南方医科大学学报 ›› 2025, Vol. 45 ›› Issue (5): 1082-1092.doi: 10.12122/j.issn.1673-4254.2025.05.22
收稿日期:2024-11-04
出版日期:2025-05-20
发布日期:2025-05-23
通讯作者:
周小龙
E-mail:sqli228@stu.ahjzu.edu.cn;xiaolong@ieee.org
作者简介:李苏强,在读硕士研究生,E-mail: sqli228@stu.ahjzu.edu.cn
基金资助:
Suqiang LI1( ), Zhouyang WANG2, Sixian CHAN2, Xiaolong ZHOU3(
), Zhouyang WANG2, Sixian CHAN2, Xiaolong ZHOU3( )
)
Received:2024-11-04
Online:2025-05-20
Published:2025-05-23
Contact:
Xiaolong ZHOU
E-mail:sqli228@stu.ahjzu.edu.cn;xiaolong@ieee.org
Supported by:摘要:
目的 提出一种基于双向稠密连接和注意力机制的多尺度颌骨囊肿分割模型(AConvLSTM U-Net),实现颌骨囊肿图像的准确自动分割。 方法 使用含有2592张颌骨囊肿图像数据集。首先,AConvLSTM U-Net在编码路径上设计移动翻转瓶颈卷积模块(MBC)以增强特征提取能力。其次,采用双路径稠密卷积(DPD)连接编码器和解码器,在跳跃连接中引入双向ConvLSTM以获取丰富的语义信息。然后,解码路径上使用基于空间和通道注意力的解码块(scSE),以提升对重要信息的关注。最后,设计了全尺寸深度监督模块(DS),并结合联合损失函数对模型进行优化,以进一步提高分割精度。 结果 AConvLSTM U-Net在颌骨囊肿病灶分割的实验结果在MCC、DSC和JSC方面分别达到93.8443%、93.9067%、88.5133%,性能均优于所有被比较的分割模型。 结论 所提出的算法在颌骨囊肿数据集上表现出较高的准确性与鲁棒性,优于多种主流方法,展现了AConvLSTM U-Net在颌骨囊肿图像分割的优越性能和辅助诊断的巨大潜力。
李苏强, 王周阳, 产思贤, 周小龙. AConvLSTM U-Net:基于双向稠密连接和注意力机制的多尺度颌骨囊肿分割模型[J]. 南方医科大学学报, 2025, 45(5): 1082-1092.
Suqiang LI, Zhouyang WANG, Sixian CHAN, Xiaolong ZHOU. AConvLSTM U-Net: a multi-scale jaw cyst segmentation model based on bidirectional dense connection and attention mechanism[J]. Journal of Southern Medical University, 2025, 45(5): 1082-1092.
| Method | DSC | MCC | JSC |
|---|---|---|---|
| Unet | 86.842% | 86.942% | 76.745% |
| BCDUNet | 89.889% | 90.090% | 83.985% |
| MedSAM | 91.870% | 92.046% | 85.806% |
| DS-TransUnet | 92.832% | 92.900% | 87.154% |
| H2Former | 92.390% | 92.540% | 86.440% |
