南方医科大学学报 ›› 2026, Vol. 46 ›› Issue (4): 946-955.doi: 10.12122/j.issn.1673-4254.2026.04.23
• • 上一篇
收稿日期:2025-09-28
出版日期:2026-04-20
发布日期:2026-04-24
通讯作者:
张煜
E-mail:wmei2933@gmail.com;yuzhang@smu.edu.cn
作者简介:王璟儿,在读硕士研究生, E -mail: wmei2933@gmail.com
基金资助:Received:2025-09-28
Online:2026-04-20
Published:2026-04-24
Contact:
Yu ZHANG
E-mail:wmei2933@gmail.com;yuzhang@smu.edu.cn
Supported by:摘要:
目的 构建一种融合临床视觉先验的掩码图像建模框架,以提升胸部X光片的语义理解与诊断性能。 方法 提出视觉先验引导的掩码图像重建框架(VP-MIM)。该方法利用放射科医生的眼动数据在掩码阶段区分诊断相关与无关区域,并据此实施注意力引导的掩码策略;同时构建金字塔注意力重建模块,在重建阶段引入多尺度监督,并结合语义感知的眼动热力图以优化特征学习。 结果 在RSNA Pneumonia与ChestXray-14两个公开数据集上,VP-MIM的预训练质量在线性评估这一实验设置中展现出显著优势。