南方医科大学学报 ›› 2026, Vol. 46 ›› Issue (3): 693-706.doi: 10.12122/j.issn.1673-4254.2026.03.23
• 基础研究 • 上一篇
向科霖1( ), 章晓云2, 黎征鹏1, 徐志为2, 刘素杰1, 柴源2,3(
), 章晓云2, 黎征鹏1, 徐志为2, 刘素杰1, 柴源2,3( )
)
收稿日期:2025-08-14
出版日期:2026-03-20
发布日期:2026-03-26
通讯作者:
柴源
E-mail:xiangkelin1024@163.com;chaizxy@163.com
作者简介:向科霖,在读硕士研究生,E-mail: xiangkelin1024@163.com
基金资助:
Kelin XIANG1( ), Xiaoyu ZHANG2, Zhengpeng LI1, Zhiwei XU2, Sujie LIU1, Yuan CHAI2,3(
), Xiaoyu ZHANG2, Zhengpeng LI1, Zhiwei XU2, Sujie LIU1, Yuan CHAI2,3( )
)
Received:2025-08-14
Online:2026-03-20
Published:2026-03-26
Contact:
Yuan CHAI
E-mail:xiangkelin1024@163.com;chaizxy@163.com
摘要:
目的 基于生物信息学与机器学习法,筛选骨关节炎中与胞葬作用相关的关键基因,探讨其诊断价值、免疫微环境特征及潜在中药治疗靶点。 方法 从GEO数据库获取骨关节炎数据集GSE55235、GSE55457和GSE117999,其中GSE55235作为训练集,GSE55457和GSE117999作为验证集。从GeneCards数据库获取胞葬相关基因集。通过差异表达分析筛选骨关节炎差异基因,并与胞葬基因取交集获得胞葬相关差异基因。对差异基因进行GO和KEGG富集分析。采用随机森林、LASSO回归和SVM三种机器学习算法筛选特征基因,并通过ROC曲线评估其诊断效能。通过qRT-PCR实验在大鼠骨关节炎模型中验证特征基因表达,利用CIBERSORT解析免疫细胞浸润情况,采用GSEA分析特征基因相关通路,运用Coremine数据库预测与特征基因相关的中药。 结果 共筛选出959个OA差异基因,其中15个与胞葬作用相关。GO和KEGG分析显示这些基因主要富集于白细胞迁移、细胞外基质、炎症通路等。机器学习筛选出UCP2、EGLN3和IL1B三个特征基因,其在训练集和验证集中均表现出良好的诊断能力。qRT-PCR显示部分特征基因表达趋势存在差异(P<0.05)。免疫浸润分析显示静止肥大细胞、休眠记忆CD4+ T细胞和活化的肥大细胞在骨关节炎患者和健康人群的浸润差异显著(P<0.05)。GSEA提示特征基因与脂肪细胞因子信号通路、硫代谢和剪接体通路密切相关。中药预测获得100味中药,以补虚、清热、活血化瘀类为主,包括枸杞子、淫羊藿、生地黄、苦参、川芎、牛膝等。 结论 胞葬相关基因在骨关节炎发病中起重要作用,UCP2、EGLN3、IL1B基因对骨关节炎具有诊断价值,预测出的枸杞子、淫羊藿、生地黄、苦参、川芎、牛膝等中药可能是防治骨关节炎的潜在药物。
向科霖, 章晓云, 黎征鹏, 徐志为, 刘素杰, 柴源. 胞葬相关基因UCP2、EGLN3、IL1B对骨关节炎的诊断价值及中药治疗预测:基于生物信息学及机器学习[J]. 南方医科大学学报, 2026, 46(3): 693-706.
Kelin XIANG, Xiaoyu ZHANG, Zhengpeng LI, Zhiwei XU, Sujie LIU, Yuan CHAI. Identification of efferocytosis-related genes in osteoarthritis and prediction of traditional Chinese medicines based on bioinformatics and machine learning[J]. Journal of Southern Medical University, 2026, 46(3): 693-706.
| GEO ID | Platform | Control (n) | Case (n) | Group |
|---|---|---|---|---|
| GSE55235 | GPL96 | 10 | 10 | Training set |
| GSE55457 | GPL96 | 10 | 10 | Validation set |
| GSE117999 | GPL20844 | 12 | 12 | Validation set |
表1 GEO数据集的基本信息和分组
Tab.1 Basic information and grouping of GEO datasets
| GEO ID | Platform | Control (n) | Case (n) | Group |
|---|---|---|---|---|