| SwinUnet | 89.683% | 89.827% | 82.686% |
| AConvLSTM U-Net | 93.039% | 92.968% | 87.183% |
表1 在JCMI数据集上与主流医学图像分割方法对比
Tab.1 Comparison with mainstream methods for medical image segmentation on the JCMI dataset
| Method | DSC | MCC | JSC |
|---|---|---|---|
| Unet | 86.842% | 86.942% | 76.745% |
| BCDUNet | 89.889% | 90.090% | 83.985% |
| MedSAM | 91.870% | 92.046% | 85.806% |
| DS-TransUnet | 92.832% | 92.900% | 87.154% |
| H2Former | 92.390% | 92.540% | 86.440% |
| SwinUnet | 89.683% | 89.827% | 82.686% |
| AConvLSTM U-Net | 93.039% | 92.968% | 87.183% |
| Methods | F1 | IoU |
|---|---|---|
| Unet | 79.37 | 66.95 |
| Unet++ | 77.54 | 64.33 |
| ResUNet | 78.25 | 64.89 |
| MedT | 76.93 | 63.89 |
| TransUNet | 79.30 | 66.92 |
| UNeXt | 79.37 | 66.95 |
| AAU-net | - | 64.26 |
| AConvLSTM U-Net | 78.87 | 65.11 |
表2 在BUSI数据集上的定性比较
Tab.2 Qualitative comparison on the BUSI dataset
| Methods | F1 | IoU |
|---|---|---|
| Unet | 79.37 | 66.95 |
| Unet++ | 77.54 | 64.33 |
| ResUNet | 78.25 | 64.89 |
| MedT | 76.93 | 63.89 |
| TransUNet | 79.30 | 66.92 |
| UNeXt | 79.37 | 66.95 |
| AAU-net | - | 64.26 |
| AConvLSTM U-Net | 78.87 | 65.11 |
| Methods | DSC | MCC | JSC | Params |
|---|---|---|---|---|
| Baseline | 89.889% | 90.090% | 83.985% | 23.03M |
| Baseline+MBC | 90.697% | 90.876% | 85.043% | 22.11M |
| Baseline+MBC+scSE | 90.792% | 90.732% | 83.136% | 22.28M |
| Baseline+MBC+DPD | 92.247% | 92.172% | 85.610% | 17.95M |
| Baseline+MBC+DPD+scSE | 92.399% | 92.322% | 85.872% | 18.10M |
| AConvLSTM U-Net | 93.039% | 92.968% | 87.183% | 18.12M |
表3 模块有效性验证结果