在仅2616个预训练样本下,于RSNA Pneumonia单标签分类中获得AUC=86.83,在ChestXray-14多标签分类中达到平均AUC(mAUC)=72.82。在全量微调实验设置中,随着预训练数据规模的扩大,VP-MIM在ChestXray-14上取得mAUC=85.49的性能,验证了其良好的可扩展性和其在实际诊断任务中的优秀性能。 结论 VP-MIM有效缓解了医学影像中MIM方法的语义损失与多尺度不足问题,提升了胸片诊断性能,并可以在数据规模扩展中保持稳定增益。
王璟儿, 张煜. 基于临床视觉先验引导的掩码图像建模可提升胸部X光诊断性能[J]. 南方医科大学学报, 2026, 46(4): 946-955.
Jinger WANG, Yu ZHANG. Visual prior-guided masked image modeling enhances chest X-ray diagnostic efficacy[J]. Journal of Southern Medical University, 2026, 46(4): 946-955.
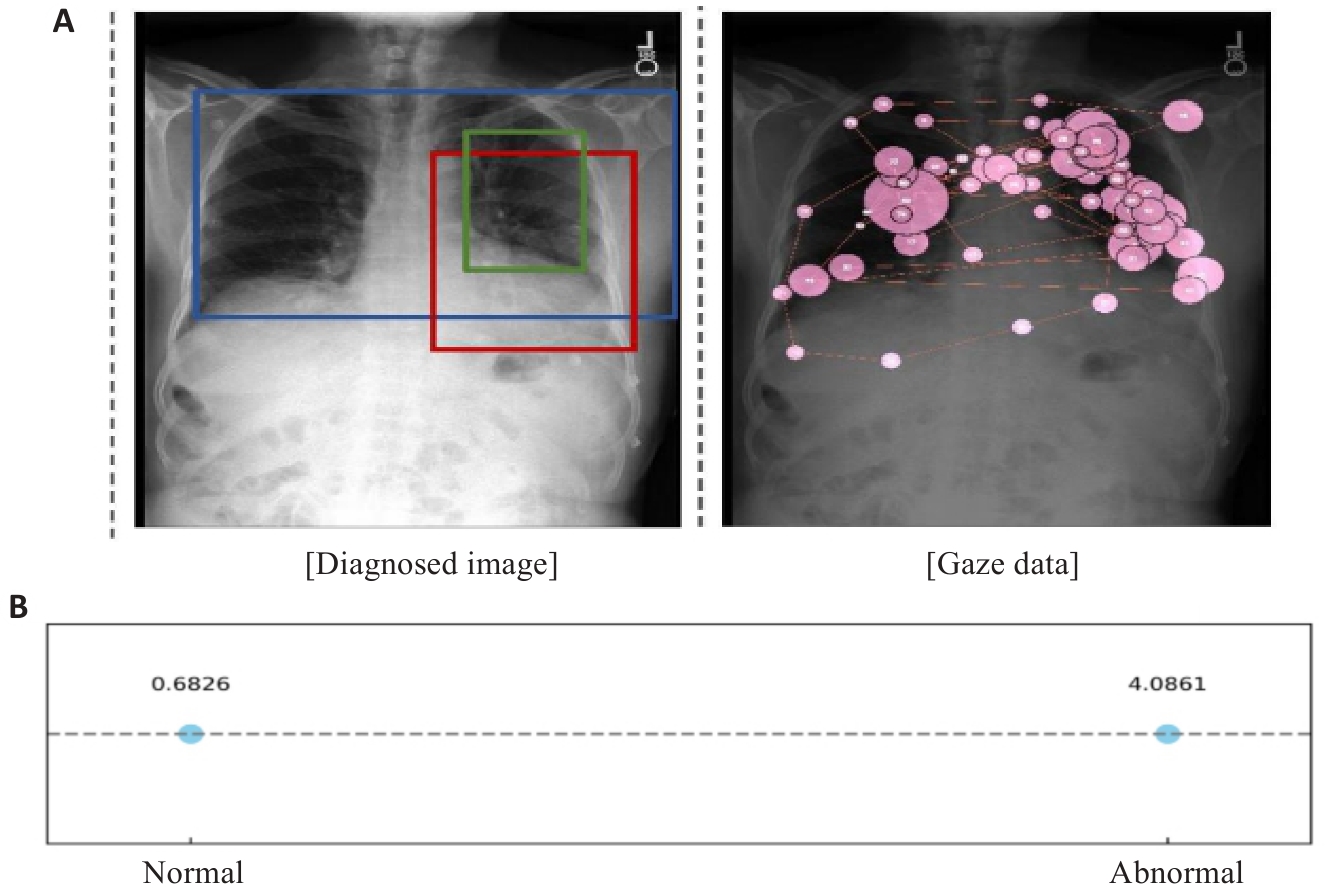
图1 不同尺度胸部异常及其与注视数据的对应关系
Fig.1 Illustration of thoracic abnormalities of varying sizes and the alignment of gaze data with these regions. A: Prominent heart. Low lung volumes.Left