretch-activated channels[J]. J Gen Physiol, 2000, 115(5): 583-98. doi:10.1085/jgp.115.5.583 |
| [29] | Boța M, Vlaia L, Jîjie AR, et al. Exploring synergistic interactions between natural compounds and conventional chemotherapeutic drugs in preclinical models of lung cancer[J]. Pharmaceuticals (Basel), 2024, 17(5): 598. doi:10.3390/ph17050598 |
| [30] | He M, Liao QY, Liu D, et al. Dihydroergotamine mesylate enhances the anti-tumor effect of sorafenib in liver cancer cells[J]. Biochem Pharmacol, 2023, 211: 115538. doi:10.1016/j.bcp.2023.115538 |
| [31] | Kumar S, Arora A, Chaudhary R, et al. Recent advances in the synthesis of acyclic nucleosides and their therapeutic applications[J]. Top Curr Chem (Cham), 2024, 382(4): 34. doi:10.1007/s41061-024-00476-7 |
| [32] | Sigmond J, Peters GJ. Pyrimidine and purine analogues, effects on cell cycle regulation and the role of cell cycle inhibitors to enhance their cytotoxicity[J]. Nucleosides Nucleotides Nucleic Acids, 2005, 24(10/11/12): 1997-2022. doi:10.1080/15257770500269556 |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||