Tab.3 Module effectiveness verification results
| Methods | DSC | MCC | JSC | Params |
|---|---|---|---|---|
| Baseline | 89.889% | 90.090% | 83.985% | 23.03M |
| Baseline+MBC | 90.697% | 90.876% | 85.043% | 22.11M |
| Baseline+MBC+scSE | 90.792% | 90.732% | 83.136% | 22.28M |
| Baseline+MBC+DPD | 92.247% | 92.172% | 85.610% | 17.95M |
| Baseline+MBC+DPD+scSE | 92.399% | 92.322% | 85.872% | 18.10M |
| AConvLSTM U-Net | 93.039% | 92.968% | 87.183% | 18.12M |
| Methods | Loss | DSC | MCC | JSC | |
|---|---|---|---|---|---|
| LossBCE | LossDice | ||||
| Baseline | √ | 89.889% | 90.090% | 83.985% | |
| √ | 0.000% | 0.000% | 0.000% | ||
| √ | √ | 90.355% | 90.284% | 84.759% | |
| Ours | √ | 93.039% | 92.968% | 87.1832% | |
| √ | 2.008% | 0.000% | 1.014% | ||
| √ | √ | 93.907% | 93.844% | 88.513% | |
表4 联合损失函数有效性对比结果
Tab.4 Evaluation of the effectiveness of the combined loss function
| Methods | Loss | DSC | MCC | JSC | |
|---|---|---|---|---|---|
| LossBCE | LossDice | ||||
| Baseline | √ | 89.889% | 90.090% | 83.985% | |
| √ | 0.000% | 0.000% | 0.000% | ||
| √ | √ | 90.355% | 90.284% | 84.759% | |
| Ours | √ | 93.039% | 92.968% | 87.1832% | |
| √ | 2.008% | 0.000% | 1.014% | ||
| √ | √ | 93.907% | 93.844% | 88.513% | |
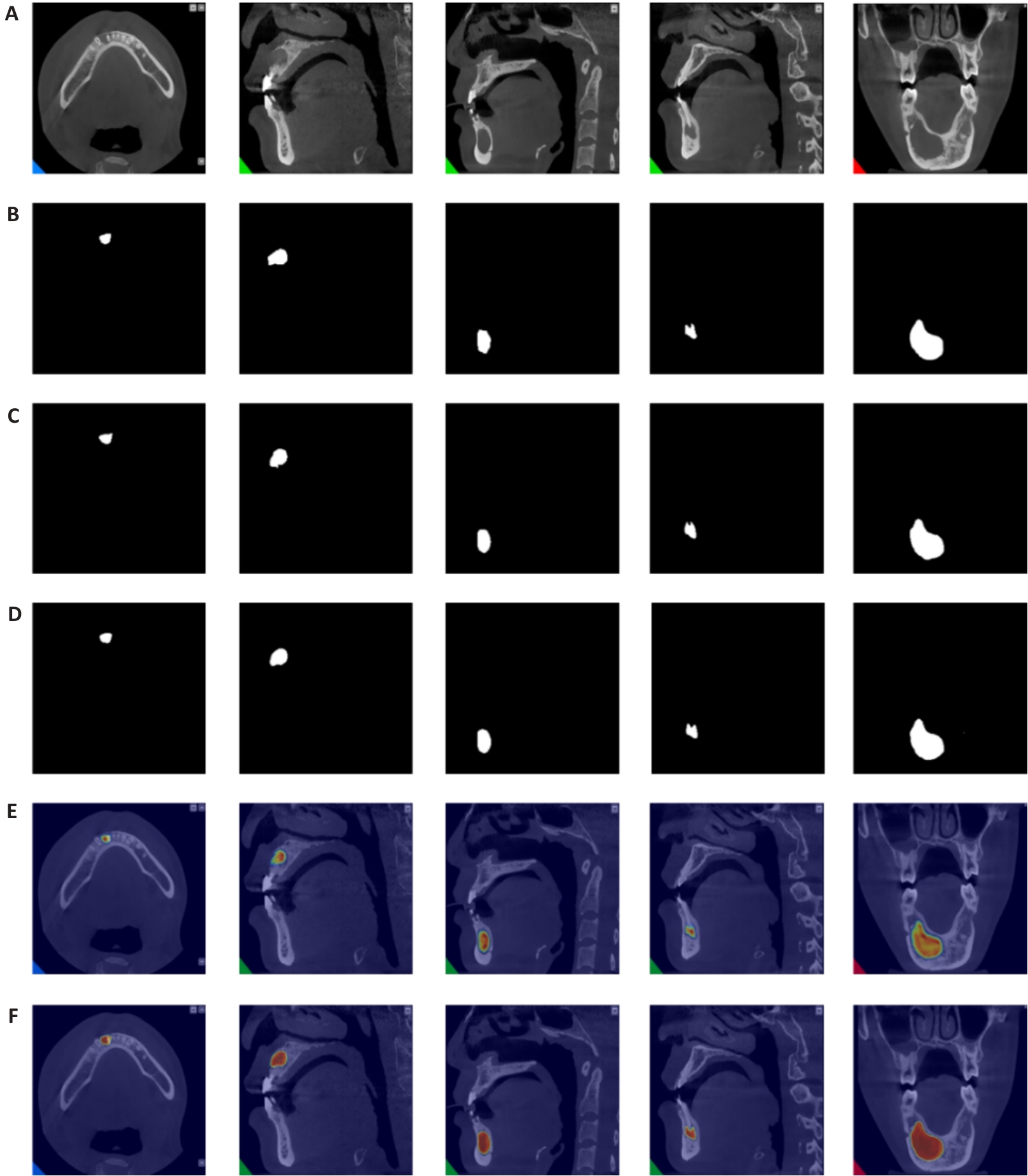
图9 颌骨囊肿可视化结果对比
Fig.9 Visual comparison of jaw cysts detection results. A: Original CT image of jaw cyst. B: Ground truth segmentation label image. C: Segmentation result by BCDUNet model. D: Segmentation result by AConvLSTM U-Net. E: Heatmap of segmentation result by the baseline model. F: Heatmap of segmentation result by AConvLSTM U-Net.
