Journal of Southern Medical University ›› 2025, Vol. 45 ›› Issue (5): 954-961.doi: 10.12122/j.issn.1673-4254.2025.05.07
Previous Articles Next Articles
Yumei ZENG1( ), Jike LI2, Zhongxi HUANG2, Yibo ZHOU3(
), Jike LI2, Zhongxi HUANG2, Yibo ZHOU3( )
)
Received:2024-09-23
Online:2025-05-20
Published:2025-05-23
Contact:
Yibo ZHOU
E-mail:41274931@qq.com;boyi365@163.com
Yumei ZENG, Jike LI, Zhongxi HUANG, Yibo ZHOU. Villin-like protein VILL suppresses proliferation of nasopharyngeal carcinoma cells by interacting with LMO7 protein[J]. Journal of Southern Medical University, 2025, 45(5): 954-961.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.j-smu.com/EN/10.12122/j.issn.1673-4254.2025.05.07
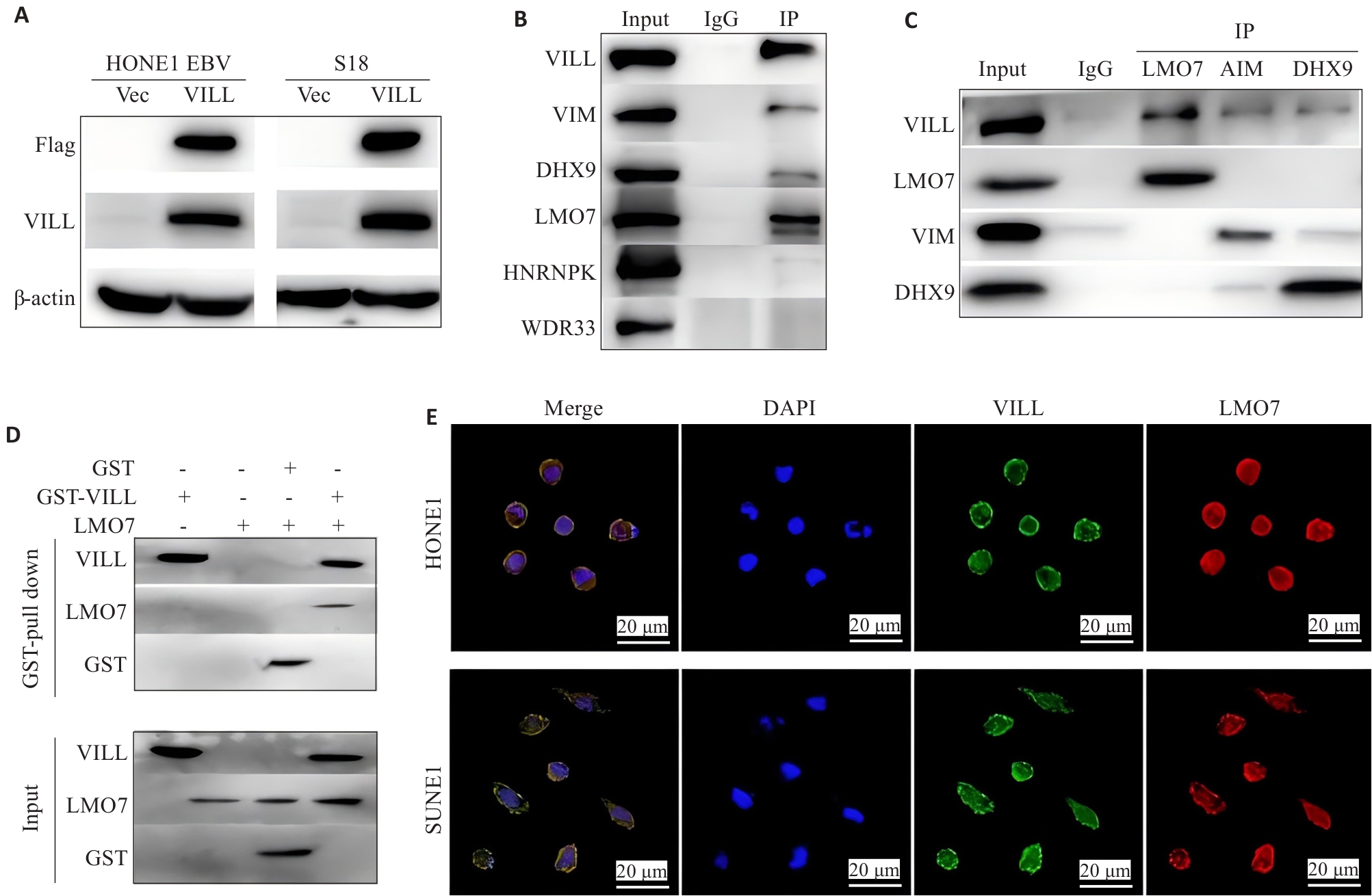
Fig.1 LMO7 directly interacts with VILL protein. A: Verification of successful construction of NPC cell lines (HONE1-EBV and S18) overexpressing VILL+Flag. B: CO-IP assay and Western blotting, using VILL as the bait, for verification of the proteins interacting with VILL. C: CO-IP assay and Western blotting, using LMO7, VIM and DHX9 as the decoys, for verification of the proteins interacting with VILL. D: GST-pull down experiment for verifying direct interaction between LMO7 and VILL. E: Immunofluorescence assay showing co-localization of LMO7 and VILL in NPC cell lines (HONE1 and SUNE1).