| GSE55235 | GPL96 | 10 | 10 | Training set |
| GSE55457 | GPL96 | 10 | 10 | Validation set |
| GSE117999 | GPL20844 | 12 | 12 | Validation set |
| Gene | Primer sequence (5'-3') |
|---|---|
| UCP2-S | СТСССААТGТТGСССGАААТ |
| UCP2-A | GAAGTGGCAAGGGAGGTCGT |
| EGLN3-S | CGCCAAGTTACATGGAGGGG |
| EGLN3-A | CCAGACAGTCATGGCGTACC |
| IL1B-S | GAACAACAAAAATGCCTCGTGC |
| IL1B-A | GACAAACCGCTTTTCCATCTTCT |
| MMP13-S | TGCATACGAGCATCCATCCC |
| MMP13-A | CGTGTCCTCAAAGTGAACCGC |
| COL2A1-S | CAGACAGTACCTTGAGACAGCATGA |
| COL2A1-A | AGGTGCGAGCGGGATTCTT |
| GAPDH-S | CTGGAGAAACCTGCCAAGTATG |
| GAPDH-A | GGTGGAAGAATGGGAGTTGCT |
表2 PCR引物序列
Tab.2 PCR primer sequences
| Gene | Primer sequence (5'-3') |
|---|---|
| UCP2-S | СТСССААТGТТGСССGАААТ |
| UCP2-A | GAAGTGGCAAGGGAGGTCGT |
| EGLN3-S | CGCCAAGTTACATGGAGGGG |
| EGLN3-A | CCAGACAGTCATGGCGTACC |
| IL1B-S | GAACAACAAAAATGCCTCGTGC |
| IL1B-A | GACAAACCGCTTTTCCATCTTCT |
| MMP13-S | TGCATACGAGCATCCATCCC |
| MMP13-A | CGTGTCCTCAAAGTGAACCGC |
| COL2A1-S | CAGACAGTACCTTGAGACAGCATGA |
| COL2A1-A | AGGTGCGAGCGGGATTCTT |
| GAPDH-S | CTGGAGAAACCTGCCAAGTATG |
| GAPDH-A | GGTGGAAGAATGGGAGTTGCT |
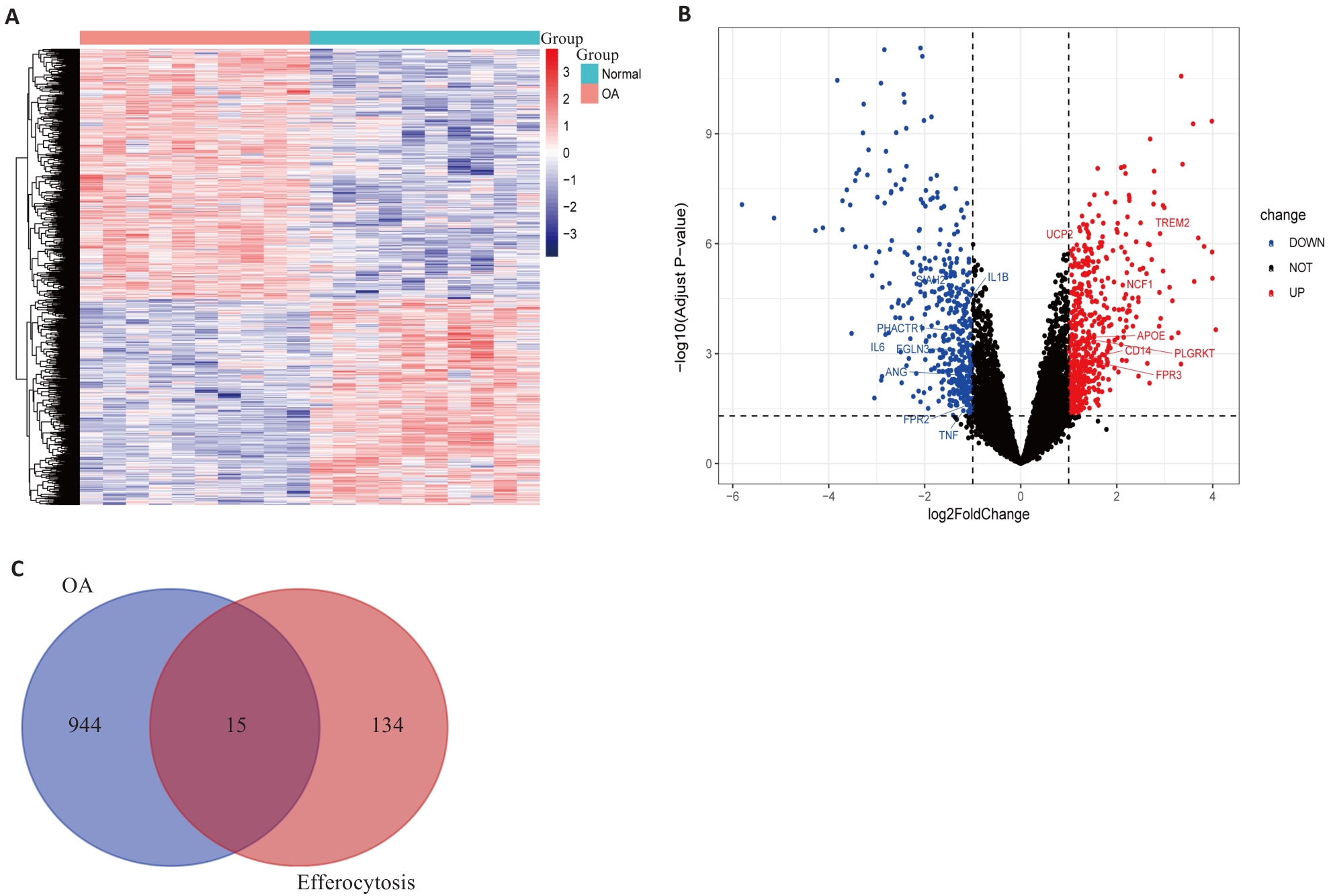
图1 OA患者与健康人群样本间的差异表达基因分析
Fig.1 Differential gene expression analysis between osteoarthritis (OA) patients and healthy controls. A: Gene expression heatmap between OA patients and healthy controls. Red represents up-regulated expression, and blue represents down-regulated expression. B: Volcano plot of differentially expressed genes (DEGs). Black dots indicate genes with no significant differences, red dots indicate significantly up-regulated DEGs, and blue dots indicate significantly down-regulated DEGs. The figure displays efferocytosis-related DEGs. C: Venn diagram of the intersection between DEGs in OA and efferocytosis-related gene sets, obtaining 15 efferocytosis-related DEGs.
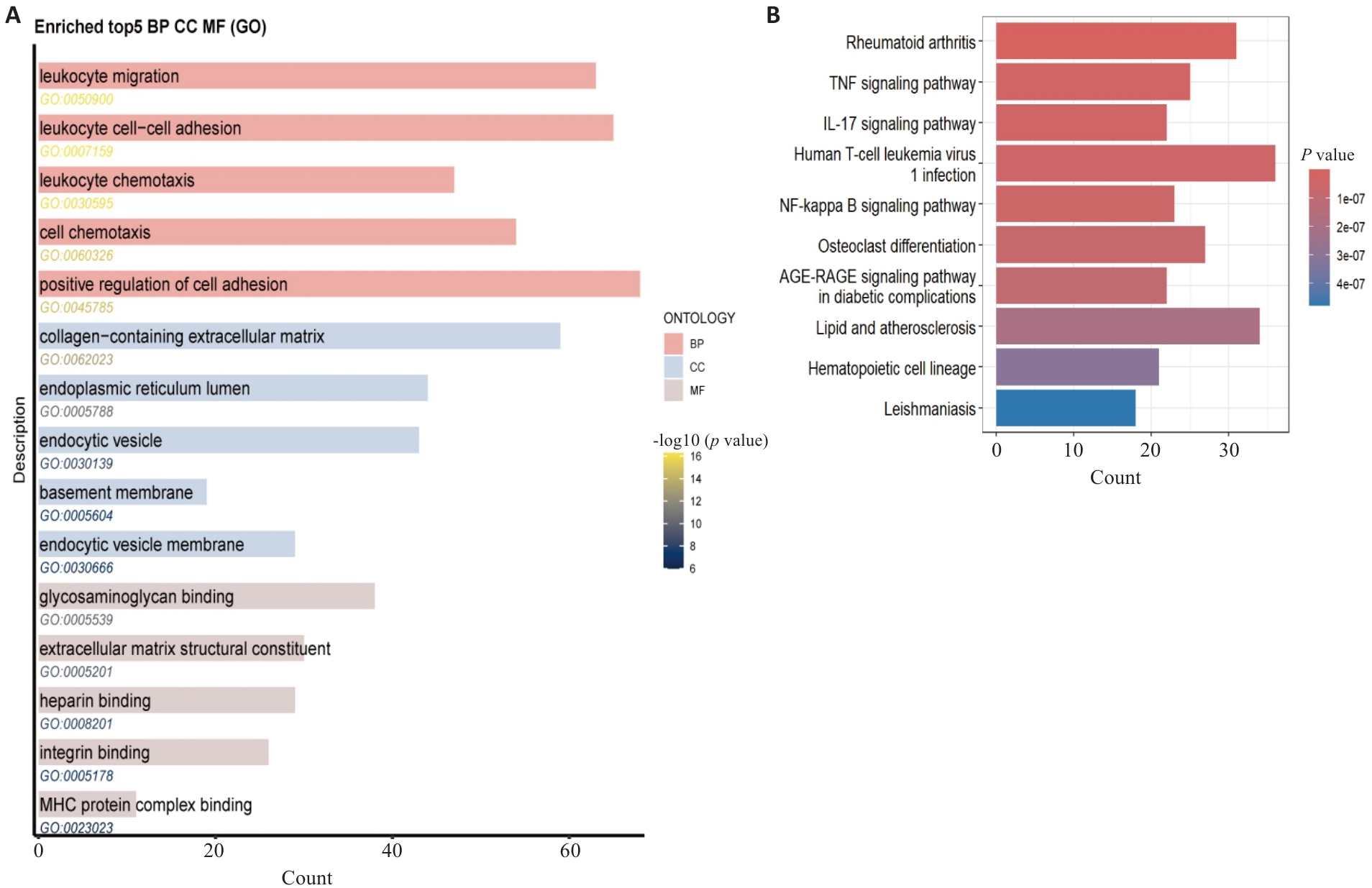
图2 骨关节炎差异基因的GO功能和KEGG通路富集分析
Fig.2 GO functional and KEGG pathway enrichment analysis of osteoarthritis DEGs. A: GO enrichment analysis results. B: KEGG pathway enrichment analysis results, displaying the signaling pathways significantly enriched by the DEGs.