| Test set | DSC | MCC | JSC |
|---|---|---|---|
| Subset 1 | 95.461% | 95.174% | 91.316% |
| Subset 2 | 93.771% | 93.395% | 88.272% |
| Subset 3 | 91.275% | 90.805% | 83.950% |
| Subset 4 | 90.688% | 90.201% | 82.963% |
| Subset 5 | 95.358% | 95.160% | 91.127% |
| Average value | 93.311% | 92.947% | 87.526% |
表5 在BUSI 数据集上的五重交叉验证结果
Tab.5 Five-fold cross-validation results on the BUSI dataset
| Test set | DSC | MCC | JSC |
|---|---|---|---|
| Subset 1 | 95.461% | 95.174% | 91.316% |
| Subset 2 | 93.771% | 93.395% | 88.272% |
| Subset 3 | 91.275% | 90.805% | 83.950% |
| Subset 4 | 90.688% | 90.201% | 82.963% |
| Subset 5 | 95.358% | 95.160% | 91.127% |
| Average value | 93.311% | 92.947% | 87.526% |
| Learning rate | DSC | MCC | JSC |
|---|---|---|---|
| 1×10-2 | 0.000% | 0.000% | 0.000% |
| 1×10-3 | 77.962% | 77.220% | 63.884% |
| 1×10-4 | 78.871% | 78.134% | 65.114% |
表6 学习率对模型精确度
Tab.6 The effect of learning rate on model accuracy
| Learning rate | DSC | MCC | JSC |
|---|---|---|---|
| 1×10-2 | 0.000% | 0.000% | 0.000% |
| 1×10-3 | 77.962% | 77.220% | 63.884% |
| 1×10-4 | 78.871% | 78.134% | 65.114% |
| Method | DSC | IoU |
|---|---|---|
| Unet | 85.75% | 75.05% |
| Unet++ | 87.76% | 78.19% |
| SwinUnet | 86.45% | 76.13% |
| Polyp-SAM | 92.00% | 87.00% |
| Polyp-SAM++ | 91.00% | 86.00% |
| TASPP-Unet | 89.67% | 87.89% |
| AConvLSTM U-Net | 93.03% | 86.43% |
表7 基准数据集CVC-ClinicDB上的比较
Tab.7 Quantitative evaluation on the benchmark CVC-ClinicDB dataset
| Method | DSC | IoU |
|---|---|---|
| Unet | 85.75% | 75.05% |
| Unet++ | 87.76% | 78.19% |
| SwinUnet | 86.45% | 76.13% |
| Polyp-SAM | 92.00% | 87.00% |
| Polyp-SAM++ | 91.00% | 86.00% |
| TASPP-Unet | 89.67% | 87.89% |
| AConvLSTM U-Net | 93.03% | 86.43% |
| 1 | Ding X, Jiang X, Zheng H, et al. MARes-Net: multi-scale attention residual network for jaw cyst image segmentation[J]. Front Bioeng Biotechnol, 2024, 12: 1454728. |
| 2 | Niu G, Zhang G, Chen JM, et al. A 3-year follow-up clinical study on the preservation for vitality of involved tooth in jaw cysts through an innovative method[J]. Sci Rep, 2024, 14(1): 128. |
| 3 | Zheng HX, Jiang XL, Xu X, et al. MFI-net: multi-level feature integration network with SE-Res2Conv encoder for jaw cyst segmentation[J]. IEEE Access, 2024, 12: 67355-67. |
| 4 | Li H, Liu Z, Jiang L, et al. Epidemiological analysis of the clinicopathologic characteristics, treatment, and prognosis of 2648 jaw cysts in West China[J]. Chin Med J: Engl, 2024, 137(9): 1124-6. |
| 5 | Tran T P, Ngoc V T N, Linh L N P, et al. Effectiveness of marsupialization on the reduction of cystic jaw lesions in children: A scoping review[J]. Oral Science International, 2025, 22(1): e1267. |
| 6 | Jiang X, Zheng H, Yuan Z, et al. HIMS-Net: Horizontal-vertical interaction and multiple side-outputs network for cyst segmentation in jaw images[J]. Math Biosci Eng, 2024, 21(3): 4036-55. |
| 7 | Guangyan W, Yanan J, Aihemaiti G, et al. Research on Cyst of Jaw Detection Algorithm Based on Alex Net Deep Learning Model. Research Square[J]. 2024. |