| Gene | hit_num | Score | Matches | Sequences | Cover | pi |
|---|---|---|---|---|---|---|
| MYH3 | 2 | 1770 | 58 | 34 | 19.5 | 5.62 |
| ACTB | 8 | 642 | 39 | 12 | 35.8 | 5.23 |
| PLEC | 13 | 524 | 23 | 23 | 5.6 | 5.74 |
| HNRNPK | 14 | 522 | 20 | 10 | 22.5 | 5.39 |
| ACTN2 | 17 | 463 | 17 | 12 | 17 | 5.31 |
| DHX9 | 19 | 419 | 25 | 20 | 16.4 | 6.41 |
| LMO7 | 21 | 398 | 21 | 10 | 16.9 | 8.73 |
| VIM | 22 | 394 | 17 | 12 | 28.5 | 5.06 |
| PABPC1 | 24 | 374 | 14 | 11 | 18.6 | 9.52 |
| RCN1 | 27 | 345 | 14 | 8 | 19.3 | 4.86 |
| DDX3X | 32 | 286 | 14 | 9 | 14.2 | 6.73 |
| DDX5 | 54 | 180 | 7 | 5 | 6.5 | 5.25 |
Tab.1 Proteins predicted to interact with villin-like protein VILL by mass spectrometry
| Gene | hit_num | Score | Matches | Sequences | Cover | pi |
|---|---|---|---|---|---|---|
| MYH3 | 2 | 1770 | 58 | 34 | 19.5 | 5.62 |
| ACTB | 8 | 642 | 39 | 12 | 35.8 | 5.23 |
| PLEC | 13 | 524 | 23 | 23 | 5.6 | 5.74 |
| HNRNPK | 14 | 522 | 20 | 10 | 22.5 | 5.39 |
| ACTN2 | 17 | 463 | 17 | 12 | 17 | 5.31 |
| DHX9 | 19 | 419 | 25 | 20 | 16.4 | 6.41 |
| LMO7 | 21 | 398 | 21 | 10 | 16.9 | 8.73 |
| VIM | 22 | 394 | 17 | 12 | 28.5 | 5.06 |