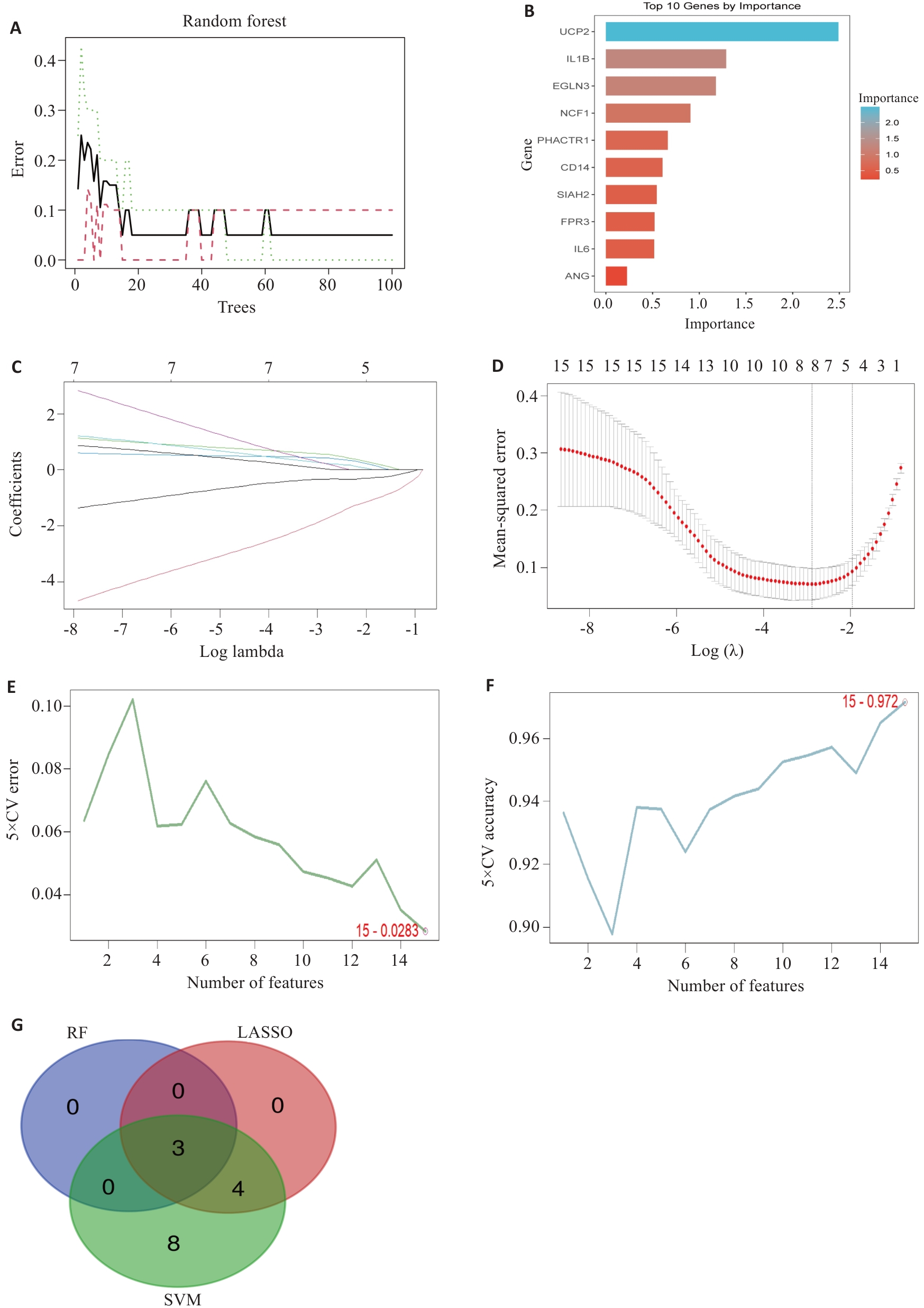
图3 机器学习筛选特征基因
Fig.3 Machine learning screening of feature genes. A, B: RF screening process and results, showing trend of model error with increasing number of decision trees (A) and gene importance ranking (B). C, D: LASSO regression screening process and results showing model coefficient path diagram (C) and trend of cross-validation error with penalty coefficient λ (D). E, F: SVM screening process and results showing model error rate with varying number of features (E) and model accuracy with varying number of features (F). G: Venn diagram of the intersection of screening results from three machine learning algorithms, ultimately obtaining 3 consensus feature genes.
| Dataset | Accuracy | Precision | Recall | F1-Score | AUC |
|---|---|---|---|---|---|
| GSE55235 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 |
| GSE55457 | 0.850 | 0.818 | 0.900 | 0.857 | 0.900 |
| GSE117999 | 0.850 | 0.818 | 0.900 | 0.857 | 0.910 |
表3 基于特征基因的分类模型性能评估
Tab.3 Performance evaluation of classification model based on the feature genes
| Dataset | Accuracy | Precision | Recall | F1-Score | AUC |
|---|---|---|---|---|---|
| GSE55235 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 |
| GSE55457 | 0.850 | 0.818 | 0.900 | 0.857 | 0.900 |
| GSE117999 | 0.850 | 0.818 | 0.900 | 0.857 | 0.910 |
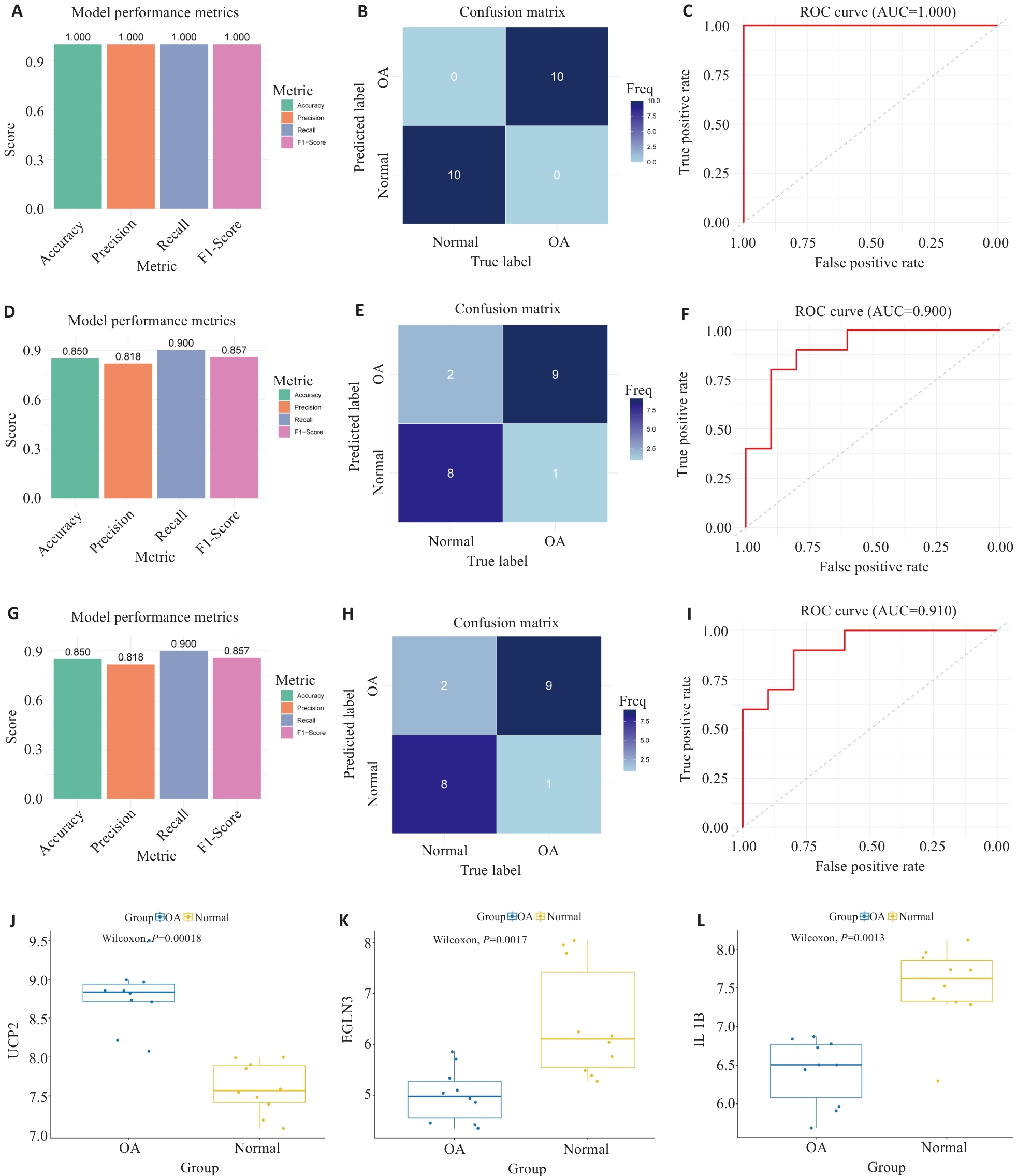
图4 模型性能评估的综合分析
Fig.4 Comprehensive analysis of model performance evaluation. A-C: Performance assessment of the diagnostic model in the training set (GSE55235), showing the accuracy, precision, recall, and F1-score diagram (A), confusion matrix (B), and ROC curve and AUC value (C). D-F: Performance assessment of the diagnostic model in the validation set (GSE55457) showing the accuracy, precision, recall, and F1-score diagram (D), confusion matrix (E), and ROC curve and AUC value (F). G-I: Performance of the diagnostic model in the validation set (GSE117999), showing the accuracy, precision, recall, and F1-score diagram (G), confusion matrix (H), and ROC curve and AUC value (I). J-L: Comparison of expression levels of the 3 feature genes between OA patients and healthy controls (J: UCP2; K: EGLN3; L: IL1B).