perihilar opacity. No large effusion. This could represent very early congestion or an early left suprahilar infiltrate. B: Mean gaze density for two kinds of areas.
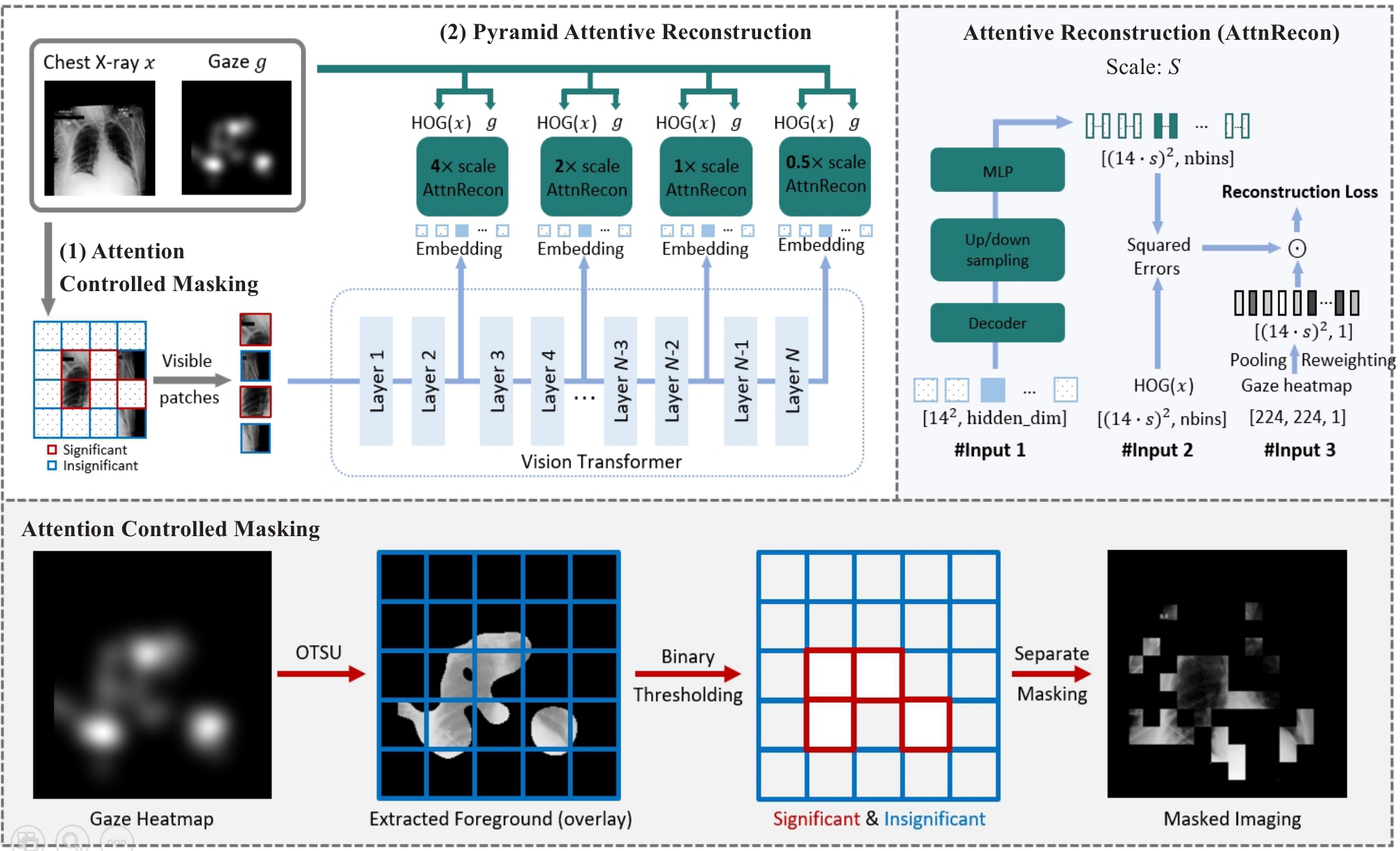
图2 VP-MIM整体框架示意图
Fig.2 Overview of the proposed VP-MIM (upper-left part) by using ViT-B/16 as the default vision backbone. VP-MIM integrates (1) attention-controlled masking to preserve clinically significant features based on radiologist attention, and (2) pyramid attentive reconstruction to reconstruct multi-scale HOG features using embeddings from selected transformer layers. The design of each attentive reconstruction module is shown in the upper-right panel.