| PABPC1 | 24 | 374 | 14 | 11 | 18.6 | 9.52 |
| RCN1 | 27 | 345 | 14 | 8 | 19.3 | 4.86 |
| DDX3X | 32 | 286 | 14 | 9 | 14.2 | 6.73 |
| DDX5 | 54 | 180 | 7 | 5 | 6.5 | 5.25 |
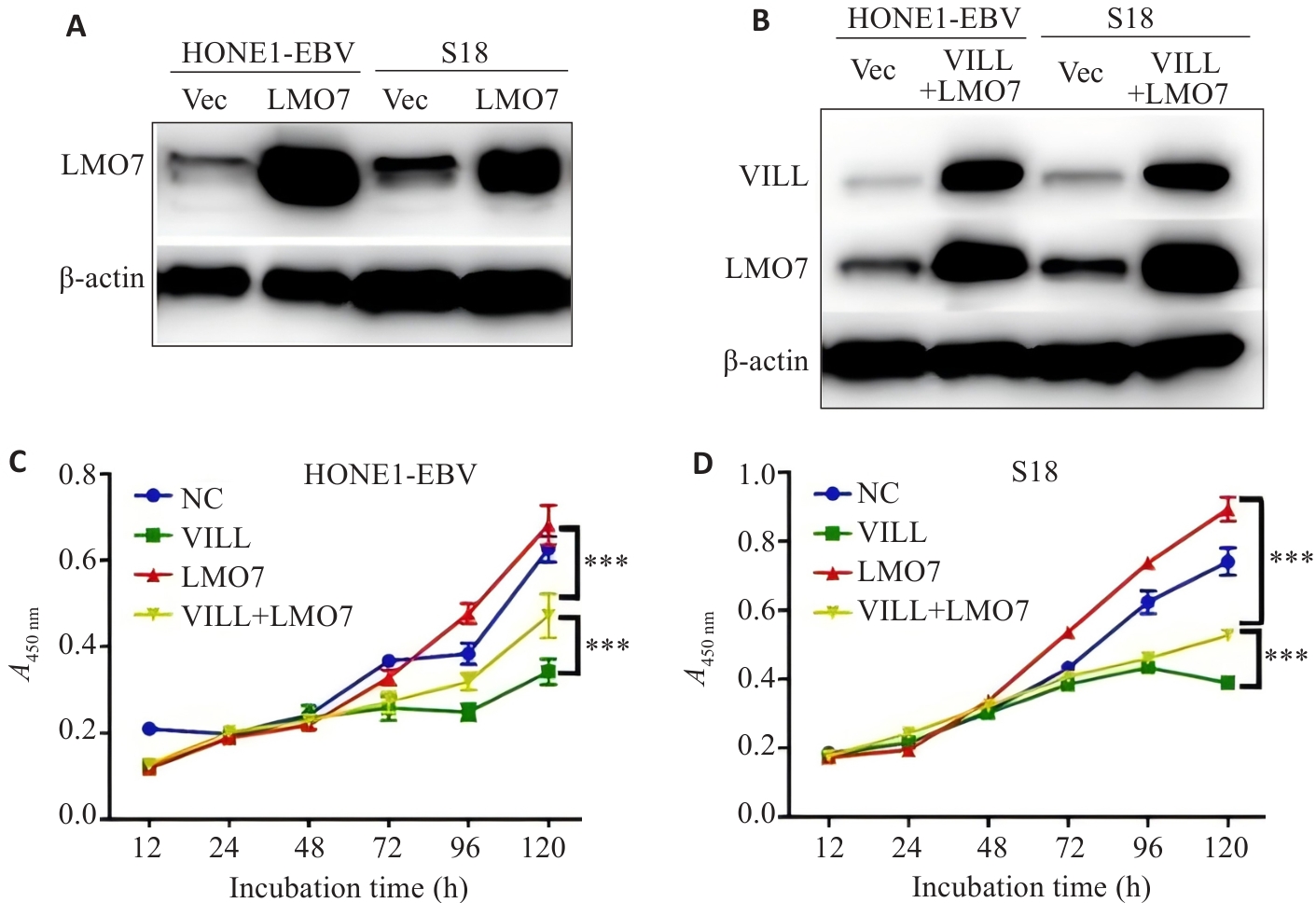
Fig.2 CCK-8 assay showing that LMO7 partially reverses the inhibitory effect of VILL on NPC cell proliferation. A,B: Western blotting for verification of successful construction of NPC cell lines (HONE1-EBV and S18) with LMO7 overexpression (A) and VILL+LMO7 double overexpression (B). C, D: CCK-8 assay showing that LMO7 partially reverses the inhibitory effect of VILL on proliferation of NPC cell lines HONE1-EBV (C) and S18 (D). ***P<0.001.
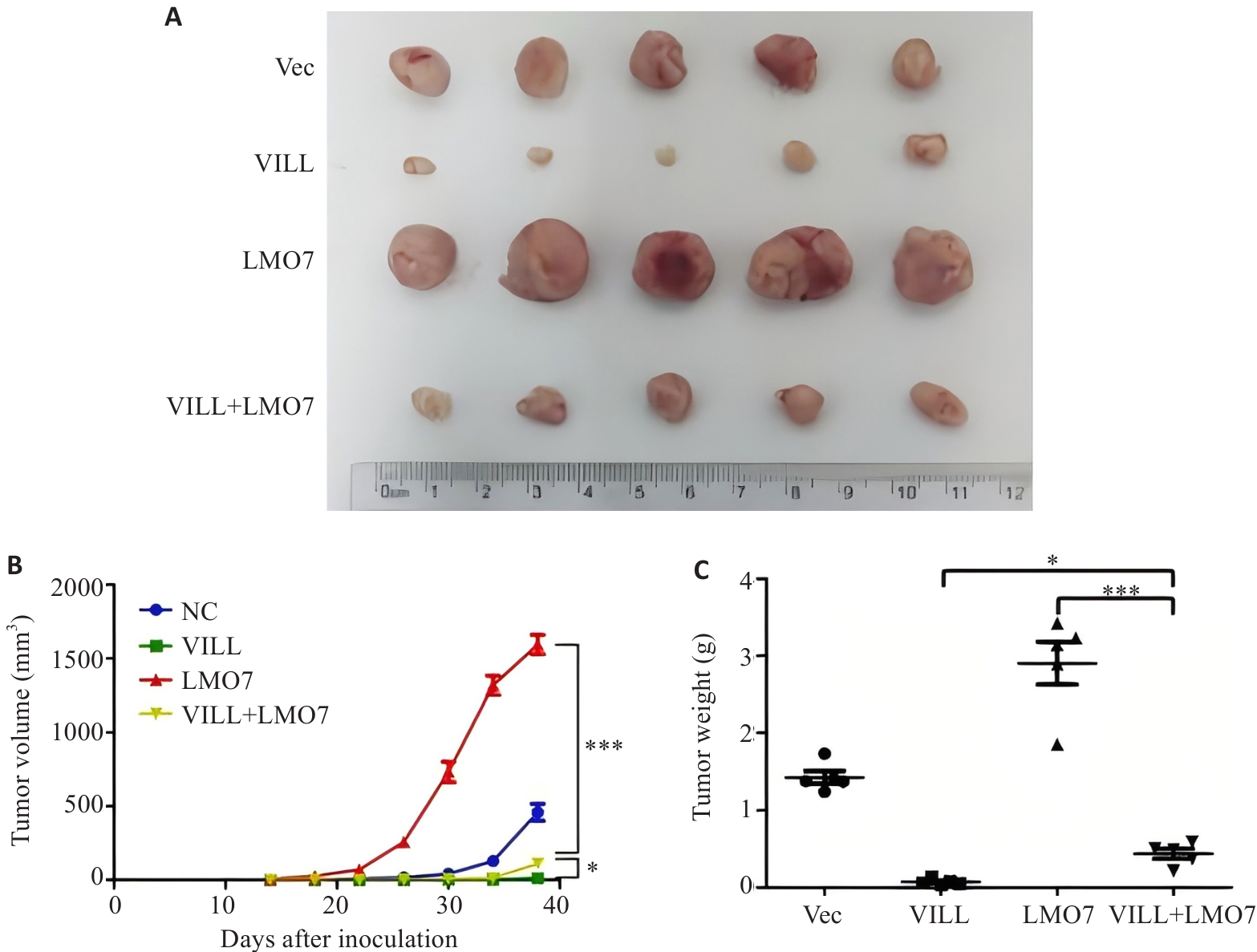
Fig.3 Subcutaneous tumor formation in nude mice showing that LMO7 partially reverses the inhibitory effect of VILL on NPC cell proliferation in vivo. A: Tumors dissected 40 days after subcutaneous implantation of NPC cells (S18) in nude mice. B: Time curve of subcutaneous tumor growth in nude mice. C: Tumor weight 40 days after subcutaneous implantation of NPC cells (S18) in nude mice. *P<0.05, ***P<0.001.
