Journal of Southern Medical University ›› 2025, Vol. 45 ›› Issue (7): 1442-1450.doi: 10.12122/j.issn.1673-4254.2025.07.10
Previous Articles Next Articles
Hongling ZHANG( ), Changyun GU(
), Changyun GU( ), Qiyan WANG, Xiaolan HUANG, Qianchong RAN, Zheng REN, Yubo LIU, Yansha LUO, Shuaiji PAN, Meiqing YANG, Jingyan JI, Xiaoye JIN(
), Qiyan WANG, Xiaolan HUANG, Qianchong RAN, Zheng REN, Yubo LIU, Yansha LUO, Shuaiji PAN, Meiqing YANG, Jingyan JI, Xiaoye JIN( )
)
Received:2025-03-25
Online:2025-07-20
Published:2025-07-17
Contact:
Xiaoye JIN
E-mail:229598103@qq.com;15060315252@qq.com;1115259825@qq.com
Supported by:Hongling ZHANG, Changyun GU, Qiyan WANG, Xiaolan HUANG, Qianchong RAN, Zheng REN, Yubo LIU, Yansha LUO, Shuaiji PAN, Meiqing YANG, Jingyan JI, Xiaoye JIN. Forensic performance and genetic background analyses of Guizhou Chuanqing population using a self-constructed microhaplotype panel[J]. Journal of Southern Medical University, 2025, 45(7): 1442-1450.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.j-smu.com/EN/10.12122/j.issn.1673-4254.2025.07.10
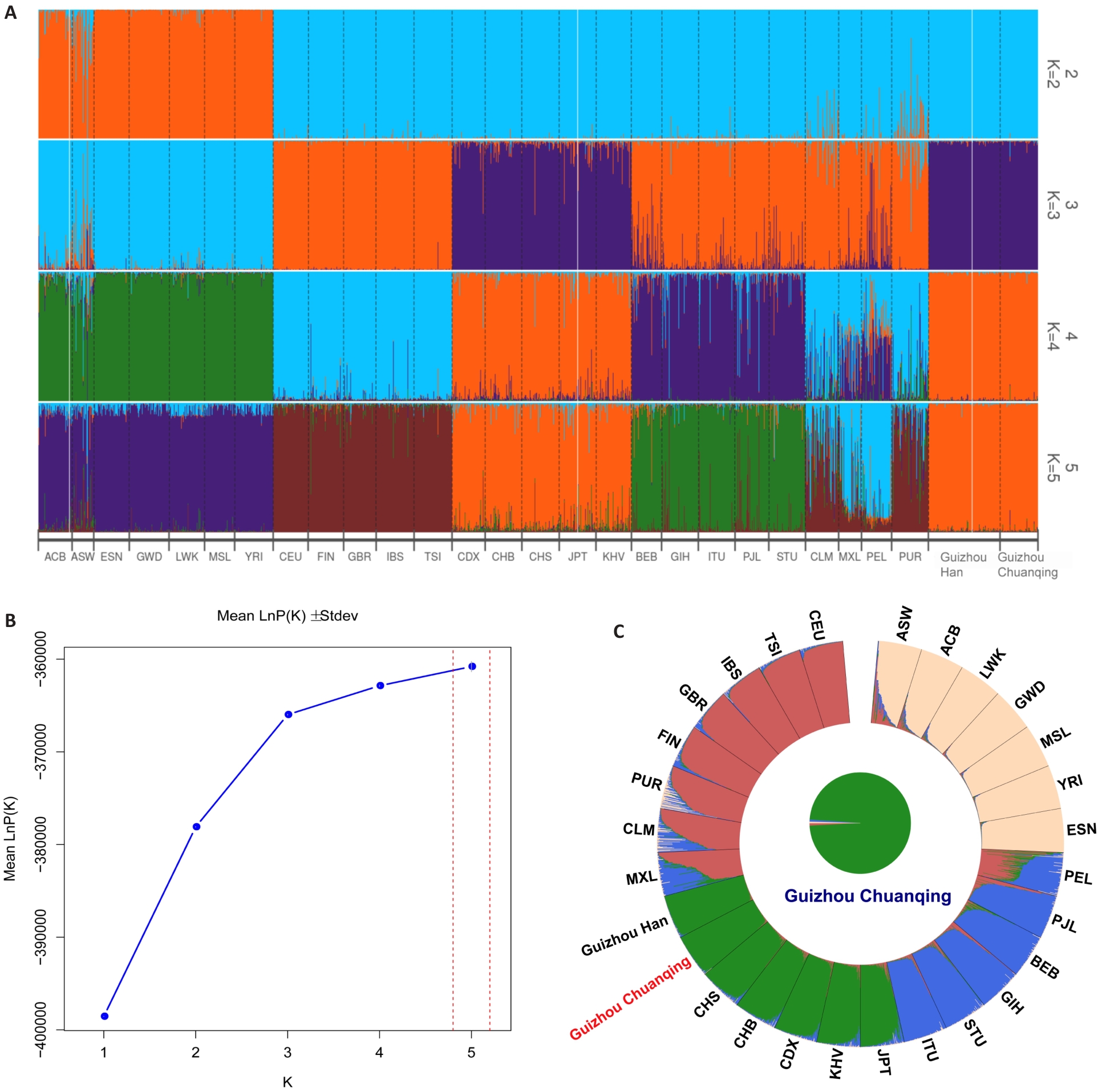
Fig.7 Genetic structure analysis of Chuanqing and the reference populations. A: Population genetic structure under different K values. B: Mean LnP(K) values across different K values. C: Genetic component dissection of Chuanqing and the reference populations at K=4.
| [1] | Yan Q. Distribution characteristics and historical causes of unreco-gnized ethnic populations in Guizhou [J]. Popular Business, 2009, (2): 27-31. |
| [2] | 李良品. 贵州方志中有关“穿青人” 及其先民族源和族称的记载[J]. 贵州民族研究, 2011, 32(2): 159-66. |
| [3] | 黄锦树. 贵州穿青人族属考析: 与畲族同出古帝少昊[J]. 韩山师范学院学报, 2013, 34(1): 21-9. |
| [4] | Liu Y, Zhang H, He G, et al. Forensic features and population genetic structure of Dong, Yi, Han, and chuanqing human populations in southwest China inferred from insertion/deletion markers[J]. Front Genet, 2020, 11: 360. doi:10.3389/fgene.2020.00360 |
| [5] | Lu J, Zhang H, Ren Z, et al. Genome-wide analysis of unrecognised ethnic group Chuanqing people revealing a close affinity with Southern Han Chinese[J]. Ann Hum Biol, 2020, 47(5): 465-71. doi:10.1080/03014460.2020.1782470 |
| [6] | 李思睿. 中间群体、边民与他者: 贵州穿青人的族群形成和认同[J]. 青海民族研究, 2017, 28(1): 61-4. doi:10.ssss/j.issn.1005-5681.2017.1.014 |
| [7] | Chen P, He G, Xing H, et al. Forensic characteristics and phylogenetic analysis of 23 Y-STR loci in the Miao population from Guizhou province, southwest China[J]. Ann Hum Biol, 2019, 46(1): 84-7. doi:10.1080/03014460.2019.1583374 |