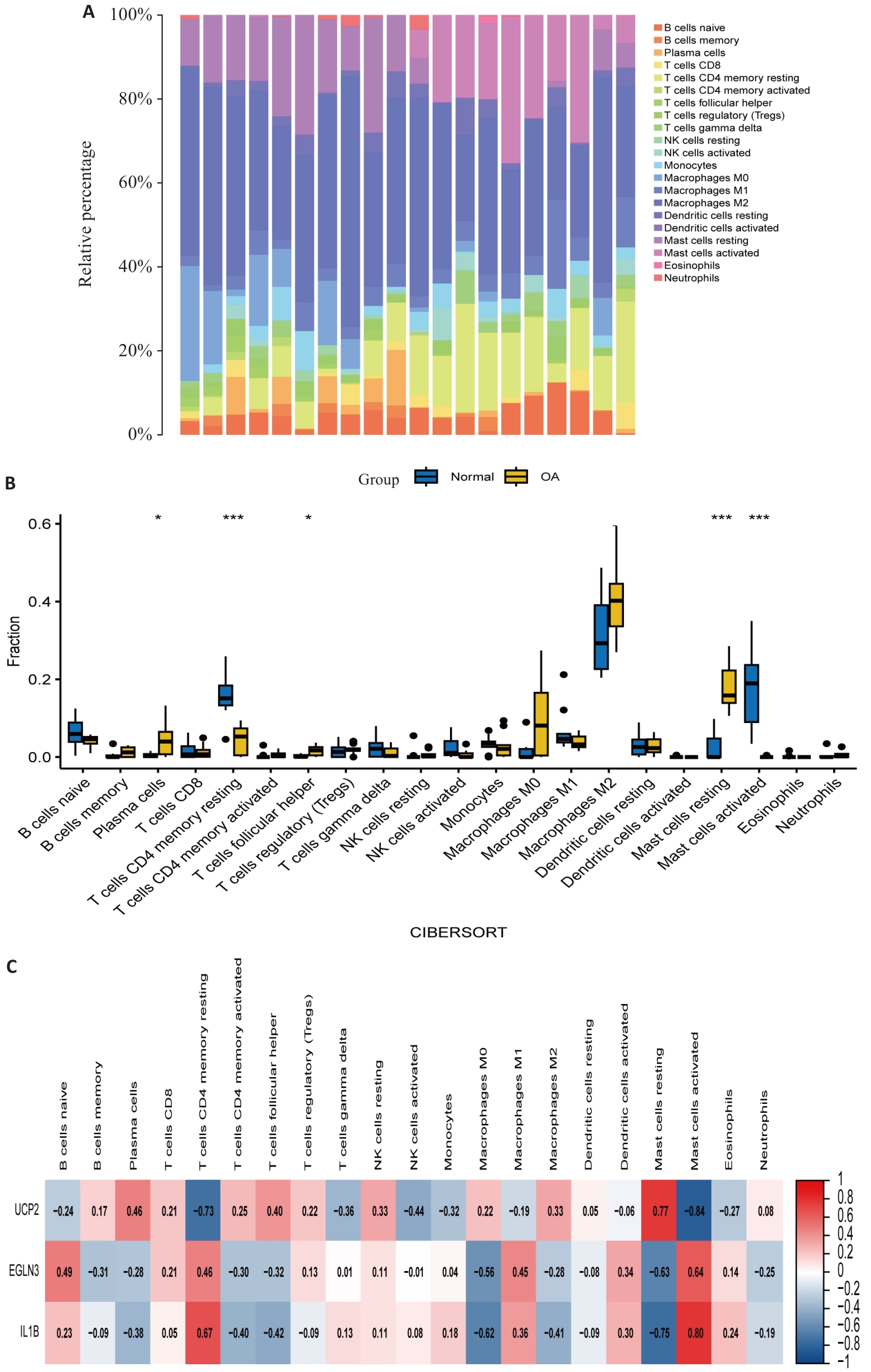
图6 免疫浸润分析图
Fig.6 Immune infiltration analysis. A: Infiltration of immune cells in OA patient samples and healthy control samples, with different colors representing different immune cells. B: Comparison of immune cell infiltration between OA patient samples and healthy control samples. C: Correlation between feature genes and immune cells, with color depth representing the strength of correlation. *P<0.05, ***P<0.001.
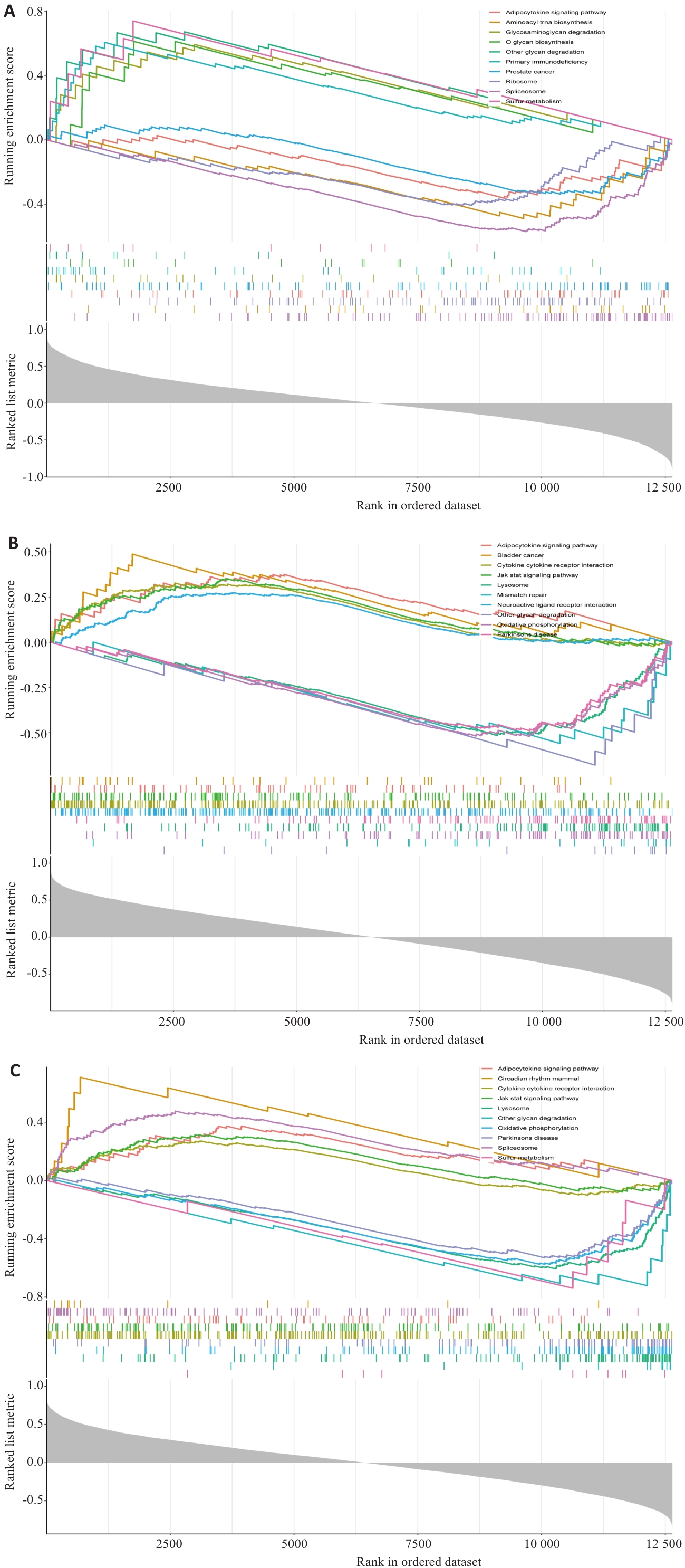
图7 特征基因GSEA富集分析结果
Fig.7 GSEA enrichment analysis results of the feature genes. A: GSEA enrichment analysis results of UCP2. B: GSEA enrichment analysis results of EGLN3. C: GSEA enrichment analysis results of IL1B.