| Methods | RSNA | CXR-14 | ||||
|---|---|---|---|---|---|---|
| 1% | 10% | 100% | 1% | 10% | 100% | |
| MAE[ | 79.19 (0.06) | 83.04 (0.01) | 84.64 (0.01) | 56.91 (0.59) | 63.73 (0.56) | 68.33 (0.34) |
| MaskFeat[ | 80.99 (0.23) | 84.29 (0.11) | 85.67 (0.19) | 61.18 (0.62) | 66.19 (1.16) | 70.21 (0.15) |
| dMAE[ | 81.74 (0.40) | 84.65 (0.10) | 85.70 (0.03) | 58.71 (1.53) | 65.41 (1.73) | 69.88 (0.42) |
| MaskAlign[ | 79.64 (0.82) | 84.56 (0.12) | 85.26 (0.01) | 58.37 (0.93) | 65.10 (3.96) | 70.52 (1.47) |
| HPM[ | 82.01 (0.03) | 84.30 (0.02) | 85.46 (0.02) | 59.93 (0.18) | 65.57 (2.48) | 70.87 (0.71) |
| AttnMA[ | 79.15 (0.98) | 84.06 (0.15) | 85.09 (0.02) | 57.55 (0.34) | 64.81 (3.42) | 70.01 (1.42) |
| DeepMIM[ | 81.89 (0.36) | 83.96 (0.12) | 85.18 (0.11) | 61.97 (0.35) | 66.41 (1.52) | 71.20 (0.64) |
| VP-MIM | 83.51 (0.08) | 85.49 (0.05) | 86.83 (0.01) | 63.48 (0.22) | 68.53 (0.68) | 72.82 (0.02) |
表1 不同MIM方法在线性评估实验设置中的定量比较
Tab.1 Quantitative comparison between all MIM methods considering the linear probing performance
| Methods | RSNA | CXR-14 | ||||
|---|---|---|---|---|---|---|
| 1% | 10% | 100% | 1% | 10% | 100% | |
| MAE[ | 79.19 (0.06) | 83.04 (0.01) | 84.64 (0.01) | 56.91 (0.59) | 63.73 (0.56) | 68.33 (0.34) |
| MaskFeat[ | 80.99 (0.23) | 84.29 (0.11) | 85.67 (0.19) | 61.18 (0.62) | 66.19 (1.16) | 70.21 (0.15) |
| dMAE[ | 81.74 (0.40) | 84.65 (0.10) | 85.70 (0.03) | 58.71 (1.53) | 65.41 (1.73) | 69.88 (0.42) |
| MaskAlign[ | 79.64 (0.82) | 84.56 (0.12) | 85.26 (0.01) | 58.37 (0.93) | 65.10 (3.96) | 70.52 (1.47) |
| HPM[ | 82.01 (0.03) | 84.30 (0.02) | 85.46 (0.02) | 59.93 (0.18) | 65.57 (2.48) | 70.87 (0.71) |
| AttnMA[ | 79.15 (0.98) | 84.06 (0.15) | 85.09 (0.02) | 57.55 (0.34) | 64.81 (3.42) | 70.01 (1.42) |
| DeepMIM[ | 81.89 (0.36) | 83.96 (0.12) | 85.18 (0.11) | 61.97 (0.35) | 66.41 (1.52) | 71.20 (0.64) |
| VP-MIM | 83.51 (0.08) | 85.49 (0.05) | 86.83 (0.01) | 63.48 (0.22) | 68.53 (0.68) | 72.82 (0.02) |
Attention- controlled masking | Pyramid attentive reconstruction | mAUC (×100) | ||
|---|---|---|---|---|
| HOG | Pyramid | Attentive | ||
| √ | 68.97 (0.16) | |||
| √ | √ | 70.52 (0.44) | ||
| √ | √ | √ | 72.18 (0.17) | |
| √ | √ | √ | √ | 72.82 (0.02) |
表2 针对关键模块的消融实验
Tab.2 Ablation study on key components
Attention- controlled masking | Pyramid attentive reconstruction | mAUC (×100) | ||
|---|---|---|---|---|
| HOG | Pyramid | Attentive | ||
| √ | 68.97 (0.16) | |||
| √ | √ | 70.52 (0.44) | ||
| √ | √ | √ | 72.18 (0.17) | |
| √ | √ | √ | √ | 72.82 (0.02) |
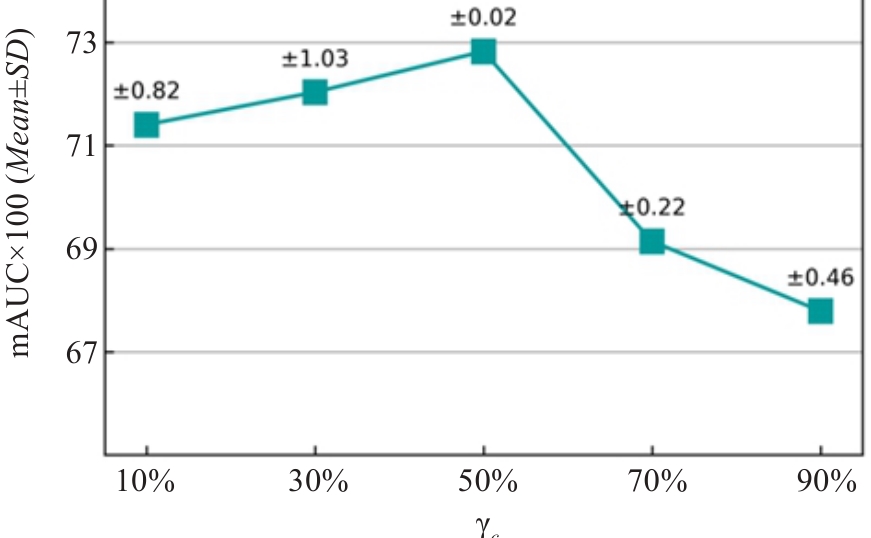
图6 针对临床掩码率的消融实验
Fig.6 Ablation study on clinical masking ratio rc. The rc changes while the global masking ratio r=0.75 remain fixed throughout pre-training.
| [1] | Bansal T, Beese R. Interpreting a chest X-ray[J]. Br J Hosp Med, 2019, 80(5): C75-9. doi:10.12968/hmed.2019.80.5.c75 |