| [8] | Luo L, Gao H, Yao L, et al. Genetic diversity, forensic feature, and phylogenetic analysis of Guizhou Tujia population via 19 X-STRs[J]. Mol Genet Genomic Med, 2020, 8(11): e1473. doi:10.1002/mgg3.1473 |
| [9] | Zhang H, Huang X, Jin X, et al. Comprehensive analyses of genetic diversities and population structure of the Guizhou Dong group based on 44 Y-markers[J]. PeerJ, 2023, 11: e16183. doi:10.7717/peerj.16183 |
| [10] | 赵勤松, 任 峥, 张红玲, 等. 贵州汉族1 9 个STR基因座多态性及法医学应用[J]. 法医学杂志, 2017, 33(4): 388-92. |
| [11] | Zhang H, Yang M, Zhang H, et al. Forensic features and phylogenetic structure survey of four populations from southwest China via the autosomal insertion/deletion markers[J]. Forensic Sci Res, 2024, 9(2): owad052. doi:10.1093/fsr/owae059 |
| [12] | Yang CH, Jin XY, Guo YX, et al. Genetic distribution analyses and population background explorations of Gansu Yugur and Guizhou Miao groups via InDel markers[J]. J Hum Genet, 2019, 64(6): 535-43. doi:10.1038/s10038-019-0595-3 |
| [13] | Chen J, He G, Ren Z, et al. Fine-scale population admixture landscape of Tai-kadai-speaking Maonan in southwest China inferred from genome-wide SNP data[J]. Front Genet, 2022, 13: 815285. doi:10.3389/fgene.2022.815285 |
| [14] | 罗 雪, 张文飞, 杨 林, 等. 贵州水族人群11个Y-SNP位点的多态性及法医学应用[J]. 法医学杂志, 2020, 36(6): 791-6. doi:10.12116/j.issn.1004-5619.2020.06.008 |
| [15] | Cui W, Chen M, Yang Y, et al. Applications of 1993 single nucleotide polymorphism loci in forensic pairwise kinship identifications and inferences[J]. Forensic Sci Int Genet, 2023, 65: 102889. doi:10.1016/j.fsigen.2023.102889 |
| [16] | Pakstis AJ, Haigh E, Cherni L, et al. 52 additional reference population samples for the 55 AISNP panel[J]. Forensic Sci Int Genet, 2015, 19: 269-71. doi:10.1016/j.fsigen.2015.08.003 |
| [17] | Kidd KK, Pakstis AJ, Speed WC, et al. Microhaplotype loci are a powerful new type of forensic marker[J]. Forensic Sci Int Genet Suppl Ser, 2013, 4(1): e123-4. doi:10.1016/j.fsigss.2013.10.063 |
| [18] | Oldoni F, Kidd KK, Podini D. Microhaplotypes in forensic genetics[J]. Forensic Sci Int Genet, 2019, 38: 54-69. doi:10.1016/j.fsigen.2020.102398 |
| [19] | Chandra D, Mishra VC, Raina A, et al. Mutation rate evaluation at 21 autosomal STR loci: Paternity testing experience[J]. Leg Med, 2022, 58: 102080. doi:10.1016/j.legalmed.2022.102080 |