| Category | TCM herbs |
|---|---|
| Exterior-releasing | Saposhnikovia divaricata, Cinnamomum cassia, Ligusticum sinense, Arctium lappa, Centipeda minima, Peucedanum praeruptorum, Angelica dahurica, Ephedra sinica |
| Heat-clearing | Rehmannia glutinosa, Sophora flavescens, Scutellaria baicalensis, Coptis chinensis, Forsythia suspensa, Portulaca oleracea, Lonicera japonica, Houttuynia cordata, Lonicera japonica stem, Arnebia euchroma, Anemarrhena asphodeloides, Morus alba leaf, Phellodendron chinense, Prunella vulgaris, Chrysanthemum indicum, Scutellaria barbata, Terminalia chebula |
| Purgative | Rheum palmatum, Euphorbia lathyris |
| Wind-damp dispelling | Angelica pubescens, Centella asiatica, Dioscorea nipponica, Sinomenium acutum, Clematis chinensis, Gentiana macrophylla, Morus alba twig, Siegesbeckia orientalis, Stephania tetrandra, Chaenomeles speciosa |
| Dampness-resolving | Pogostemon cablin |
| Dampness-draining | Sargassum, Poria cocos, Smilax glabra, Alisma orientale, Akebia quinata |
| Internal-warming | Fresh Ginger Rhizome, Dried Ginger Rhizome, Aconitum carmichaelii, Evodia rutaecarpa, Cnidium monnieri, Cinnamomum cassia |
| Qi-regulating | Magnolia officinalis flower, Magnolia officinalis bark, Inula racemosa, Aucklandia lappa, Citrus aurantium |
| Hemostatic | Artemisia argyi |
| Blood-activating | Ligusticum chuanxiong, Achyranthes bidentata, Salvia miltiorrhiza, Paeonia lactiflora, Carthamus tinctorius, Panax notoginseng, Curcuma longa, Panax notoginseng flower, Crocus sativus, Sargentodoxa cuneata, Caesalpinia sappan, Cyathula officinalis, Anemone raddeana |
| Phlegm-resolving | Ginkgo biloba |
| Tranquilizing | Polygala tenuifolia |
| Wind-extinguishing | Gastrodia elata |
| Tonic | Lycium barbarum, Epimedium brevicornum, Panax ginseng leaf, Panax ginseng, Atractylodes macrocephala, Fish maw, Angelica sinensis, Ganoderma lucidum, Rehmannia glutinosa, Red ginseng, Glycyrrhiza uralensis, Cordyceps sinensis, Eucommia ulmoides, Cornus officinalis, Polygonatum sibiricum, Ophiopogon japonicus, Gynostemma pentaphyllum, Coriolus versicolor, Dioscorea opposita, Houttuynia cordata, Hippophae rhamnoides, Bletilla striata, Agrimonia pilosa, Ligustrum lucidum |
| Astringent | Schisandra chinensis, Rosa laevigata, Punica granatum peel, Ephedra sinica root, Terminalia chebula |
表4 中药预测分类表
Tab.4 Classification of predicted traditional Chinese medicines
| Category | TCM herbs |
|---|---|
| Exterior-releasing | Saposhnikovia divaricata, Cinnamomum cassia, Ligusticum sinense, Arctium lappa, Centipeda minima, Peucedanum praeruptorum, Angelica dahurica, Ephedra sinica |
| Heat-clearing | Rehmannia glutinosa, Sophora flavescens, Scutellaria baicalensis, Coptis chinensis, Forsythia suspensa, Portulaca oleracea, Lonicera japonica, Houttuynia cordata, Lonicera japonica stem, Arnebia euchroma, Anemarrhena asphodeloides, Morus alba leaf, Phellodendron chinense, Prunella vulgaris, Chrysanthemum indicum, Scutellaria barbata, Terminalia chebula |
| Purgative | Rheum palmatum, Euphorbia lathyris |
| Wind-damp dispelling | Angelica pubescens, Centella asiatica, Dioscorea nipponica, Sinomenium acutum, Clematis chinensis, Gentiana macrophylla, Morus alba twig, Siegesbeckia orientalis, Stephania tetrandra, Chaenomeles speciosa |
| Dampness-resolving | Pogostemon cablin |
| Dampness-draining | Sargassum, Poria cocos, Smilax glabra, Alisma orientale, Akebia quinata |
| Internal-warming | Fresh Ginger Rhizome, Dried Ginger Rhizome, Aconitum carmichaelii, Evodia rutaecarpa, Cnidium monnieri, Cinnamomum cassia |
| Qi-regulating | Magnolia officinalis flower, Magnolia officinalis bark, Inula racemosa, Aucklandia lappa, Citrus aurantium |
| Hemostatic | Artemisia argyi |
| Blood-activating | Ligusticum chuanxiong, Achyranthes bidentata, Salvia miltiorrhiza, Paeonia lactiflora, Carthamus tinctorius, Panax notoginseng, Curcuma longa, Panax notoginseng flower, Crocus sativus, Sargentodoxa cuneata, Caesalpinia sappan, Cyathula officinalis, Anemone raddeana |
| Phlegm-resolving | Ginkgo biloba |
| Tranquilizing | Polygala tenuifolia |
| Wind-extinguishing | Gastrodia elata |
| Tonic | Lycium barbarum, Epimedium brevicornum, Panax ginseng leaf, Panax ginseng, Atractylodes macrocephala, Fish maw, Angelica sinensis, Ganoderma lucidum, Rehmannia glutinosa, Red ginseng, Glycyrrhiza uralensis, Cordyceps sinensis, Eucommia ulmoides, Cornus officinalis, Polygonatum sibiricum, Ophiopogon japonicus, Gynostemma pentaphyllum, Coriolus versicolor, Dioscorea opposita, Houttuynia cordata, Hippophae rhamnoides, Bletilla striata, Agrimonia pilosa, Ligustrum lucidum |