| 8 | 梁利渡, 张浩杰, 鲁 倩, 等. aFaster RCNN: 一种基于平扫 CT 的多疾病阶段胰腺病灶检测模型[J]. 南方医科大学学报, 2023, 43(5): 755. DOI: 10.12122/j.issn.1673-4254.2023.05.11 |
| 9 | 吴雪扬, 张 煜, 张 华, 等. 基于注意力机制和多模态特征融合的猕猴脑磁共振图像全脑分割[J]. 南方医科大学学报, 2023, 43(12): 2118. DOI: 10.12122/j.issn.1673-4254.2023.12.17 |
| 10 | 黄品瑜, 钟丽明, 郑楷宜, 等. 多期相 CT 合成辅助的腹部多器官图像分割[J].南方医科大学学报, 2024, 44(1): 83. DOI: 10.12122/j.issn.1673-4254.2024.01.10 |
| 11 | Pham DL, Xu C, Prince JL. Current methods in medical image segmentation[J]. Annu Rev Biomed Eng, 2000, 2: 315-37. |
| 12 | Varga-Szemes A, Muscogiuri G, Schoepf UJ, et al. Clinical feasibility of a myocardial signal intensity threshold-based semi-automated cardiac magnetic resonance segmentation method[J]. Eur Radiol, 2016, 26(5): 1503-11. |
| 13 | Sankur B. Survey over image thresholding techniques and quantitative performance evaluation[J]. J Electron Imaging, 2004, 13(1): 146. |
| 14 | Otsu N. A threshold selection method from gray-level histograms[J]. IEEE Trans Syst, Man, Cybern, 9(1): 62-6. |
| 15 | Kapur JN, Sahoo PK, Wong AKC. A new method for gray-level picture thresholding using the entropy of the histogram[J]. Comput Vis Graph Image Process, 1985, 29(3): 273-85. |
| 16 | Adams R, Bischof L. Seeded region growing[J]. IEEE Trans Pattern Anal Machine Intell, 16(6): 641-7. |
| 17 | Mehnert A, Jackway P. An improved seeded region growing algorithm[J]. Pattern Recognit Lett, 1997, 18(10): 1065-71. |
| 18 | Fan J, Yau DY, Elmagarmid AK, et al. Automatic image segmentation by integrating color-edge extraction and seeded region growing[J]. IEEE Trans Image Process, 2001, 10(10): 1454-66. |
| 19 | Bezdek JC, Ehrlich R, Full W. FCM: The fuzzy c-means clustering algorithm[J]. Comput Geosci, 1984, 10(2/3): 191-203. |
| 20 | Ng HP, Ong SH, Foong KWC, et al. Medical image segmentation using K-means clustering and improved watershed algorithm[C]// IEEE Southwest Symposium on Image Analysis and Interpretation. Denver, CO. IEEE, 2006, 61-65. |
| 21 | Sulaiman S, Mat Isa N. Adaptive fuzzy-K-means clustering algorithm for image segmentation[J]. IEEE Trans Consumer Electron, 56(4): 2661-8. |
| 22 | Cumani A. Edge detection in multispectral images[J]. CVGIP Graph Models Image Process, 1991, 53(1): 40-51. |
| 23 | Canny J. A computational approach to edge detection[J]. IEEE Trans.Pattern Anal. Mach. Intell, 1986, 8(6): 679-98. |
| 24 | Long J, Shelhamer E, Darrell T. Fully convolutional networks for semantic segmentation[C]//2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). June 7-12, 2015. Boston, MA, USA. IEEE, 2015: 3431-40. |
| 25 | Ronneberger O, Fischer P, Brox T. U-Net: convolutional networks for biomedical image segmentation[C]//Medical Image Computing and Computer-Assisted Intervention (MICCAI). Cham: Springer International Publishing, 2015: 234-41. |
| 26 | Ozturk B, Taspinar YS, Koklu M, et al. Automatic segmentation of the maxillary sinus on cone beam computed tomographic images with U-Net deep learning model[J]. Eur Arch Oto Rhino Laryngol, 2024, 281(11): 6111-21. |