| [20] | Wen D, Xing H, Liu Y, et al. The application of short and highly polymorphic microhaplotype loci in paternity testing and sibling testing of temperature-dependent degraded samples[J]. Front Genet, 2022, 13: 983811. doi:10.3389/fgene.2022.983811 |
| [21] | Fan H, Xie Q, Wang L, et al. Microhaplotype and Y-SNP/STR (MY): a novel MPS-based system for genotype pattern recognition in two-person DNA mixtures[J]. Forensic Sci Int Genet, 2022, 59: 102705. doi:10.1016/j.fsigen.2022.102705 |
| [22] | Cheung EYY, Phillips C, Eduardoff M, et al. Performance of ancestry-informative SNP and microhaplotype markers[J]. Forensic Sci Int Genet, 2019, 43: 102141. doi:10.1016/j.fsigen.2019.102141 |
| [23] | de Barros Rodrigues ML, Rodrigues MP, Norton HL, et al. Large-scale selection of highly informative microhaplotypes for ancestry inference and population specific informativeness[J]. Forensic Sci Int Genet, 2025, 74: 103153. doi:10.1016/j.fsigen.2024.103153 |
| [24] | de la Puente M, Ruiz-Ramírez J, Ambroa-Conde A, et al. Broadening the applicability of a custom multi-platform panel of microhaplotypes: bio-geographical ancestry inference and expanded reference data[J]. Front Genet, 2020, 11: 581041. doi:10.3389/fgene.2020.581041 |
| [25] | Huang SN, Sheng MC, Li Z, et al. Inferring bio-geographical ancestry with 35 microhaplotypes[J]. Forensic Sci Int, 2022, 341: 111509. doi:10.1016/j.forsciint.2022.111509 |
| [26] | Pakstis AJ, Gandotra N, Speed WC, et al. The population genetics characteristics of a 90 locus panel of microhaplotypes[J]. Hum Genet, 2021, 140(12): 1753-73. doi:10.1007/s00439-021-02382-0 |
| [27] | Yu WS, Feng YS, Kang KL, et al. Screening of highly discriminative microhaplotype markers for individual identification and mixture deconvolution in East Asian populations[J]. Forensic Sci Int Genet, 2022, 59: 102720. doi:10.1016/j.fsigen.2022.102720 |
| [28] | Zou X, He GL, Liu J, et al. Screening and selection of 21 novel microhaplotype markers for ancestry inference in ten Chinese subpopulations[J]. Forensic Sci Int Genet, 2022, 58: 102687. doi:10.1016/j.fsigen.2022.102687 |
| [29] | Gu C, Huo W, Huang X, et al. Developmental and validation of a novel small and high-efficient panel of microhaplotypes for forensic genetics by the next generation sequencing[J]. BMC Genomics, 2024, 25(1): 958. doi:10.1186/s12864-024-10880-4 |