| Astringent | Schisandra chinensis, Rosa laevigata, Punica granatum peel, Ephedra sinica root, Terminalia chebula |
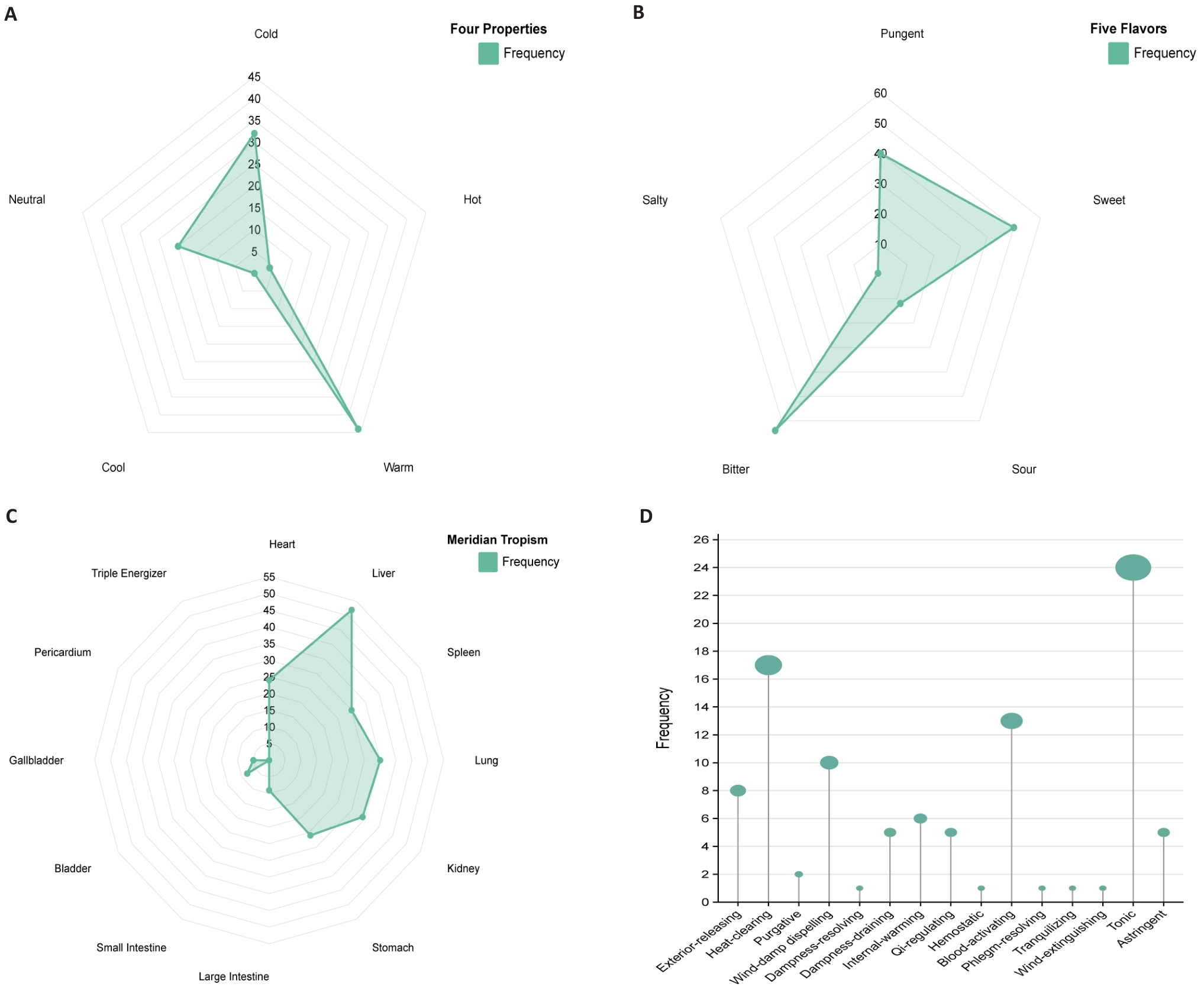
图8 中药预测图
Fig.8 Prediction of traditional Chinese medicines. A: Radar chart of four properties frequency. B: Radar chart of five flavors frequency. C: Radar chart of meridian tropism frequency. D: Frequency diagram of efficacy classification.
| [1] | Hunter DJ, March L, Chew M. Osteoarthritis in 2020 and beyond: a lancet commission[J]. Lancet, 2020, 396(10264): 1711-2. doi:10.1016/s0140-6736(20)32230-3 |
| [2] | Katz JN, Arant KR, Loeser RF. Diagnosis and treatment of hip and knee osteoarthritis: a review[J]. JAMA, 2021, 325(6): 568-78. doi:10.1001/jama.2020.22171 |
| [3] | Richard MJ, Driban JB, McAlindon TE. Pharmaceutical treatment of osteoarthritis[J]. Osteoarthritis Cartilage, 2023, 31(4): 458-66. doi:10.1016/j.joca.2022.11.005 |
| [4] | Del Sordo L, Blackler GB, Philpott HT, et al. Impaired efferocytosis by synovial macrophages in patients with knee osteoarthritis[J]. Arthritis Rheumatol, 2023, 75(5): 685-96. doi:10.1002/art.42412 |
| [5] | Xiong W, Han Z, Ding SL, et al. In situ remodeling of efferocytosis via lesion-localized microspheres to reverse cartilage senescence[J]. Adv Sci: Weinh, 2024, 11(19): e2400345. doi:10.1002/advs.202400345 |
| [6] | Guillem-Llobat P, Marín M, Rouleau M, et al. New insights into the pro-inflammatory and osteoclastogenic profile of circulating monocytes in osteoarthritis patients[J]. Int J Mol Sci, 2024, 25(3): 1710. doi:10.3390/ijms25031710 |
| [7] | Kourtzelis I, Mitroulis I, von Renesse J, et al. From leukocyte recruitment to resolution of inflammation: the cardinal role of integrins[J]. J Leukoc Biol, 2017, 102(3): 677-83. doi:10.1189/jlb.3mr0117-024r |
| [8] | Wang Z, Wang B, Zhang J, et al. Chemokine (C-C motif) ligand 2/chemokine receptor 2 (CCR2) axis blockade to delay chondrocyte hypertrophy as a therapeutic strategy for osteoarthritis[J]. Med Sci Monit, 2021, 27: e930053. doi:10.12659/msm.930053 |
| [9] | Koch AE, Burrows JC, Haines GK, et al. Immunolocalization of endothelial and leukocyte adhesion molecules in human rheumatoid and osteoarthritic synovial tissues[J]. Lab Invest, 1991, 64(3): 313-20. doi:doi:10.1084/jem.173.3.779 |
| [10] | 韩建建, 徐 鹏, 韩礼纲, 等. 关节镜清理术联合透明质酸钠对膝骨关节炎病人关节液细胞间黏附分子-1、白细胞介素-6的影响[J]. 蚌埠医学院学报, 2021, 46(2): 222-5. doi:10.13898/j.cnki.issn.1000-2200.2021.02.022 |
| [11] | Wojdasiewicz P, Poniatowski ŁA, Kotela A, et al. The chemokine CX3CL1 (fractalkine) and its receptor CX3CR1: occurrence and potential role in osteoarthritis[J]. Arch Immunol Ther Exp, 2014, 62(5): 395-403. doi:10.1007/s00005-014-0275-0 |
| [12] | Chen Y, Mehmood K, Chang YF, et al. The molecular mechanisms of glycosaminoglycan biosynthesis regulating chondrogenesis and endochondral ossification[J]. Life Sci, 2023, 335: 122243. doi:10.1016/j.lfs.2023.122243 |
| [13] | Okamura Y, Mishima S, Kashiwakura JI, et al. The dual regulation of substance P-mediated inflammation via human synovial mast cells in rheumatoid arthritis[J]. Allergol Int, 2017, 66: S9-S20. doi:10.1016/j.alit.2017.03.002 |
| [14] | Mustonen AM, Nieminen P. Fatty acids and oxylipins in osteoarthritis and rheumatoid arthritis: a complex field with significant potential for future treatments[J]. Curr Rheumatol Rep, 2021, 23(6): 41. doi:10.1007/s11926-021-01007-9 |
| [15] | Wang Y, Liu J, Huang B, et al. Mathematical modeling and application of IL-1β/TNF signaling pathway in regulating chondrocyte apoptosis[J]. Front Cell Dev Biol, 2023, 11: 1288431. doi:10.3389/fcell.2023.1288431 |
| [16] | Malemud CJ. Matrix metalloproteinases and synovial joint pathology[M]//Matrix Metalloproteinases and Tissue Remodeling in Health and Disease: Target Tissues and Therapy. Amsterdam: Elsevier, 2017: 305-25. doi:10.1016/bs.pmbts.2017.03.003 |
| [17] | Miao Y, Dong Y, Huang P, et al. Increasing UCP2 expression and decreasing NOX1/4 expression maintain chondrocyte phenotype by reducing reactive oxygen species production[J]. Oncotarget, 2017, 8(38): 63750-63. doi:10.18632/oncotarget.18908 |
| [18] | Park D, Han CZ, Elliott MR, et al. Continued clearance of apoptotic cells critically depends on the phagocyte Ucp2 protein[J]. Nature, 2011, 477(7363): 220-4. doi:10.1038/nature10340 |