| Variable | n | VILL expression | P (Fisher's Exact Test) | |
|---|---|---|---|---|
| Low | High | |||
| Nasopharyngeal tissues | 67 | 0 (0.0) | 67 (100.0) | P<0.001 |
| NPC | 136 | 99 (72.8) | 37 (27.2) | |
Tab.2 Expression profiles of VILL in 67 non-cancerous naso-pharyngeal tissues and 136 NPC tissues (n, %)
| Variable | n | VILL expression | P (Fisher's Exact Test) | |
|---|---|---|---|---|
| Low | High | |||
| Nasopharyngeal tissues | 67 | 0 (0.0) | 67 (100.0) | P<0.001 |
| NPC | 136 | 99 (72.8) | 37 (27.2) | |
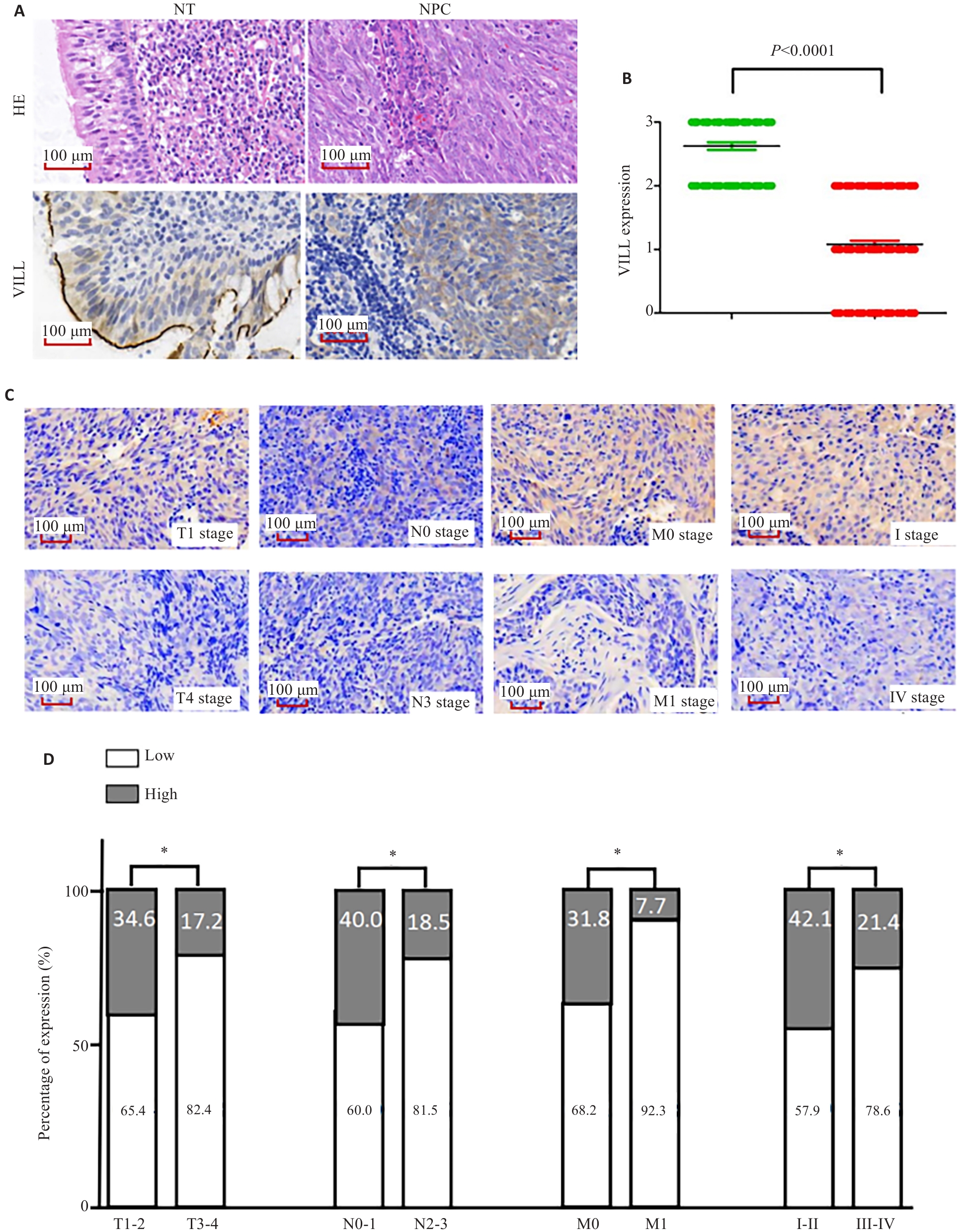
Fig.4 Expression profile of VILL in NPC tissue specimens and its correlation with clinicopathological features of the patients. A, B: Immunohistochemistry showing significant down-regulation of VILL expression in NPC tissues. C: Representative maps of VILL expression in NPC tissues in different TNM stages. D: Positivity rate of VILL in NPC tissues in different TNM stages. *P<0.05.