| [2] | Bao H, Dong L, Piao S, et al. BEiT: BERT pre-training of image transformers[C/OL]//The Tenth International Conference on Learning Representations (ICLR). 2022. Available from: |
| [3] | He KM, Chen XL, Xie SN, et al. Masked autoencoders are scalable vision learners[C]//2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR). June 18-24, 2022, New Orleans, LA, USA. IEEE, 2022: 16000-09. doi:10.1109/cvpr52688.2022.01553 |
| [4] | Huang SC, Shen LY, Lungren MP, et al. GLoRIA: a multimodal global-local representation learning framework for label-efficient medical image recognition[C]//2021 IEEE/CVF International Conference on Computer Vision (ICCV). October 10-17, 2021, Montreal, QC, Canada. IEEE, 2022: 3922-31. doi:10.1109/iccv48922.2021.00391 |
| [5] | Li JX, Wang YQ, Wang S, et al. Multiscale attention guided network for COVID-19 diagnosis using chest X-ray images[J]. IEEE J Biomed Health Inform, 2021, 25(5): 1336-46. doi:10.1109/jbhi.2021.3058293 |
| [6] | Wang S, Ouyang X, Liu TM, et al. Follow my eye: using gaze to supervise computer-aided diagnosis[J]. IEEE Trans Med Imag, 2022, 41(7): 1688-98. doi:10.1109/tmi.2022.3146973 |
| [7] | Ibragimov B, Mello-Thoms C. The use of machine learning in eye tracking studies in medical imaging: a review[J]. IEEE J Biomed Health Inform, 2024, 28(6): 3597-612. doi:10.1109/jbhi.2024.3371893 |
| [8] | Karargyris A, Kashyap S, Lourentzou I, et al. Creation and validation of a chest X-ray dataset with eye-tracking and report dictation for AI development[J]. Sci Data, 2021, 8(1): 92. doi:10.1038/s41597-021-00863-5 |
| [9] | Alsharid M, Cai YF, Sharma H, et al. Gaze-assisted automatic captioning of fetal ultrasound videos using three-way multi-modal deep neural networks[J]. Med Image Anal, 2022, 82: 102630. doi:10.1016/j.media.2022.102630 |
| [10] | Zhao ZH, Wang S, Wang Q, et al. Mining gaze for contrastive learning toward computer-assisted diagnosis[J]. Proc AAAI Conf Artif Intell, 2024, 38(7): 7543-51. doi:10.1609/aaai.v38i7.28586 |
| [11] | Kong Y, Wang S, Cai JD, et al. Gaze-DETR: using expert gaze to reduce false positives in vulvovaginal candidiasis screening[C]//Medical Image Computing and Computer Assisted Intervention- MICCAI 2024. Cham: Springer, 2024: 133-43. doi:10.1007/978-3-031-72083-3_13 |
| [12] | Bigolin Lanfredi R, Zhang MY, Auffermann WF, et al. REFLACX, a dataset of reports and eye-tracking data for localization of abnormalities in chest x-rays[J]. Sci Data, 2022, 9(1): 350. doi:10.1038/s41597-022-01441-z |
| [13] | Dosovitskiy A, Beyer L, Kolesnikov A, et al. An image is worth 16×16 words: transformers for image recognition at scale[C/OL]//The Ninth International Conference on Learning Representations (ICLR). 2021. Available from: |
| [14] | Dalal N, Triggs B. Histograms of oriented gradients for human detection[C]//2005 IEEE Computer Society Conference on Computer Vision and Pattern Recognition (CVPR'05). June 20-25, 2005, San Diego, CA, USA. IEEE, 2005: 886-93. doi:10.1109/cvpr.2005.4 |