| [19] | Charlier E, Relic B, Deroyer C, et al. Insights on molecular mechanisms of chondrocytes death in osteoarthritis[J]. Int J Mol Sci, 2016, 17(12): E2146. doi:10.3390/ijms17122146 |
| [20] | Hu S, Zhang C, Ni L, et al. Stabilization of HIF-1α alleviates osteoarthritis via enhancing mitophagy[J]. Cell Death Dis, 2020, 11(6): 481. doi:10.1038/s41419-020-2680-0 |
| [21] | Xia X, Wang S, Ni B, et al. Hypoxic gastric cancer-derived exosomes promote progression and metastasis via miR-301a-3p/PHD3/HIF-1α positive feedback loop[J]. Oncogene, 2020, 39(39): 6231-44. doi:10.1038/s41388-020-01425-6 |
| [22] | Jin Y, Pan Y, Zheng S, et al. Inactivation of EGLN3 hydroxylase facilitates Erk3 degradation via autophagy and impedes lung cancer growth[J]. Oncogene, 2022, 41(12): 1752-66. doi:10.1038/s41388-022-02203-2 |
| [23] | Escribese MM, Sierra-Filardi E, Nieto C, et al. The prolyl hydroxylase PHD3 identifies proinflammatory macrophages and its expression is regulated by activin A[J]. J Immunol, 2012, 189(4): 1946-54. doi:10.4049/jimmunol.1201064 |
| [24] | Samad TA, Moore KA, Sapirstein A, et al. Interleukin-1β-mediated induction of Cox-2 in the CNS contributes to inflammatory pain hypersensitivity[J]. Nature, 2001, 410(6827): 471-5. doi:10.1038/35068566 |
| [25] | Kapoor M, Martel-Pelletier J, Lajeunesse D, et al. Role of proinflammatory cytokines in the pathophysiology of osteoarthritis[J]. Nat Rev Rheumatol, 2011, 7(1): 33-42. doi:10.1038/nrrheum.2010.196 |
| [26] | Sun K, Jing X, Guo J, et al. Mitophagy in degenerative joint diseases[J]. Autophagy, 2021, 17(9): 2082-92. doi:10.1080/15548627.2020.1822097 |
| [27] | Liu Y, Lu T, Liu Z, et al. Six macrophage-associated genes in synovium constitute a novel diagnostic signature for osteoarthritis[J]. Front Immunol, 2022, 13: 936606. doi:10.3389/fimmu.2022.936606 |
| [28] | Wang XY, Han XL, Ma JJ, et al. 5-hydroxymethylfurfural attenuates osteoarthritis by upregulating of glucose metabolism in chondrocytes[J]. Phytomedicine, 2025, 139: 156499. doi:10.1016/j.phymed.2025.156499 |
| [29] | Hao GH, Han SQ, Xiao ZG, et al. Synovial mast cells and osteoarthritis: current understandings and future perspectives[J]. Heliyon, 2024, 10(24): e41003. doi:10.1016/j.heliyon.2024.e41003 |
| [30] | Metcalfe DD, Baram D, Mekori YA. Mast cells[J]. Physiol Rev, 1997, 77(4): 1033-79. doi:10.1152/physrev.1997.77.4.1033 |
| [31] | Fang L, Liu K, Liu C, et al. Tumor accomplice: T cell exhaustion induced by chronic inflammation[J]. Front Immunol, 2022, 13: 979116. doi:10.3389/fimmu.2022.979116 |
| [32] | Holter DB, Zahalka S, Brösamlen J, et al. Mast cells activated in vitro can modulate macrophage polarization and antibacterial responses[J]. J Allergy Clin Immunol, 2025, 156(3): 754-73. doi:10.1016/j.jaci.2025.02.040 |
| [33] | Mihlan M, Wissmann S, Gavrilov A, et al. Neutrophil trapping and nexocytosis, mast cell-mediated processes for inflammatory signal relay[J]. Cell, 2024, 187(19): 5316-35.e28. doi:10.1016/j.cell.2024.07.014 |
| [34] | Poon IKH, Ravichandran KS. Targeting efferocytosis in inflam-maging[J]. Annu Rev Pharmacol Toxicol, 2024, 64: 339-57. doi:10.1146/annurev-pharmtox-032723-110507 |
| [35] | Wherry EJ, Kurachi M. Molecular and cellular insights into T cell exhaustion[J]. Nat Rev Immunol, 2015, 15(8): 486-99. doi:10.1038/nri3862 |
| [36] | Wang TT, He CQ. Pro-inflammatory cytokines: The link between obesity and osteoarthritis[J]. Cytokine Growth Factor Rev, 2018, 44: 38-50. doi:10.1016/j.cytogfr.2018.10.002 |
| [37] | Zhang C, Lin Y, Yan CH, et al. Adipokine signaling pathways in osteoarthritis[J]. Front Bioeng Biotechnol, 2022, 10: 865370. doi:10.3389/fbioe.2022.865370 |
| [38] | Zheng S, Li YT, Yin L, et al. Identification of sulfur metabolism-related gene signature in osteoarthritis and TM9SF2's sustenance effect on M2 macrophages' phagocytic activity[J]. J Orthop Surg Res, 2024, 19(1): 62. doi:10.1186/s13018-023-04384-2 |
| [39] | Peffers MJ, Chabronova A, Balaskas P, et al. SnoRNA signatures in cartilage ageing and osteoarthritis[J]. Sci Rep, 2020, 10(1): 10641. doi:10.1038/s41598-020-67446-z |
| [40] | Song M, Pang L, Zhang M, et al. Cdc2-like kinases: structure, biological function, and therapeutic targets for diseases[J]. Signal Transduct Target Ther, 2023, 8(1): 148. doi:10.1038/s41392-023-01409-4 |
| [41] | 杨 雷, 伍搏宇, 熊 辉, 等. 基于基因芯片技术结合CIBERSORT反卷积算法研究骨关节炎的免疫机制及潜在中药靶向预测[J]. 中国免疫学杂志, 2022, 38(3): 288-95. doi:10.3969/j.issn.1000-484X.2022.03.006 |