| Characteristics | Case | VILL expression | χ2 | P | |
|---|---|---|---|---|---|
| Low | High | ||||
| Gender | |||||
| Female | 39 | 27 (69.2) | 12 (30.8) | 0.144 | 0.705 |
| Male | 97 | 72 (74.2) | 25 (25.8) | ||
| Age (years) | |||||
| <50 | 62 | 44 (71.0) | 18 (29.0) | 0.060 | 0.807 |
| ≥50 | 74 | 55 (74.3) | 19 (25.7) | ||
| T classification | |||||
| T1-T2 | 78 | 51 (65.4) | 27 (34.6) | 4.231 | 0.040 |
| T3-T4 | 58 | 48 (82.8) | 10 (17.2) | ||
| N classification | |||||
| N0-N1 | 55 | 33 (60.0) | 22 (40.0) | 6.587 | 0.010 |
| N2-N3 | 81 | 66 (81.5) | 15 (18.5) | ||
| M classification | |||||
| M0 | 110 | 75 (68.2) | 35 (31.8) | 5.023 | 0.013 |
| M1 | 26 | 24 (92.3) | 2 (7.7) | ||
| Clinical stage | |||||
| Ⅰ-Ⅱ | 38 | 22 (57.9) | 16 (42.1) | 4.913 | 0.027 |
| III-IV | 98 | 77 (78.6) | 21 (21.4) | ||
Tab.3 Correlation between VILL expression level and clinico-pathological features of NPC patients (n, %)
| Characteristics | Case | VILL expression | χ2 | P | |
|---|---|---|---|---|---|
| Low | High | ||||
| Gender | |||||
| Female | 39 | 27 (69.2) | 12 (30.8) | 0.144 | 0.705 |
| Male | 97 | 72 (74.2) | 25 (25.8) | ||
| Age (years) | |||||
| <50 | 62 | 44 (71.0) | 18 (29.0) | 0.060 | 0.807 |
| ≥50 | 74 | 55 (74.3) | 19 (25.7) | ||
| T classification | |||||
| T1-T2 | 78 | 51 (65.4) | 27 (34.6) | 4.231 | 0.040 |
| T3-T4 | 58 | 48 (82.8) | 10 (17.2) | ||
| N classification | |||||
| N0-N1 | 55 | 33 (60.0) | 22 (40.0) | 6.587 | 0.010 |
| N2-N3 | 81 | 66 (81.5) | 15 (18.5) | ||
| M classification | |||||
| M0 | 110 | 75 (68.2) | 35 (31.8) | 5.023 | 0.013 |
| M1 | 26 | 24 (92.3) | 2 (7.7) | ||
| Clinical stage | |||||
| Ⅰ-Ⅱ | 38 | 22 (57.9) | 16 (42.1) | 4.913 | 0.027 |
| III-IV | 98 | 77 (78.6) | 21 (21.4) | ||
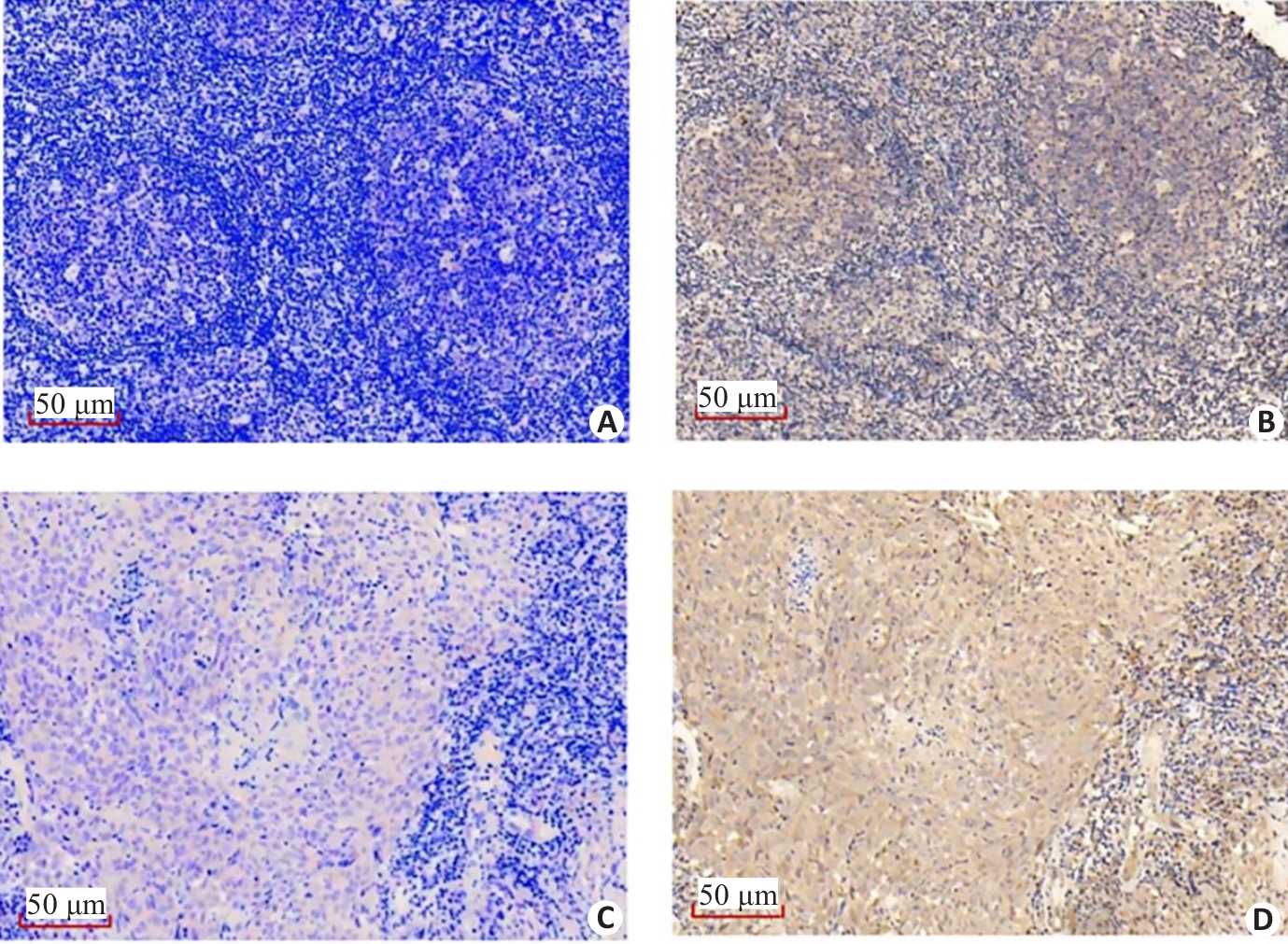
Fig.5 Immunohistochemistry showing the correlation between the expressions of VILL and LMO7 in NPC tissue specimens. VILL is lowly expressed (A) while LMO7 is highly expressed (B) in a NPC tissue specimen. VILL (C) and LMO7 (D) are both highly expressed in a NPC tissue specimen.
| 1 | Chen YP, Chan ATC, Le QT, et al. Nasopharyngeal carcinoma[J]. Lancet, 2019, 394(10192): 64-80. |
| 2 | Sung H, Ferlay J, Siegel RL, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries[J]. CA Cancer J Clin, 2021, 71(3): 209-49. |
| 3 | Han BF, Zheng RS, Zeng HM, et al. Cancer incidence and mortality in China, 2022[J]. J Natl Cancer Cent, 2024, 4(1): 47-53. |
| 4 | Liang YY, Deng XB, Lin XT, et al. RASSF1A inhibits PDGFB-driven malignant phenotypes of nasopharyngeal carcinoma cells in a YAP1-dependent manner[J]. Cell Death Dis, 2020, 11(10): 855. |
| 5 | Zhao Y, Hong XH, Li K, et al. ZNF582 hypermethylation promotes metastasis of nasopharyngeal carcinoma by regulating the transcription of adhesion molecules Nectin-3 and NRXN3[J]. Cancer Commun: Lond, 2020, 40(12): 721-37. |