| [15] | Chen SH, Qin J, Ji X, et al. Automatic scoring of multiple semantic attributes with multi-task feature leverage: a study on pulmonary nodules in CT images[J]. IEEE Trans Med Imag, 2017, 36(3): 802-14. doi:10.1109/tmi.2016.2629462 |
| [16] | Shin HC, Roth HR, Gao MC, et al. Deep convolutional neural networks for computer-aided detection: CNN architectures, dataset characteristics and transfer learning[J]. IEEE Trans Med Imag, 2016, 35(5): 1285-98. doi:10.1109/tmi.2016.2528162 |
| [17] | Ghalati MK, Nunes A, Ferreira H, et al. Texture analysis and its applications in biomedical imaging: a survey[J]. IEEE Rev Biomed Eng, 2022, 15: 222-46. doi:10.1109/rbme.2021.3115703 |
| [18] | Li CY, Wang XY, Eberl S, et al. Supervised variational model with statistical inference and its application in medical image segmentation[J]. IEEE Trans Biomed Eng, 2015, 62(1): 196-207. doi:10.1109/tbme.2014.2344660 |
| [19] | Huang XW, Zhang YL, Meng L, et al. Identification of ultrasonic echolucent carotid plaques using discrete Fréchet distance between bimodal gamma distributions[J]. IEEE Trans Biomed Eng, 2018, 65(5): 949-55. doi:10.1109/tbme.2017.2676129 |
| [20] | Otsu N. A threshold selection method from gray-level histograms[J]. IEEE Trans Syst Man Cybern, 1979, 9(1): 62-6. doi:10.1109/tsmc.1979.4310076 |
| [21] | Johnson AEW, Pollard TJ, Berkowitz SJ, et al. MIMIC-CXR, a de-identified publicly available database of chest radiographs with free-text reports[J]. Sci Data, 2019, 6(1): 317. doi:10.1038/s41597-019-0322-0 |
| [22] | Shih G, Wu CC, Halabi SS, et al. Augmenting the national institutes of health chest radiograph dataset with expert annotations of possible pneumonia[J]. Radiol Artif Intell, 2019, 1(1): e180041. doi:10.1148/ryai.2019180041 |
| [23] | Wang XS, Peng YF, Lu L, et al. ChestX-ray: hospital-scale chest X-ray database and benchmarks on weakly supervised classification and localization of common Thorax diseases[M]//Deep Learning and Convolutional Neural Networks for Medical Imaging and Clinical Informatics. Cham: Springer International Publishing, 2019: 369-92. doi:10.1007/978-3-030-13969-8_18 |
| [24] | Xue HW, Gao P, Li HY, et al. Stare at what you see: masked image modeling without reconstruction[C]//2023 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR). June 17-24, 2023, Vancouver, BC, Canada. IEEE, 2023: 22732-41. doi:10.1109/cvpr52729.2023.02177 |
| [25] | Ren SC, Wei FY, Zhang SAZ, et al. DeepMIM: deep supervision for masked image modeling[C]//2025 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV). February 26 - March 6, 2025, Tucson, AZ, USA. IEEE, 2025: 879-88. doi:10.1109/wacv61041.2025.00095 |