| 27 | Xu L, Qiu K, Li K, et al. Automatic segmentation of ameloblastoma on ct images using deep learning with limited data[J]. BMC Oral Health, 2024, 24(1): 55. |
| 28 | Su S, Jia X, Zhan L, et al. Automatic tooth periodontal ligament segmentation of cone beam computed tomography based on instance segmentation network[J]. Heliyon, 2024, 10(2): e24097. |
| 29 | Azad R, Asadi-Aghbolaghi M, Fathy M, et al. Bi-directional ConvLSTM U-Net with densley connected convolutions[C]//2019 IEEE/CVF International Conference on Computer Vision Workshop (ICCVW). October 27-28, 2019. Seoul, Korea (South). IEEE, 2019. |
| 30 | Huang G, Liu Z, Van Der Maaten L, et al. Densely connected convolutional networks[C]//2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). July 21-26, 2017. Honolulu, HI. IEEE, 2017: 4700-4708. |
| 31 | Sønderby SK, Sønderby CK, Nielsen H, et al. Convolutional LSTM networks for subcellular localization of proteins[C]//Algorithms for Computational Biology. Cham: Springer International Publishing, 2015: 68-80. |
| 32 | Tan MX, Le QV. Efficientnet: Rethinking model scaling for convolutional neural networks[C]//International conference on machine learning (ICML). 2019: 6105-14. |
| 33 | Chollet F. Xception: deep learning with depthwise separable convolutions[C]//2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). July 21-26, 2017. Honolulu, HI. IEEE, 2017: 1251-1258. |
| 34 | Hu J, Shen L, Sun G. Squeeze-and-excitation networks[C]//2018 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). June 18-23, 2018. Salt Lake City, UT. IEEE, 2018: 7132-41. |
| 35 | Ioffe S, Szegedy C, Paranhos L, et al. Batch normalization: Accelerating deep network training by reducing internal covariate shift[C]//International conference on machine learning (ICML). 2015: 448-56. |
| 36 | Ramachandran P, Zoph B, Le QV. Searching for activation functions[J]. arXiv preprint arXiv:, 2017. |
| 37 | Szegedy C, Liu W, Jia YQ, et al. Going deeper with convolutions[C]//2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). June 7-12, 2015. Boston, MA, USA. IEEE, 2015: 1-9. |
| 38 | Mao AQ, Mohri M, Zhong YT. Cross-entropy loss functions: theoretical analysis and applications[C]// International Conference on Machine Learning (ICML). Honolulu, Hawaii, USA, 2023, 992(26): 23803-28. |
| 39 | Li XY, Sun XF, Meng YX, et al. Dice loss for data-imbalanced NLP tasks[J]. 2019: 1911.02855. . |
| 40 | Al-Dhabyani W, Gomaa M, Khaled H, et al. Dataset of breast ultrasound images[J]. Data Brief, 2020, 28: 104863. |
| 41 | Jiang M, Zhai FH, Kong J. A novel deep learning model DDU-net using edge features to enhance brain tumor segmentation on MR images[J]. Artif Intell Med, 2021, 121: 102180. |
| 42 | Yang YY, Feng C, Wang RF. Automatic segmentation model combining U-Net and level set method for medical images[J]. Expert Syst Appl, 2020, 153: 113419. |
| 43 | Zhao C, Shuai RJ, Ma L, et al. Segmentation of dermoscopy images based on deformable 3D convolution and ResU-NeXt++[J]. Med Biol Eng Comput, 2021, 59(9): 1815-32. |