| 6 | Chen Y, Zhao Y, Yang X, et al. USP44 regulates irradiation-induced DNA double-strand break repair and suppresses tumorigenesis in nasopharyngeal carcinoma[J]. Nat Commun, 2022, 13(1): 501. |
| 7 | Wang YQ, Wu DH, Wei D, et al. TEAD4 is a master regulator of high-risk nasopharyngeal carcinoma[J]. Sci Adv, 2023, 9(1): eadd0960. |
| 8 | Li ZX, Zheng ZQ, Yang PY, et al. WTAP-mediated m6A modification of lncRNA DIAPH1-AS1 enhances its stability to facilitate nasopharyngeal carcinoma growth and metastasis[J]. Cell Death Differ, 2022, 29(6): 1137-51. |
| 9 | Chen B, Huang R, Xia T, et al. The m6A reader IGF2BP3 preserves NOTCH3 mRNA stability to sustain Notch3 signaling and promote tumor metastasis in nasopharyngeal carcinoma[J]. Oncogene, 2023, 42(48): 3564-74. |
| 10 | Zhang B, Li J, Wang Y, et al. Deubiquitinase USP7 stabilizes KDM5B and promotes tumor progression and cisplatin resistance in nasopharyngeal carcinoma through the ZBTB16/TOP2A axis[J]. Cell Death Differ, 2024, 31(3): 309-21. |
| 11 | Sun XS, Zhang K, Peng XZ, et al. HDAC4 mediated LHPP deacetylation enhances its destabilization and promotes the proliferation and metastasis of nasopharyngeal carcinoma[J]. Cancer Lett, 2023, 562: 216158. |
| 12 | Luo SD, Tsai HT, Hwang CF, et al. Aberrant miR-874-3p/leptin/EGFR/c-Myc signaling contributes to nasopharyngeal carcinoma pathogenesis[J]. J Exp Clin Cancer Res, 2022, 41(1): 215. |
| 13 | Wu P, Hou X, Peng M, et al. Circular RNA circRILPL1 promotes nasopharyngeal carcinoma malignant progression by activating the Hippo-YAP signaling pathway[J]. Cell Death Differ, 2023, 30(7): 1679-94. |
| 14 | Bruce JP, To KF, Lui VWY, et al. Whole-genome profiling of nasopharyngeal carcinoma reveals viral-host co-operation in inflammatory NF-κB activation and immune escape[J]. Nat Commun, 2021, 12(1): 4193. |
| 15 | Patel SA, Hirosue S, Rodrigues P, et al. The renal lineage factor PAX8 controls oncogenic signalling in kidney cancer[J]. Nature, 2022, 606(7916): 999-1006. |
| 16 | Schrag D, Beer TM, McDonnell CH, et al. Blood-based tests for multicancer early detection (PATHFINDER): a prospective cohort study[J]. Lancet, 2023, 402(10409): 1251-60. |
| 17 | Huang H, Liu A, Liang Y, et al. A urinary assay for mutation and methylation biomarkers in the diagnosis and recurrence prediction of non-muscle invasive bladder cancer patients[J]. BMC Med, 2023, 21(1): 357. |
| 18 | Jiang W, Liu N, Chen XZ, et al. Genome-wide identification of a methylation gene panel as a prognostic biomarker in nasopharyngeal carcinoma[J]. Mol Cancer Ther, 2015, 14(12): 2864-73. |
| 19 | Guo S, Diep D, Plongthongkum N, et al. Identification of methylation haplotype blocks aids in deconvolution of heterogeneous tissue samples and tumor tissue-of-origin mapping from plasma DNA[J]. Nat Genet, 2017, 49(4): 635-42. |
| 20 | Silacci P, Mazzolai L, Gauci C, et al. Gelsolin superfamily proteins: key regulators of cellular functions[J]. Cell Mol Life Sci CMLS, 2004, 61(19): 2614-23. |