| [30] | 邹 星. 法医物证族源推断的新微单倍型遗传标记研究[D]. 成都: 四川大学, 2021. |
| [31] | Byrska-Bishop M, Evani US, Zhao XF, et al. High-coverage whole-genome sequencing of the expanded 1000 Genomes Project cohort including 602 trios[J]. Cell, 2022, 185(18): 3426-40.e19. |
| [32] | Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data[J]. Bioinformatics, 2014, 30(15): 2114-20. doi:10.1093/bioinformatics/btu170 |
| [33] | Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform[J]. Bioinformatics, 2009, 25(14): 1754-60. doi:10.1093/bioinformatics/btp324 |
| [34] | Gouy A, Zieger M. STRAF: a convenient online tool for STR data evaluation in forensic genetics[J]. Forensic Sci Int Genet, 2017, 30: 148-51. doi:10.1016/j.fsigen.2017.07.007 |
| [35] | Rousset F. genepop'007: a complete re-implementation of the genepop software for Windows and Linux[J]. Mol Ecol Resour, 2008, 8(1): 103-6. doi:10.1111/j.1471-8286.2007.01931.x |
| [36] | Ota T. DISPAN: genetic distance and phylogenetic analysis[J]. Pennsylvania State University, University Park, PA, 1993. |
| [37] | Kumar S, Stecher G, Li M, et al. MEGA X: molecular evolutionary genetics analysis across computing platforms[J]. Mol Biol Evol, 2018, 35(6): 1547-9. doi:10.1093/molbev/msy096 |
| [38] | Falush D, Stephens M, Pritchard JK. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies[J]. Genetics, 2003, 164(4): 1567-87. doi:10.1093/genetics/164.4.1567 |
| [39] | Earl DA, VonHoldt BM. STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method[J]. Conserv Genet Resour, 2012, 4(2): 359-61. doi:10.1007/s12686-011-9548-7 |
| [40] | Jakobsson M, Rosenberg NA. CLUMPP: a cluster matching and permutation program for dealing with label switching and multimo-dality in analysis of population structure[J]. Bioinformatics, 2007, 23(14): 1801-6. doi:10.1093/bioinformatics/btm233 |
| [41] | Francis RM. Pophelper: an R package and web app to analyse and visualize population structure[J]. Mol Ecol Resour, 2017, 17(1): 27-32. doi:10.1111/1755-0998.12509 |
| [42] | Chen S, Lei C, Zhao X, et al. AncestryPainter 2.0: visualizing ancestry composition and admixture history graph[J]. Genome Biol Evol, 2024, 16(11): evae249. doi:10.1093/gbe/evae249 |
| [43] | 雷 勇. 社会历史、宗教生活与族群身份的建构: 以黔西北穿青人为例[J]. 青海民族研究, 2012, 23(4): 11-6. doi:10.3969/j.issn.1005-5681.2012.04.003 |
| [44] | 周成勋. 穿青人民族认同问题研究[D]. 贵阳: 贵州民族大学, 2013. |
| No related articles found! |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||