| 44 | Ma J, He Y, Li F, et al. Segment anything in medical images[J]. Nat Commun, 2024, 15(1): 654. |
| 45 | Lin AL, Chen BZ, Xu JY, et al. DS-TransUNet: dual swin transformer U-Net for medical image segmentation[J]. IEEE Trans Instrum Meas, 2022, 71: 1-15. |
| 46 | He A, Wang K, Li T, et al. H2Former: an efficient hierarchical hybrid transformer for medical image segmentation[J]. IEEE Trans Med Imaging, 2023, 42(9): 2763-75. |
| 47 | Cao H, Wang YY, Chen J, et al. Swin-unet: Unet-like pure transformer for medical image segmentation[C]//European conference on computer vision (ECCV). Cham: Springer Nature Switzerland, 2022: 205-18. |
| 48 | Zhou ZW, Rahman Siddiquee MM, Tajbakhsh N, et al. UNet++: A nested U-Net architecture for medical image segmentation[C]//Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support. Cham: Springer International Publishing, 2018: 3-11. |
| 49 | Zhang ZX, Liu QJ, Wang YH. Road extraction by deep residual U-net[J]. IEEE Geosci Remote Sensing Lett, 15(5): 749-53. |
| 50 | Valanarasu JMJ, Oza P, Hacihaliloglu I, et al. Medical transformer: gated axial-attention for medical image segmentation[C]//Medical Image Computing and Computer Assisted Intervention (MICCAI). Cham: Springer International Publishing, 2021: 36-46. |
| 51 | Chen JN, Lu YY, Yu QH, et al. TransUNet: transformers make strong encoders for medical image segmentation[J]. 2021: 2102. 04306. . |
| 52 | Valanarasu JMJ, Patel VM. UNeXt: MLP-based rapid medical image segmentation network[C]//Medical Image Computing and Computer Assisted Intervention (MICCAI). Cham: Springer Nature Switzerland, 2022: 23-33. |
| 53 | Chen G, Li L, Dai Y, et al. AAU-Net: an adaptive attention U-Net for breast lesions segmentation in ultrasound images[J]. IEEE Trans Med Imaging, 2023, 42(5): 1289-300. |
| 54 | Selvaraju RR, Cogswell M, Das A, et al. Grad-CAM: visual explanations from deep networks via gradient-based localization[C]//2017 IEEE International Conference on Computer Vision (ICCV). October 22-29, 2017. Venice. IEEE, 2017: 618-26. |
| [1] | 郑子瑜, 杨夏颖, 吴圣杰, 张诗婕, 吕国荣, 柳培忠, 王珺, 何韶铮. 多特征融合的产时超声胎方位识别模型[J]. 南方医科大学学报, 2025, 45(7): 1563-1570. |
| [2] | 计寰宇, 王蕊, 高盛祥, 车文刚. SG-UNet:基于全局注意力和自校准卷积增强的黑色素瘤分割模型[J]. 南方医科大学学报, 2025, 45(6): 1317-1326. |
| [3] | 任煜瀛, 黄凌霄, 杜方, 姚新波. 基于改进RT-DETR的多尺度特征融合的高效轻量皮肤病理检测方法[J]. 南方医科大学学报, 2025, 45(2): 409-421. |
| [4] | 巩 高, 曹 石, 肖 慧, 方威扬, 阙与清, 刘子蔚, 陈超敏. 深度注意力机制结合临床特征预测肝细胞癌微血管浸润[J]. 南方医科大学学报, 2023, 43(5): 839-851. |
| [5] | 吴雪扬, 张 煜, 张 华, 钟 涛. 基于注意力机制和多模态特征融合的猕猴脑磁共振图像全脑分割[J]. 南方医科大学学报, 2023, 43(12): 2118-2125. |
| [6] | 邹青清, 王梦虹, 陆紫箫, 赵英华, 冯前进. 基于多序列MRI的3D关系注意力网络预测HLA-B27阴性中轴性脊柱关节病[J]. 南方医科大学学报, 2023, 43(11): 1955-1964. |
| [7] | 钟友闻, 车文刚, 高盛祥. 轻型多尺度黑色素瘤目标检测网络模型的建立:基于注意力机制调控[J]. 南方医科大学学报, 2022, 42(11): 1662-1671. |
| [8] | 张晓玥, 王永雄, 张佳鹏, 孙洪鑫, 王 东, 陈 羽, 周 志. 基于激活层前置压缩激励残差网络的早期胃癌筛查算法[J]. 南方医科大学学报, 2021, 41(11): 1616-1622. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||