| 21 | Uhlén M, Fagerberg L, Hallström BM, et al. Proteomics. tissue-based map of the human proteome[J]. Science, 2015, 347(6220): 1260419. |
| 22 | Mahajan-Miklos S, Cooley L. The villin-like protein encoded by the Drosophila quail gene is required for actin bundle assembly during oogenesis[J]. Cell, 1994, 78(2): 291-301. |
| 23 | Kang CK, Lee TH. Medaka villin 1-like protein (VILL) is associated with the formation of microvilli induced by decreasing salinities in the absorptive ionocytes[J]. Front Zool, 2014, 11(1): 2. |
| 24 | Zeng Q, Jiang T, Wang J. Role of LMO7 in cancer (review)[J]. Oncol Rep, 2024, 52(3): 117. |
| 25 | Karlsson T, Kvarnbrink S, Holmlund C, et al. LMO7 and LIMCH1 interact with LRIG proteins in lung cancer, with prognostic implications for early-stage disease[J]. Lung Cancer, 2018, 125: 174-84. |
| 26 | Liu X, Yuan H, Zhou J, et al. LMO7 as an unrecognized factor promoting pancreatic cancer progression and metastasis[J]. Front Cell Dev Biol, 2021, 9: 647387. |
| 27 | Takahashi M, Lio CJ, Campeau A, et al. The tumor suppressor kinase DAPK3 drives tumor-intrinsic immunity through the STING-IFN-β pathway[J]. Nat Immunol, 2021, 22(4): 485-96. |
| 28 | Dai S, Peng Y, Wang G, et al. LIM domain only 7: a novel driver of immune evasion through regulatory T cell differentiation and chemotaxis in pancreatic ductal adenocarcinoma[J]. Cell Death Differ, 2025, 32(2): 271-90. |
| [1] | Lu TAO, Zhuoli WEI, Yueyue WANG, Ping XIANG. CEACAM6 inhibits proliferation and migration of nasopharyngeal carcinoma cells by suppressing epithelial-mesenchymal transition [J]. Journal of Southern Medical University, 2025, 45(3): 566-576. |
| [2] | Jinyu LIU, Shujun LIANG, Yu ZHANG. A multi-scale supervision and residual feedback optimization algorithm for improving optic chiasm and optic nerve segmentation accuracy in nasopharyngeal carcinoma CT images [J]. Journal of Southern Medical University, 2025, 45(3): 632-642. |
| [3] | Liangjun XUE, Qiuyu TAN, Jingwen XU, Lu FENG, Wenjin LI, Liang YAN, Yulei LI. MiR-6838-5p overexpression inhibits proliferation of breast cancer MCF-7 cells by downregulating DDR1 expression [J]. Journal of Southern Medical University, 2024, 44(9): 1677-1684. |
| [4] | Yidan PANG, Ya LIU, Siai CHEN, Jinglei ZHANG, Jin ZENG, Yuanming PAN, Juan AN. Biological role of SPAG5 in the malignant proliferation of gastric cancer cells [J]. Journal of Southern Medical University, 2024, 44(8): 1497-1507. |
| [5] | Xiaofan CONG, Teng CHEN, Shuo LI, Yuanyuan WANG, Longyun ZHOU, Xiaolong LI, Pei ZHANG, Xiaojin SUN, Surong ZHAO. Dihydroartemisinin enhances sensitivity of nasopharyngeal carcinoma HNE1/DDP cells to cisplatin-induced apoptosis by promoting ROS production [J]. Journal of Southern Medical University, 2024, 44(8): 1553-1560. |