| [26] | Wei C, Fan HQ, Xie SN, et al. Masked feature prediction for self-supervised visual pre-training[C]//2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR). June 18-24, 2022, New Orleans, LA, USA. IEEE, 2022: 14648-58. doi:10.1109/cvpr52688.2022.01426 |
| [27] | Wu QL, Ye H, Gu Y, et al. Denoising masked autoencoders help robust classification[C/OL]//The Eleventh International Conference on Learning Representations(ICLR). May 1-5, 2023, Kigali, Rwanda. Available from: |
| [28] | Wang HC, Song KY, Fan JS, et al. Hard patches mining for masked image modeling[C]//2023 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR). June 17-24, 2023, Vancouver, BC, Canada. IEEE, 2023: 10375-85. doi:10.1109/cvpr52729.2023.01000 |
| [29] | Selvaraju RR, Cogswell M, Das A, et al. Grad-CAM: visual explanations from deep networks via gradient-based localization[J]. Int J Comput Vis, 2020, 128(2): 336-59. doi:10.1007/s11263-019-01228-7 |
| [30] | Yosinski J, Clune J, Bengio Y, et al. How transferable are features in deep neural networks[C]//The Twenty-Eighth Annual Conference on Neural Information Processing Systems(NeurIPS). December 8-13, 2014, Montreal, QC, Canda. IEEE 2014, 27: 3320-28. |
| [31] | Duan JX, Zhang MW, Song MH, et al. Eye tracking-enhanced deep learning for medical image analysis: a systematic review on data efficiency, interpretability, and multimodal integration[J]. Bioengineering, 2025, 12(9): 954. doi:10.3390/bioengineering12090954 |
| [32] | Wang B, Pan HY, Aboah A, et al. GazeGNN: a gaze-guided graph neural network for chest X-ray classification[C]//2024 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV). January 3-8, 2024, Waikoloa, HI, USA. IEEE, 2024: 2183-92. doi:10.1109/wacv57701.2024.00219 |
| [33] | Ma C, Zhao L, Chen YZ, et al. Eye-gaze-guided vision transformer for rectifying shortcut learning[J]. IEEE Trans Med Imag, 2023, 42(11): 3384-94. doi:10.1109/tmi.2023.3287572 |
| [34] | Chen ZH, Liu Z, Song YJ. Gaze-guided vision transformer for chest X-ray image classification[J]. Biomed Signal Process Control, 2026, 111: 108298. doi:10.1016/j.bspc.2025.108298 |
| [35] | Suara S, Jha A, Sinha P, et al. Is grad-CAM explainable in Medical images? [C]//Computer Vision and Image Processing. Cham: Springer, 2024: 124-35. doi:10.1007/978-3-031-58181-6_11 |
| [36] | Santos R, Pedrosa J, Mendonça AM, et al. Grad-CAM: The impact of large receptive fields and other caveats[J]. Comput Vis Image Underst, 2025, 258: 104383. doi:10.1016/j.cviu.2025.104383 |
| [37] | Xiang WZ, Liu C, Yu HY, et al. Wavelet-driven masked image modeling: a path to efficient visual representation[J]. Proc AAAI Conf Artif Intell, 2025, 39(8): 8611-9. doi:10.1609/aaai.v39i8.32930 |
| [38] | Liu C, Cheng YZ, Tamura S. Masked image modeling-based boundary reconstruction for 3D medical image segmentation[J]. Comput Biol Med, 2023, 166: 107526. doi:10.1016/j.compbiomed.2023.107526 |