| [42] | 唐仕荣, 巫永华, 刘恩岐, 等. 枸杞多糖的提取分级及其氧自由基清除能力分析[J]. 食品科技, 2018, 43(10): 251-6. |
| [43] | 蔡松涛, 孙京涛, 魏 瑄. 枸杞多糖抑制核因子κB(NF-κB)通路降低骨关节炎软骨细胞炎性细胞因子水平[J]. 细胞与分子免疫学杂志, 2018, 34(11): 989-93. |
| [44] | 赵 飞, 丁 冬, 巩 凡, 等. 枸杞多糖干预膝骨关节炎模型兔关节软骨组织CD151和基质金属蛋白酶3的表达[J]. 中国组织工程研究, 2020, 24(17): 2648-53. doi:10.3969/j.issn.2095-4344.2562 |
| [45] | Mi B, Wang J, Liu Y, et al. Icariin activates autophagy via down-regulation of the NF‑κB signaling-mediated apoptosis in chondro-cytes[J]. Front Pharmacol, 2018, 9: 605. doi:10.3389/fphar.2018.00605 |
| [46] | Huang H, Zhang ZF, Qin FW, et al. Icariin inhibits chondrocyte apoptosis and angiogenesis by regulating the TDP-43 signaling pathway[J]. Molec Gen & Gen Med, 2019, 7(4): e00586. doi:10.1002/mgg3.586 |
| [47] | Zhu PT, Wu Y, Yang AH, et al. Catalpol suppressed proliferation, growth and invasion of CT26 colon cancer by inhibiting inflammation and tumor angiogenesis[J]. Biomed Pharmacother, 2017, 95: 68-76. doi:10.1016/j.biopha.2017.08.049 |
| [48] | Zeng YF, Wang R, Bian Y, et al. Catalpol attenuates IL-1β induced matrix catabolism, apoptosis and inflammation in rat chondrocytes and inhibits cartilage degeneration[J]. Med Sci Monit, 2019, 25: 6649-59. doi:10.12659/msm.916209 |
| [49] | Lu S, Xiao X, Cheng M. Matrine inhibits IL-1β-induced expression of matrix metalloproteinases by suppressing the activation of MAPK and NF-κB in human chondrocytes in vitro [J]. Int J Clin Exp Pathol, 2015, 8(5): 4764-72. doi:10.1016/S1359-6349(09)71986-X |
| [50] | Wu DY, Zhu XY, Kang XD, et al. The protective effect of sophocarpine in osteoarthritis: an in vitro and in vivo study[J]. Int Immunopharmacol, 2019, 67: 145-51. doi:10.1016/j.intimp.2018.11.046 |
| [51] | 梁桂洪, 曾令烽, 潘建科, 等. 川芎嗪干预骨性关节炎的机制研究进展[J]. 中华中医药杂志, 2020, 35(12): 6228-32. |
| [52] | 李飞龙, 谢平金, 柴生颋, 等. 川芎嗪对膝骨性关节炎大鼠软骨VEGF表达的影响[J]. 中国骨质疏松杂志, 2018, 24(7): 904-9. |
| [53] | 朱海泉, 刘子敏, 孟祥圣, 等. 川芎嗪抑制NF-κB P65磷酸化对LPS诱导的骨关节炎软骨细胞凋亡和炎症反应的调节作用[J]. 中国免疫学杂志, 2019, 35(2): 181-5. |
| [54] | 高 坤, 张 勇, 陈大宇, 等. 牛膝总皂苷干预兔膝骨关节炎滑膜液来源细胞因子的表达[J]. 中国组织工程研究, 2019, 23(33): 5317-21. |
| [55] | 孙雪莲, 刘 渊, 周红海. 牛膝总皂苷对兔膝骨关节炎软骨组织形态变化及关节液中IL-1β、TGF-β1含量的影响[J]. 中药新药与临床药理, 2016, 27(3): 321-6. |
| [56] | Xu X, Zhang X, Diao Y, et al. Achyranthes bidentate saponins protect rat articular chondrocytes against interleukin-1β-induced inflammation and apoptosis in vitro[J]. Kaohsiung J Med Sci, 2017,33(2):62-8. doi:10.1016/j.kjms.2016.11.004 |
| [57] | Wei B, Zhang Y, Tang L, et al. Protective effects of quercetin against inflammation and oxidative stress in a rabbit model of knee osteoarthritis[J]. Drug Dev Res, 2019, 80(3): 360-7. doi:10.1002/ddr.21510 |
| [1] | 陶红成, 梁富凯, 黄文波, 范思奇, 曾平. 酒精暴露与股骨头坏死的潜在关联:基于机器学习构建诊断模型[J]. 南方医科大学学报, 2026, 46(3): 559-569. |
| [2] | 邱佳惠, 陈萌, 满如, 陈鑫, 邱东瑞, 常启同, 马洪玉. 化瘀通便汤通过抑制PI3K/Akt信号通路促进慢传输型便秘大鼠肠道Cajal间质细胞的胞葬作用[J]. 南方医科大学学报, 2026, 46(2): 293-300. |
| [3] | 陈君尧, 陈泽宇, 林钊杰, 方梦浩, 沈超英, 许琦, 张晓怡, 卢鲁. 饮茶对胃肠道疾病风险的双重作用:基于可解释机器学习与大语言模型的联合预测辅助模型[J]. 南方医科大学学报, 2026, 46(2): 353-361. |
| [4] | 崔运能, 冯敏清, 姚亮凤, 严杰文, 李闻瀚, 黄燕平. 基于欠采样的影像组学机器学习模型术前预测子宫肌瘤高强度聚焦超声消融效果[J]. 南方医科大学学报, 2026, 46(1): 141-149. |
| [5] | 王喆, 孔柯瑜, 金明昊, 伍信儒, 范文轩, 翟赞京, 胡子豪, 牛琳, 齐岩松, 徐永胜. 叶酸预处理髌下脂肪垫来源间充质干细胞来源的外泌体调控巨噬细胞极化[J]. 南方医科大学学报, 2026, 46(1): 166-174. |
| [6] | 黄启智, 谢戴鹏, 姚霖彤, 李洽轩, 吴少伟, 周海榆. 肿瘤微环境特异性CT影像组学标签预测非小细胞肺癌免疫治疗疗效[J]. 南方医科大学学报, 2025, 45(9): 1903-1918. |
| [7] | 王莹, 李静, 王伊迪, 华明钰, 胡玮彬, 张晓智. 原发性肝癌患者的临床结局与治疗反应预测模型:基于失巢凋亡和免疫基因[J]. 南方医科大学学报, 2025, 45(9): 1967-1979. |
| [8] | 姜君, 封硕, 孙银贵, 安燕. 经尿道前列腺钬激光剜除术后低体温风险预测模型:基于逻辑回归、决策树和支持向量机[J]. 南方医科大学学报, 2025, 45(9): 2019-2025. |
| [9] | 陈鑫源, 吴成挺, 李瑞迪, 潘雪芹, 张耀丹, 陶俊宇, 林才志. 双术汤通过P53/SLC7A11/GPX4通路诱导胃癌细胞铁死亡[J]. 南方医科大学学报, 2025, 45(7): 1363-1371. |
| [10] | 庞金龙, 赵新丽, 张振, 王豪杰, 周星琦, 杨玉梅, 李姗姗, 常小强, 李锋, 李娴. 皮肤黑色素瘤中MMRN2高表达促进肿瘤细胞的侵袭和迁移并与不良预后相关[J]. 南方医科大学学报, 2025, 45(7): 1479-1489. |
| [11] | 王康, 李海宾, 余靖, 孟源, 张虹丽. ELFN1高表达是结肠癌的预后生物标志物并促进结肠癌细胞的增殖转移[J]. 南方医科大学学报, 2025, 45(7): 1543-1553. |
| [12] | 陈梅妹, 王洋, 雷黄伟, 张斐, 黄睿娜, 杨朝阳. 基于多种机器学习算法和语音情绪特征的阈下抑郁辨识模型构建[J]. 南方医科大学学报, 2025, 45(4): 711-717. |
| [13] | 高志, 吴傲, 胡仲翔, 孙培养. 类风湿性关节炎中氧化应激与免疫浸润的生物信息学分析[J]. 南方医科大学学报, 2025, 45(4): 862-870. |
| [14] | 富丽萍, 袁立霞, 王杰, 陈学蓝, 柯桂芝, 黄煜, 杨心仪, 刘刚. 近十年低强度脉冲超声在肌骨疾病治疗中的应用进展[J]. 南方医科大学学报, 2025, 45(3): 661-668. |
| [15] | 付长龙, 陈若岚, 徐诗淇, 游锦欣, 林晴, 黄艳峰. 巴戟天多糖通过靶向lncRNA XIST调控糖酵解-焦亡延缓小鼠骨关节炎软骨细胞退变[J]. 南方医科大学学报, 2025, 45(12): 2541-2550. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||