| [6] | Yuanyuan WANG, Teng CHEN, Xiaofan CONG, Yiran LI, Rui CHEN, Pei ZHANG, Xiaojin SUN, Surong ZHAO. Pristimerin enhances cisplatin-induced apoptosis in nasopharyngeal carcinoma cells via ROS-mediated deactivation of the PI3K/AKT signaling pathway [J]. Journal of Southern Medical University, 2024, 44(5): 904-912. |
| [7] | SHEN Mengdi, ZHAO Na, DENG Xiaojing, DENG Min. High expression of COX6B2 in gastric cancer is associated with poor long-term prognosis and promotes cell proliferation and cell cycle progression by inhibiting p53 signaling [J]. Journal of Southern Medical University, 2024, 44(2): 289-297. |
| [8] | Yue HU, Yu ZENG, Linjing WANG, Zhiwei LIAO, Jianming TAN, Yanhao KUANG, Pan GONG, Bin QI, Xin ZHEN. Performance of multi-modality and multi-classifier fusion models for predicting radiation-induced oral mucositis in patients with nasopharyngeal carcinoma [J]. Journal of Southern Medical University, 2024, 44(12): 2434-2442. |
| [9] | Peipei ZHAO, Zhigang ZHOU, Yuanyuan YANG, Shusheng HUANG, Yixuan TU, Jian TU. Ferroptosis inducer Erastin inhibits proliferation of liver cancer cells in vitro by down-regulating ACSL4 [J]. Journal of Southern Medical University, 2024, 44(11): 2131-2136. |
| [10] | CHENG Yang, HE Xuxu, WANG Lian, XU Yibo, SHEN Mengdi, ZHANG Wenjing, XIA Yongsheng, ZHANG Jie, ZHANG Min, WANG Yijun, HU Jianguo, ZHANG Jun. HSDL2 overexpression promotes rectal cancer progression by regulating cancer cell cycle and promoting cell proliferation [J]. Journal of Southern Medical University, 2023, 43(4): 544-551. |
| [11] | ZHANG Haoxuan, LU Jin, JIANG Chengyi, FANG Meifang. Construction and evaluation of an artificial intelligence-based risk prediction model for death in patients with nasopharyngeal cancer [J]. Journal of Southern Medical University, 2023, 43(2): 271-279. |
| [12] | WANG Shaoxin, CUI Lihong, LI Hui, LIU Xinyao, LI Xiaowei, WANG Xiaohui. UBE2W overexpression promotes proliferation of intestinal mucosal cells in mice with chemically induced colitis [J]. Journal of Southern Medical University, 2023, 43(12): 2023-2028. |
| [13] | HU Tong, GOU Wenfeng, REN Zhonghao, LIU Gaiting, LI Yiliang, ZUO Daiying, HOU Wenbin. Icaritin increases radiosensitivity of nasopharyngeal carcinoma cells by regulating iron death [J]. Journal of Southern Medical University, 2023, 43(10): 1665-1673. |
| [14] | FAN Xiaoli, WANG Juan, WANG Liming. Licochalcone A induces cell cycle arrest in human lung squamous carcinoma cells via the PI3K/Akt signaling pathway [J]. Journal of Southern Medical University, 2023, 43(1): 111-116. |
| [15] | LIN Jie, OU Huohui, WANG Weidong, MA Jing, ZHANG Weijie, LIU Qingbo. Overexpression of CLEC5A inhibits cell proliferation and metastasis and reverses epithelial-mesenchymal transition in hepatocellular carcinoma [J]. Journal of Southern Medical University, 2023, 43(1): 85-91. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||