| [39] | Sultana J, Qin RW, Yin ZZ. Seeing through expert's eyes: leveraging radiologist eye gaze and speech report with graph neural networks for chest X-ray image classification[C]//Computer Vision-ACCV 2024. Singapore: Springer, 2025: 142-58. doi:10.1007/978-981-96-0901-7_9 |
| [40] | Pershin I, Mustafaev T, Ibragimova D, et al. Changes in radiologists' gaze patterns against lung X-rays with different abnormalities: a randomized experiment[J]. J Digit Imaging, 2023, 36(3): 767-75. doi:10.1007/s10278-022-00760-2 |
| [41] | Cho K, Kim KD, Nam Y, et al. CheSS: chest X-ray pre-trained model via self-supervised contrastive learning[J]. J Digit Imaging, 2023, 36(3): 902-10. doi:10.1007/s10278-023-00782-4 |
| [42] | Shurrab S, Guerra-Manzanares A, Shamout FE. Multimodal masked Siamese network improves chest X-ray representation learning[J]. Sci Rep, 2024, 14: 22516. doi:10.1038/s41598-024-74043-x |
| [43] | Kong LJ, Ma MQ, Chen GY, et al. Understanding masked autoencoders via hierarchical latent variable models[C]//2023 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR). June 17-24, 2023, Vancouver, BC, Canada. IEEE, 2023: 7918-28. doi:10.1109/cvpr52729.2023.00765 |
| [44] | Yang QS, Li WY, Li BP, et al. MRM: masked relation modeling for medical image pre-training with genetics[C]//2023 IEEE/CVF International Conference on Computer Vision (ICCV). October 1-6, 2023, Paris, France. IEEE, 2024: 21395-405. doi:10.1109/iccv51070.2023.01961 |
| [45] | Dai PY, Ou YF, Yang YQ, et al. SaSaMIM: synthetic anatomical semantics-aware masked image modeling for colon tumor segmentation in non-contrast abdominal computed tomography[C]//Medical Image Computing and Computer Assisted Intervention- MICCAI 2024. Cham: Springer, 2024: 567-78. doi:10.1007/978-3-031-72120-5_53 |
| [46] | Xie YT, Gu L, Harada T, et al. MedIM: boost medical image representation via radiology report-guided masking[C]//Medical Image Computing and Computer Assisted Intervention-MICCAI 2023. Cham: Springer, 2023: 13-23. doi:10.1007/978-3-031-43907-0_2 |
| [47] | Wang WX, Wang J, Chen C, et al. FreMIM: Fourier transform meets masked image modeling for medical image segmentation[C]//2024 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV). January 3-8, 2024, Waikoloa, HI, USA. IEEE, 2024: 7845-55. doi:10.1109/wacv57701.2024.00768 |
| [48] | Xing ZH, Zhu L, Yu LQ, et al. Hybrid masked image modeling for 3D medical image segmentation[J]. IEEE J Biomed Health Inform, 2024, 28(4): 2115-25. doi:10.1109/jbhi.2024.3360239 |
| [1] | 梁业东, 朱雄峰, 黄美燕, 张文聪, 郭翰宇, 冯前进. CRAKUT:融合对比区域注意力机制与临床先验知识的U-Transformer用于放射学报告生成[J]. 南方医科大学学报, 2025, 45(6): 1343-1352. |
| [2] | 李青峰,邢潇丹,冯前进. 基于耦合的卷积-图卷积神经网络的阿尔茨海默病的磁共振诊断方法[J]. 南方医科大学学报, 2020, 40(04): 531-537. |
| [3] | 梁翠霞,李明强,边兆英,吕闻冰,曾栋,马建华. 基于深度学习特征的乳腺肿瘤分类模型评估[J]. 南方医科大学学报, 2019, 39(01): 88-. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
